6B7Q
 
 | |
6B7M
 
 | |
6B7O
 
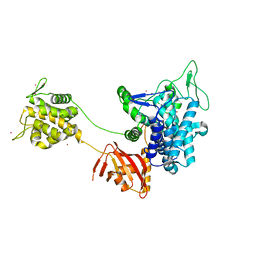
 | | Crystal structure of Legionella effector sdeD (lpg2509) H67A in complex with ADP-ribosylated Ubiquitin | | Descriptor: | Polyubiquitin-C, SdeD (lpg2509) H67A, [(2R,3S,4R,5R)-5-(6-AMINOPURIN-9-YL)-3,4-DIHYDROXY-OXOLAN-2-YL]METHYL [HYDROXY-[[(2R,3S,4R,5S)-3,4,5-TRIHYDROXYOXOLAN-2-YL]METHOXY]PHOSPHORYL] HYDROGEN PHOSPHATE | | Authors: | Mao, Y, Akturk, A, Wasilko, J. | | Deposit date: | 2017-10-04 | | Release date: | 2018-04-18 | | Last modified: | 2020-01-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector.
Nature, 557, 2018
|
|
7X8G
 
 | |
7X8B
 
 | |
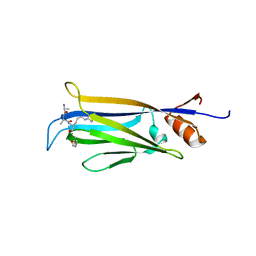
7X8F
 | |
7W5H
 
 | | The structure of trichobrasilenol synthase TaTC6 in complex with FsPP | | Descriptor: | GLYCEROL, MAGNESIUM ION, MALONATE ION, ... | | Authors: | Chen, C, Wang, T, Yang, Y, Zhang, L, Ko, T, Huang, J, Guo, R. | | Deposit date: | 2021-11-30 | | Release date: | 2022-08-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural insights into the cyclization of unusual brasilane-type sesquiterpenes.
Int.J.Biol.Macromol., 209, 2022
|
|
7FBB
 
 | | De novo design protein D12 with MBP tag | | Descriptor: | Maltodextrin-binding protein,de novo designed protein D12 | | Authors: | Bin, H. | | Deposit date: | 2021-07-09 | | Release date: | 2021-12-22 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.307 Å) | | Cite: | A backbone-centred energy function of neural networks for protein design.
Nature, 602, 2022
|
|
7FBD
 
 | | De novo design protein D53 with MBP tag | | Descriptor: | Maltodextrin-binding protein,De novo design protein D53 | | Authors: | Bin, H. | | Deposit date: | 2021-07-09 | | Release date: | 2021-12-22 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | A backbone-centred energy function of neural networks for protein design.
Nature, 602, 2022
|
|
7FBC
 
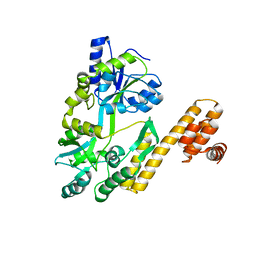 | | De novo design protein D22 with MBP tag | | Descriptor: | Maltodextrin-binding protein,De novo design protein D22 | | Authors: | Bin, H. | | Deposit date: | 2021-07-09 | | Release date: | 2021-12-22 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A backbone-centred energy function of neural networks for protein design.
Nature, 602, 2022
|
|
4LVA
 
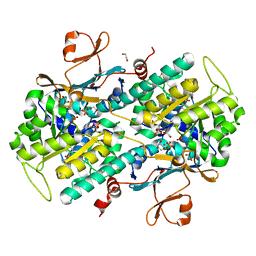 | | Fragment-based Identification of Amides Derived From trans-2-(Pyridin-3-yl)cyclopropanecarboxylic Acid as Potent Inhibitors of Human Nicotinamide Phosphoribosyltransferase (NAMPT) | | Descriptor: | 1,2-ETHANEDIOL, N-(4-{[4-(pyrrolidin-1-yl)piperidin-1-yl]sulfonyl}benzyl)-2H-pyrido[4,3-e][1,2,4]thiadiazin-3-amine 1,1-dioxide, Nicotinamide phosphoribosyltransferase, ... | | Authors: | Giannetti, A.M, Zheng, X, Skelton, N, Wang, W, Bravo, B, Feng, Y, Gunzner-Toste, J, Ho, Y, Hua, R, Wang, C, Zhao, Q, Liederer, B.M, Liu, Y, O'Brien, T, Oeh, J, Sampath, D, Shen, Y, Wang, L, Wu, H, Xiao, Y, Yuen, P, Zak, M, Zhao, G, Dragovich, P.S. | | Deposit date: | 2013-07-26 | | Release date: | 2013-09-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Identification of amides derived from 1H-pyrazolo[3,4-b]pyridine-5-carboxylic acid as potent inhibitors of human nicotinamide phosphoribosyltransferase (NAMPT).
Bioorg.Med.Chem.Lett., 23, 2013
|
|
5ZK1
 
