6EGA
 
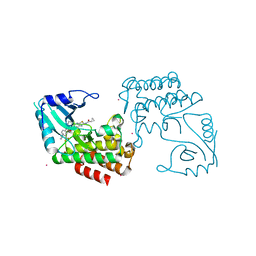 | | IRAK4 in complex with a type II inhibitor | | Descriptor: | 3-{2-[(cyclopropanecarbonyl)amino][1,3]thiazolo[5,4-b]pyridin-5-yl}-N-{4-[(piperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzamide, COBALT (II) ION, Interleukin-1 receptor-associated kinase 4, ... | | Authors: | Wang, L, Wu, H. | | Deposit date: | 2018-08-19 | | Release date: | 2019-02-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.512 Å) | | Cite: | Conformational flexibility and inhibitor binding to unphosphorylated interleukin-1 receptor-associated kinase 4 (IRAK4).
J.Biol.Chem., 294, 2019
|
|
6EGF
 
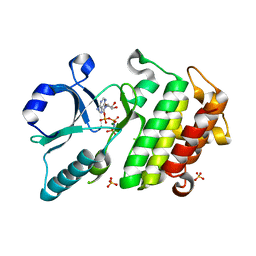 | |
7LF6
 
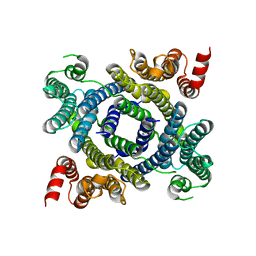 | | Structure of lysosomal membrane protein | | Descriptor: | Endosomal/lysosomal potassium channel TMEM175 | | Authors: | Shen, C, Fu, T.M, Wang, L.F, Rawson, S, Wu, H. | | Deposit date: | 2021-01-15 | | Release date: | 2022-01-26 | | Last modified: | 2023-08-09 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | pH regulates potassium conductance and drives a constitutive proton current in human TMEM175.
Sci Adv, 8, 2022
|
|
6EGD
 
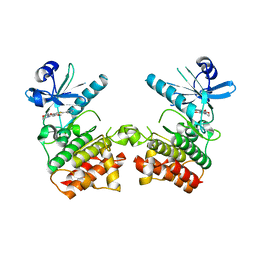 | |
6EGE
 
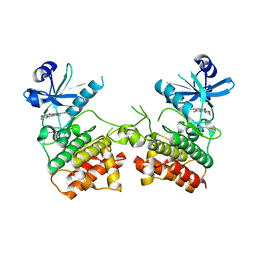 | |
6EG9
 
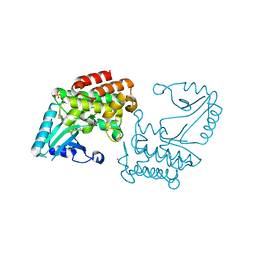 | | IRAK4 in complex with Ponatinib | | Descriptor: | 3-(imidazo[1,2-b]pyridazin-3-ylethynyl)-4-methyl-N-{4-[(4-methylpiperazin-1-yl)methyl]-3-(trifluoromethyl)phenyl}benzam ide, COBALT (II) ION, Interleukin-1 receptor-associated kinase 4, ... | | Authors: | Wang, L, Wu, H. | | Deposit date: | 2018-08-19 | | Release date: | 2019-02-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.414 Å) | | Cite: | Conformational flexibility and inhibitor binding to unphosphorylated interleukin-1 receptor-associated kinase 4 (IRAK4).
J.Biol.Chem., 294, 2019
|
|
7T32
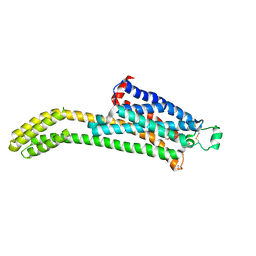 
 | | CryoEM structure of the adenosine 2A receptor-BRIL/Anti BRIL Fab complex with ZM241385 | | Descriptor: | 4-{2-[(7-amino-2-furan-2-yl[1,2,4]triazolo[1,5-a][1,3,5]triazin-5-yl)amino]ethyl}phenol, Adenosine receptor A2a/Soluble cytochrome b562 Fusion Protein | | Authors: | Zhang, K.H, Wu, H, Hoppe, N, Manglik, A, Cheng, Y.F. | | Deposit date: | 2021-12-06 | | Release date: | 2022-08-10 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Fusion protein strategies for cryo-EM study of G protein-coupled receptors.
Nat Commun, 13, 2022
|
|
3JBL
 
