6HNN
 
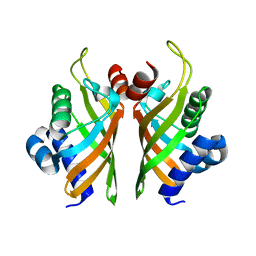 | | Crystal structure of wild-type IdmH, a putative polyketide cyclase from Streptomyces antibioticus | | Descriptor: | Putative polyketide cyclase IdmH | | Authors: | Drulyte, I, Obajdin, J, Trinh, C, Hemsworth, G.R, Berry, A. | | Deposit date: | 2018-09-16 | | Release date: | 2019-11-06 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the putative cyclase IdmH from the indanomycin nonribosomal peptide synthase/polyketide synthase.
Iucrj, 6, 2019
|
|
6RSY
 
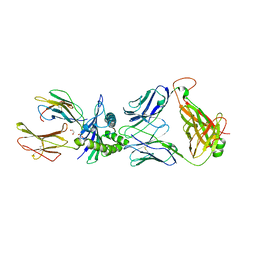 | |
7ZT2
 
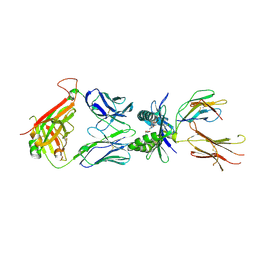 | | Structure of E8 TCR in complex with human MR1 bound to 5-OP-RU | | Descriptor: | 1,2-ETHANEDIOL, 1-deoxy-1-({2,6-dioxo-5-[(E)-propylideneamino]-1,2,3,6-tetrahydropyrimidin-4-yl}amino)-D-ribitol, Beta-2-microglobulin, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT9
 
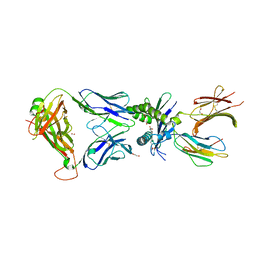 | | Structure of E8 TCR in complex in human MR1 bound to 4FBA | | Descriptor: | 1,2-ETHANEDIOL, 4-METHYLBENZOIC ACID, Beta-2-microglobulin, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT4
 
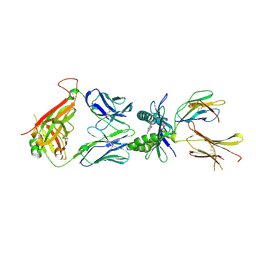 | | Structure of E8 TCR in complex with human MR1 bound to 6FP | | Descriptor: | 2-azanyl-6-methyl-3~{H}-pteridin-4-one, Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, ... | | Authors: | Karuppiah, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT8
 
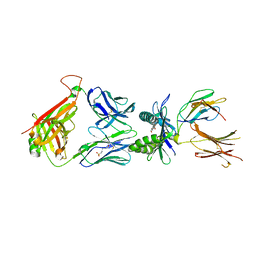 | | Structure of E8 TCR in complex in human MR1 bound to 3FBA | | Descriptor: | 1,2-ETHANEDIOL, 3-methylbenzoic acid, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT5
 
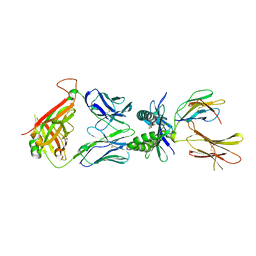 | | Structure of E8 TCR in complex in human MR1 bound to 3FSA | | Descriptor: | 3-methanoyl-2-oxidanyl-benzoic acid, Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, ... | | Authors: | Karuppiah, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT3
 
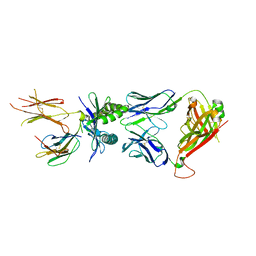 | | Structure of E8 TCR in complex in human MR1 K43A | | Descriptor: | Beta-2-microglobulin, Major histocompatibility complex class I-related gene protein, TCR alpha, ... | | Authors: | Karuppiah, V, Srikannathasan, V, Robinson, R.A. | | Deposit date: | 2022-05-09 | | Release date: | 2023-06-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
J.Exp.Med., 220, 2023
|
|
7ZT7
 
