5TCZ
 
 | |
5K9R
 
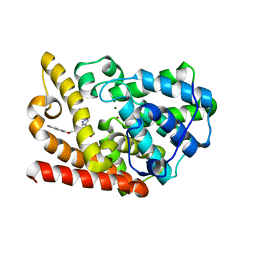 | | PDE10a with imidazopyrazine inhibitor | | Descriptor: | 4-[5-[1-(2-methoxyethyl)pyrazol-4-yl]-2-(quinolin-2-yloxymethyl)imidazo[1,2-a]pyrazin-8-yl]morpholine, MAGNESIUM ION, ZINC ION, ... | | Authors: | Gibbs, A.G, Schubert, C. | | Deposit date: | 2016-06-01 | | Release date: | 2016-08-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery of novel potent imidazo[1,2-b]pyridazine PDE10a inhibitors.
Bioorg.Med.Chem.Lett., 26, 2016
|
|
2M4Z
 
 | |
2M50
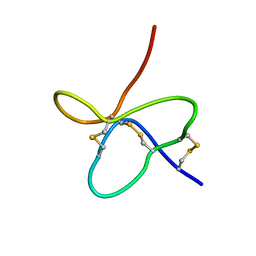 
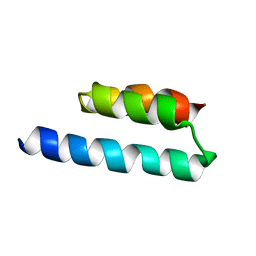
 | | Analysis of the structural and molecular basis of voltage-sensitive sodium channel inhibition by the spider toxin, Huwentoxin-IV (-TRTX-Hh2a). | | Descriptor: | Mu-theraphotoxin-Hh2a | | Authors: | Gibbs, A, Minassian, N, Flinspach, M, Wickenden, A. | | Deposit date: | 2013-02-12 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-29 | | Method: | SOLUTION NMR | | Cite: | Analysis of the Structural and Molecular Basis of Voltage-sensitive Sodium Channel Inhibition by the Spider Toxin Huwentoxin-IV ( mu-TRTX-Hh2a).
J.Biol.Chem., 288, 2013
|
|
2M4X
 
 | |
2N35
 
 | |
2M9P
 
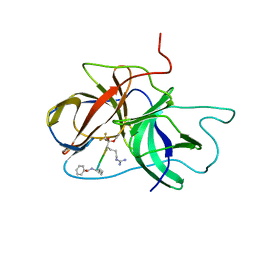 | |
2M9Q
 
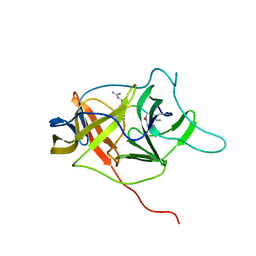 | |
3RO4
 
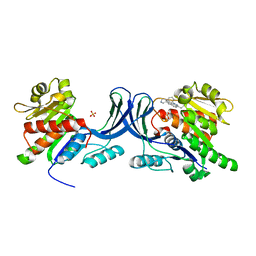 | |
3NBW
 
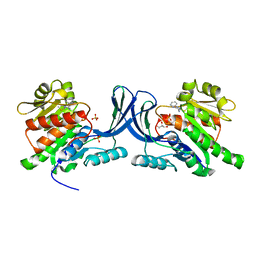 | | X-ray structure of ketohexokinase in complex with a pyrazole compound | | Descriptor: | 5-amino-3-(methylsulfanyl)-1-phenyl-1H-pyrazole-4-carbonitrile, GLYCEROL, Ketohexokinase, ... | | Authors: | Abad, M.C, Gibbs, A.C, Spurlino, J.C. | | Deposit date: | 2010-06-04 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.341 Å) | | Cite: | Electron density guided fragment-based lead discovery of ketohexokinase inhibitors.
J.Med.Chem., 53, 2010
|
|
3NC2
 