 | | Cryo-EM Structure of the Activated NAIP2/NLRC4 Inflammasome Reveals Nucleated Polymerization | | Descriptor: | NLR family CARD domain-containing protein 4 | | Authors: | Zhang, L, Chen, S, Ruan, J, Wu, J, Tong, A.B, Yin, Q, Li, Y, David, L, Lu, A, Wang, W.L, Marks, C, Ouyang, Q, Zhang, X, Mao, Y, Wu, H. | | Deposit date: | 2015-09-05 | | Release date: | 2015-10-21 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY (4.5 Å) | | Cite: | Cryo-EM structure of the activated NAIP2-NLRC4 inflammasome reveals nucleated polymerization.
Science, 350, 2015
|
|
1D0A
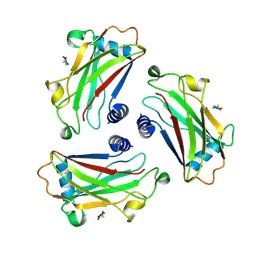 
 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 (TRAF2) IN COMPLEX WITH A HUMAN OX40 PEPTIDE | | Descriptor: | OX40L RECEPTOR PEPTIDE, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED PROTEIN 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-09 | | Release date: | 2000-03-08 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
1D0J
 
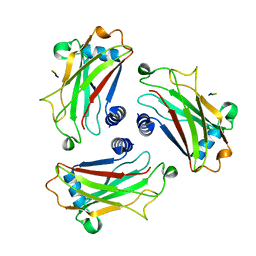 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 IN COMPLEX WITH A M4-1BB PEPTIDE | | Descriptor: | 4-1BB LIGAND RECEPTOR, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED PROTEIN 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-10 | | Release date: | 2000-03-08 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
1CZZ
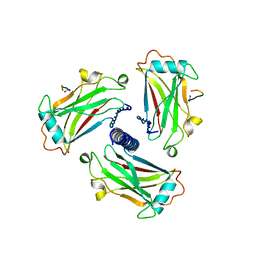 
 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 IN COMPLEX WITH A 17-RESIDUE CD40 PEPTIDE | | Descriptor: | CD 40 PEPTIDE, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED PROTEIN 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-07 | | Release date: | 2000-03-08 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
7RAV
 
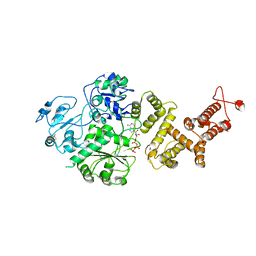 | | Cryo-EM structure of the unliganded form of NLR family apoptosis inhibitory protein 5 (NAIP5) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Baculoviral IAP repeat-containing protein 1e | | Authors: | Paidimuddala, B, Cao, J, Xie, Q, Wu, H, Zhang, L. | | Deposit date: | 2021-07-02 | | Release date: | 2023-01-11 | | Last modified: | 2023-03-01 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Mechanism of NAIP-NLRC4 inflammasome activation revealed by cryo-EM structure of unliganded NAIP5.
Nat.Struct.Mol.Biol., 30, 2023
|
|
1D01
 
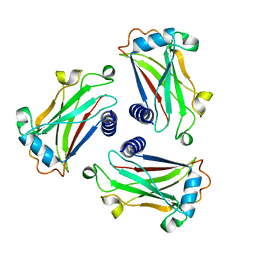 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 IN COMPLEX WITH A HUMAN CD30 PEPTIDE | | Descriptor: | CD30 PEPTIDE, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED FACTOR 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-07 | | Release date: | 2003-12-02 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
1D00
 