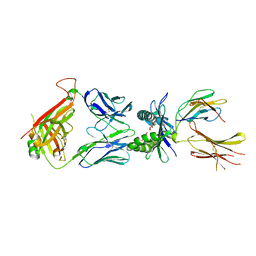 | |
6TMO
 
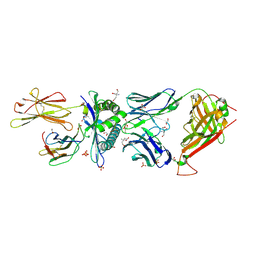 | | Structure determination of an enhanced affinity TCR, a24b17, in complex with HLA-A*02:01 presenting a MART-1 peptide, EAAGIGILTV | | Descriptor: | 1,2-ETHANEDIOL, Alpha chain of engineered high affinity T-cell receptor, Beta chain of high affinity engineered T-cell receptor, ... | | Authors: | Rizkallah, P.J, Cole, D.K. | | Deposit date: | 2019-12-05 | | Release date: | 2020-10-07 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Molecular Rules Underpinning Enhanced Affinity Binding of Human T Cell Receptors Engineered for Immunotherapy.
Mol Ther Oncolytics, 18, 2020
|
|
4BWL
 
 | | Structure of the Y137A mutant of E. coli N-acetylneuraminic acid lyase in complex with pyruvate, N-acetyl-D-mannosamine and N- acetylneuraminic acid | | Descriptor: | 2-(ACETYLAMINO)-2-DEOXY-D-MANNOSE, 5-(acetylamino)-3,5-dideoxy-D-glycero-D-galacto-non-2-ulosonic acid, N-ACETYLNEURAMINATE LYASE, ... | | Authors: | Campeotto, I, Phillips, S.E.V, Pearson, A.R, Nelson, A, Berry, A. | | Deposit date: | 2013-07-03 | | Release date: | 2014-02-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Reaction Mechanism of N-Acetylneuraminic Acid Lyase Revealed by a Combination of Crystallography, Qm/Mm Simulation and Mutagenesis.
Acs Chem.Biol., 9, 2014
|
|
4EV4
 
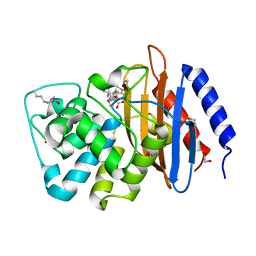 | | Crystal structure of serratia fonticola carbapenemase SFC-1 E166A mutant with the acylenzyme intermediate of meropenem | | Descriptor: | (4R,5S)-3-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-5-[(2S,3R)-3-hydroxy-1-oxobutan-2-yl]-4-methyl-4,5-d ihydro-1H-pyrrole-2-carboxylic acid, 1,2-ETHANEDIOL, Carbapenem-hydrolizing beta-lactamase SFC-1 | | Authors: | Fonseca, F, Spencer, J. | | Deposit date: | 2012-04-25 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The basis for carbapenem hydrolysis by class A beta-lactamases: a combined investigation using crystallography and simulations.
J.Am.Chem.Soc., 134, 2012
|
|
4EQI
 
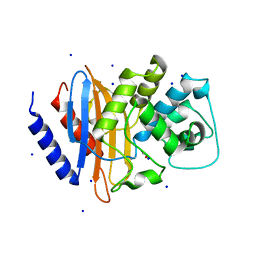 | | Crystal structure of serratia fonticola carbapenemase SFC-1 | | Descriptor: | 1,2-ETHANEDIOL, Carbapenem-hydrolizing beta-lactamase SFC-1, SODIUM ION | | Authors: | Fonseca, F, Spencer, J. | | Deposit date: | 2012-04-18 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | The basis for carbapenem hydrolysis by class A beta-lactamases: a combined investigation using crystallography and simulations.
J.Am.Chem.Soc., 134, 2012
|
|
4EUZ
 
 | | Crystal structure of serratia fonticola carbapenemase SFC-1 S70A-Meropenem complex | | Descriptor: | (4R,5S,6S)-3-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-6-[(1R)-1-hydroxyethyl]-4-methyl-7-oxo-1-azabicyclo[3.2.0]hept-2-ene-2-carboxylic acid, 1,2-ETHANEDIOL, Carbapenem-hydrolizing beta-lactamase SFC-1, ... | | Authors: | Fonseca, F, Spencer, J. | | Deposit date: | 2012-04-25 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | The basis for carbapenem hydrolysis by class A beta-lactamases: a combined investigation using crystallography and simulations.
J.Am.Chem.Soc., 134, 2012
|
|