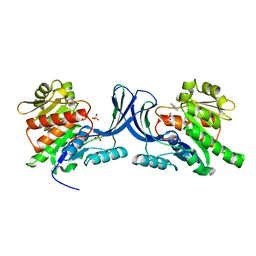 | |
3NCA
 
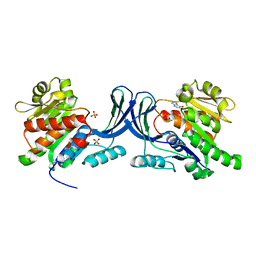 | | X-ray structure of ketohexokinase in complex with a thieno pyridinol compound | | Descriptor: | Ketohexokinase, SULFATE ION, thieno[3,2-b]pyridin-7-ol | | Authors: | Abad, M.C, Gibbs, A.C, Spurlino, J.C. | | Deposit date: | 2010-06-04 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Electron density guided fragment-based lead discovery of ketohexokinase inhibitors.
J.Med.Chem., 53, 2010
|
|
3NC9
 
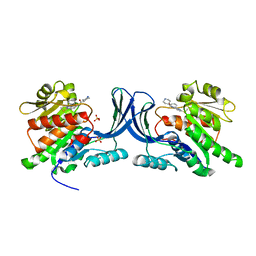 | |
3NBV
 
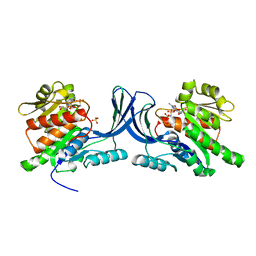 | |
1RY3
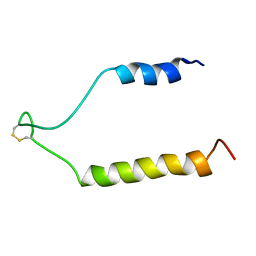 
 | | NMR Solution Structure of the Precursor for Carnobacteriocin B2, an Antimicrobial Peptide from Carnobacterium piscicola | | Descriptor: | Bacteriocin carnobacteriocin B2 | | Authors: | Sprules, T, Kawulka, K.E, Gibbs, A.C, Wishart, D.S, Vederas, J.C. | | Deposit date: | 2003-12-19 | | Release date: | 2004-05-04 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the precursor for carnobacteriocin B2, an antimicrobial peptide from Carnobacterium piscicola.
Eur.J.Biochem., 271, 2004
|
|
1CW6
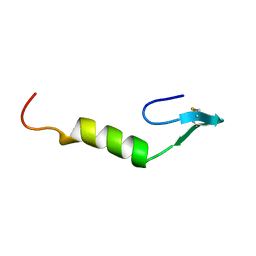 
 | | REFINED SOLUTION STRUCTURE OF LEUCOCIN A | | Descriptor: | TYPE IIA BACTERIOCIN LEUCOCIN A | | Authors: | Wang, Y, Henz, M.E, Gallagher, N.L.F, Chai, S, Yan, L.Z, Gibbs, A.C, Stiles, M.E, Wishart, D.S, Vederas, J.C. | | Deposit date: | 1999-08-25 | | Release date: | 1999-09-08 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of carnobacteriocin B2 and implications for structure-activity relationships among type IIa bacteriocins from lactic acid bacteria.
Biochemistry, 38, 1999
|
|
1CW5
 
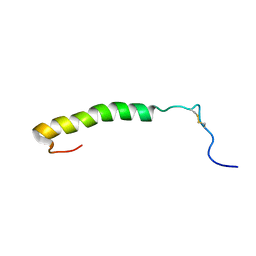 | | SOLUTION STRUCTURE OF CARNOBACTERIOCIN B2 | | Descriptor: | TYPE IIA BACTERIOCIN CARNOBACTERIOCIN B2 | | Authors: | Wang, Y, Henz, M.E, Gallagher, N.L.F, Chai, S, Yan, L.Z, Gibbs, A.C, Stiles, M.E, Wishart, D.S, Vederas, J.C. | | Deposit date: | 1999-08-25 | | Release date: | 1999-09-07 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of carnobacteriocin B2 and implications for structure-activity relationships among type IIa bacteriocins from lactic acid bacteria.
Biochemistry, 38, 1999
|
|