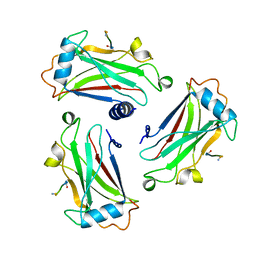 | | STRUCTURE OF TNF RECEPTOR ASSOCIATED FACTOR 2 IN COMPLEX WITH A 5-RESIDUE CD40 PEPTIDE | | Descriptor: | B-CELL SURFACE ANTIGEN CD40, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED PROTEIN 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-07 | | Release date: | 2000-03-29 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
6WM4
 
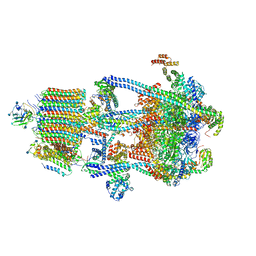 | | Human V-ATPase in state 3 with SidK and ADP | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ADENOSINE-5'-DIPHOSPHATE, Renin receptor, ... | | Authors: | Wang, L, Wu, H, Fu, T.M. | | Deposit date: | 2020-04-20 | | Release date: | 2020-11-11 | | Last modified: | 2020-11-18 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structures of a Complete Human V-ATPase Reveal Mechanisms of Its Assembly.
Mol.Cell, 80, 2020
|
|
6WLZ
 
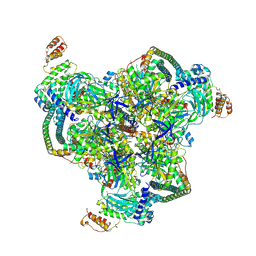 | | The V1 region of human V-ATPase in state 1 (focused refinement) | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, SidK, V-type proton ATPase catalytic subunit A, ... | | Authors: | Wang, L, Wu, H, Fu, T.M. | | Deposit date: | 2020-04-20 | | Release date: | 2020-11-11 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structures of a Complete Human V-ATPase Reveal Mechanisms of Its Assembly.
Mol.Cell, 80, 2020
|
|
6WLW
 
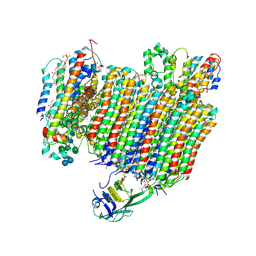 | | The Vo region of human V-ATPase in state 1 (focused refinement) | | Descriptor: | (2~{S})-2-$l^{4}-azanyl-3-[[(2~{R})-3-octadecanoyloxy-2-oxidanyl-propoxy]-oxidanyl-oxidanylidene-$l^{6}-phosphanyl]oxy-propanoic acid, 1,2-DICAPROYL-SN-PHOSPHATIDYL-L-SERINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Wang, L, Wu, H, Fu, T.-M. | | Deposit date: | 2020-04-20 | | Release date: | 2020-11-11 | | Last modified: | 2020-11-25 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structures of a Complete Human V-ATPase Reveal Mechanisms of Its Assembly.
Mol.Cell, 80, 2020
|
|
6WM2
 
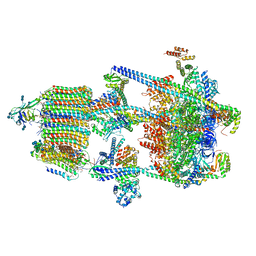 | | Human V-ATPase in state 1 with SidK and ADP | | Descriptor: | (2~{S})-2-$l^{4}-azanyl-3-[[(2~{R})-3-octadecanoyloxy-2-oxidanyl-propoxy]-oxidanyl-oxidanylidene-$l^{6}-phosphanyl]oxy-propanoic acid, 1,2-DICAPROYL-SN-PHOSPHATIDYL-L-SERINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Wang, L, Wu, H, Fu, T.M. | | Deposit date: | 2020-04-20 | | Release date: | 2020-11-11 | | Last modified: | 2020-11-25 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structures of a Complete Human V-ATPase Reveal Mechanisms of Its Assembly.
Mol.Cell, 80, 2020
|
|
1CZY
 