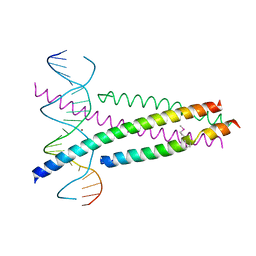 | | Crystal Structure of the CRTC2(SeMet)-CREB-CRE complex | | Descriptor: | CREB-regulated transcription coactivator 2, Cyclic AMP-responsive element-binding protein 1, DNA (5'-D(*CP*TP*TP*GP*GP*CP*TP*GP*AP*CP*GP*TP*CP*AP*GP*CP*CP*AP*AP*G)-3'), ... | | Authors: | Xiang, S, Zhai, L, Valencia-Swain, J. | | Deposit date: | 2018-03-22 | | Release date: | 2018-06-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structural Insights into the CRTC2-CREB Complex Assembly on CRE.
J. Mol. Biol., 430, 2018
|
|
5ZKO
 
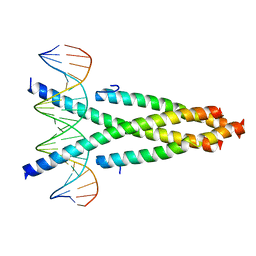 | | Crystal structure of the CRTC2-CREB-CRE complex | | Descriptor: | CREB-regulated transcription coactivator 2, Cyclic AMP-responsive element-binding protein 1, DNA (5'-D(*CP*TP*TP*GP*GP*CP*TP*GP*AP*CP*GP*TP*CP*AP*GP*CP*CP*AP*AP*G)-3') | | Authors: | Xiang, S, Zhai, L, Valecia-Swain, J. | | Deposit date: | 2018-03-24 | | Release date: | 2018-06-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structural Insights into the CRTC2-CREB Complex Assembly on CRE.
J. Mol. Biol., 430, 2018
|
|
7XC3
 
 | |
4FQ1
 
 | | Crystal Structure of PGT121 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Fab heavy chain, Fab light chain | | Authors: | Scharf, L, Bjorkman, P.J. | | Deposit date: | 2012-06-24 | | Release date: | 2012-11-14 | | Last modified: | 2021-05-19 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Complex-type N-glycan recognition by potent broadly neutralizing HIV antibodies.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
7XP1
 
 | | Crystal structure of PmiR from Pseudomonas aeruginosa | | Descriptor: | ALPHA-METHYLISOCITRIC ACID, GLYCEROL, Probable transcriptional regulator, ... | | Authors: | Zhang, Y.X, Liang, H.H, Gan, J.H. | | Deposit date: | 2022-05-02 | | Release date: | 2023-04-12 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | PmiR senses 2-methylisocitrate levels to regulate bacterial virulence in Pseudomonas aeruginosa.
Sci Adv, 8, 2022
|
|
2M4X
 
 | |
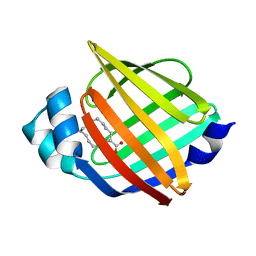
7YF1
 | |
2M4Z
 
 | |
8HB2
 
 | |
8HBB
 
 | |
8HAZ
 
 | |
8G9J
 
 | |
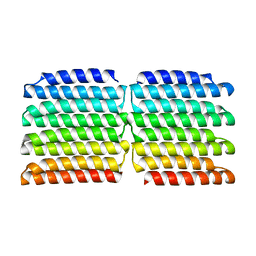
8G9K
 | |
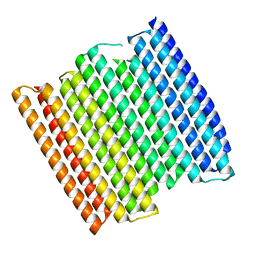
8GA6
 | |