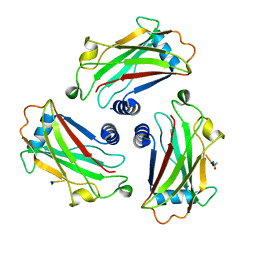 | | CRYSTAL STRUCTURE OF THE COMPLEX BETWEEN THE TRAF DOMAIN OF HUMAN TRAF2 AND AN LMP1 BINDING PEPTIDE | | Descriptor: | LATENT MEMBRANE PROTEIN 1, TUMOR NECROSIS FACTOR RECEPTOR ASSOCIATED PROTEIN 2 | | Authors: | Ye, H, Park, Y.C, Kreishman, M, Kieff, E, Wu, H. | | Deposit date: | 1999-09-07 | | Release date: | 2000-03-08 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structural basis for the recognition of diverse receptor sequences by TRAF2.
Mol.Cell, 4, 1999
|
|
1WM7
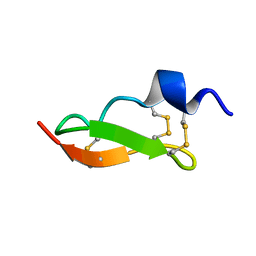 
 | | Solution Structure of BmP01 from the Venom of Scorpion Buthus martensii Karsch, 9 structures | | Descriptor: | Neurotoxin BmP01 | | Authors: | Wu, G, Li, Y, Wei, D, He, F, Jiang, S, Hu, G, Wu, H, Chen, X. | | Deposit date: | 2004-07-05 | | Release date: | 2004-07-27 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of BmP01 from the Venom of Scorpion Buthus martensii Karsch
Biochem.Biophys.Res.Commun., 276, 2000
|
|
7LFH
 
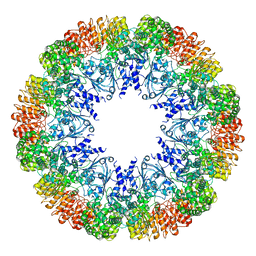 | |
2EX4
 
 | | Crystal Structure of Human methyltransferase AD-003 in complex with S-adenosyl-L-homocysteine | | Descriptor: | S-ADENOSYL-L-HOMOCYSTEINE, adrenal gland protein AD-003 | | Authors: | Min, J.R, Wu, H, Zeng, H, Loppnau, P, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Plotnikov, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2005-11-07 | | Release date: | 2005-11-15 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Crystal Structure of Human AD-003 protein in complex with S-adenosyl-L-homocysteine
To be Published
|
|
1M2S
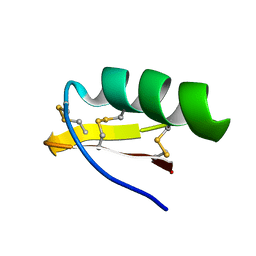 
 | | Solution Structure of A New Potassium Channels Blocker from the Venom of Chinese Scorpion Buthus martensi Karsch | | Descriptor: | Toxin BmTX3 | | Authors: | Wang, Y, Li, M, Zhang, N, Wu, G, Hu, G, Wu, H. | | Deposit date: | 2002-06-25 | | Release date: | 2004-04-06 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The solution structure of BmTx3B, a member of the scorpion toxin subfamily alpha-KTx 16
Proteins, 58, 2005
|
|
6NHW
 
 | | Structure of the transmembrane domain of the Death Receptor 5 - Dimer of Trimer | | Descriptor: | Tumor necrosis factor receptor superfamily member 10B | | Authors: | Chou, J.J, Pan, L, Fu, Q, Zhao, L, Chen, W, Piai, A, Fu, T, Wu, H. | | Deposit date: | 2018-12-24 | | Release date: | 2019-02-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Higher-Order Clustering of the Transmembrane Anchor of DR5 Drives Signaling.
Cell, 176, 2019
|
|
6NHY
 
 | | Structure of the transmembrane domain of the Death Receptor 5 mutant (G217Y) - Trimer Only | | Descriptor: | Tumor necrosis factor receptor superfamily member 10B | | Authors: | Chou, J.J, Pan, L, Zhao, L, Chen, W, Piai, A, Fu, T, Wu, H, Liu, Z. | | Deposit date: | 2018-12-24 | | Release date: | 2019-02-27 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Higher-Order Clustering of the Transmembrane Anchor of DR5 Drives Signaling.
Cell, 176, 2019
|
|
