[English] 日本語
 Yorodumi
Yorodumi- EMDB-0243: Improved model derived from cryo-EM map of Staphylococcus aureus ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0243 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Improved model derived from cryo-EM map of Staphylococcus aureus large ribosomal subunit | |||||||||
 Map data Map data | resolution 2.65 A | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosomal RNA / ribosome / pathogen / ribosomal protein / cryo-EM | |||||||||
| Function / homology |  Function and homology information Function and homology informationlarge ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding ...large ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / mRNA binding / RNA binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.65 Å | |||||||||
 Authors Authors | Eyal Z / Cimicata G | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2015 Journal: Proc Natl Acad Sci U S A / Year: 2015Title: Structural insights into species-specific features of the ribosome from the pathogen Staphylococcus aureus. Authors: Zohar Eyal / Donna Matzov / Miri Krupkin / Itai Wekselman / Susanne Paukner / Ella Zimmerman / Haim Rozenberg / Anat Bashan / Ada Yonath /   Abstract: The emergence of bacterial multidrug resistance to antibiotics threatens to cause regression to the preantibiotic era. Here we present the crystal structure of the large ribosomal subunit from ...The emergence of bacterial multidrug resistance to antibiotics threatens to cause regression to the preantibiotic era. Here we present the crystal structure of the large ribosomal subunit from Staphylococcus aureus, a versatile Gram-positive aggressive pathogen, and its complexes with the known antibiotics linezolid and telithromycin, as well as with a new, highly potent pleuromutilin derivative, BC-3205. These crystal structures shed light on specific structural motifs of the S. aureus ribosome and the binding modes of the aforementioned antibiotics. Moreover, by analyzing the ribosome structure and comparing it with those of nonpathogenic bacterial models, we identified some unique internal and peripheral structural motifs that may be potential candidates for improving known antibiotics and for use in the design of selective antibiotic drugs against S. aureus. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0243.map.gz emd_0243.map.gz | 126.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0243-v30.xml emd-0243-v30.xml emd-0243.xml emd-0243.xml | 51.9 KB 51.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_0243.png emd_0243.png | 164.4 KB | ||
| Filedesc metadata |  emd-0243.cif.gz emd-0243.cif.gz | 10.6 KB | ||
| Others |  emd_0243_additional.map.gz emd_0243_additional.map.gz emd_0243_half_map_1.map.gz emd_0243_half_map_1.map.gz emd_0243_half_map_2.map.gz emd_0243_half_map_2.map.gz | 105.9 MB 106.1 MB 106.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0243 http://ftp.pdbj.org/pub/emdb/structures/EMD-0243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0243 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0243 | HTTPS FTP |
-Related structure data
| Related structure data |  6hmaMC  7asmC  7asnC  7asoC  7aspC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
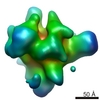
Map
| File |  Download / File: emd_0243.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0243.map.gz / Format: CCP4 / Size: 134.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | resolution 2.65 A | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.858 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
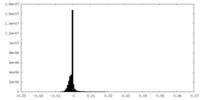
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: resolution 2.87 A
| File | emd_0243_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | resolution 2.87 A | ||||||||||||
| Projections & Slices |
| ||||||||||||
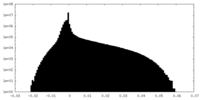
| Density Histograms |
-Half map: #2
| File | emd_0243_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
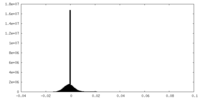
| Density Histograms |
-Half map: #1
| File | emd_0243_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
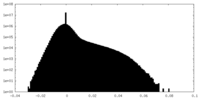
| Density Histograms |
- Sample components
Sample components
+Entire : Staphylococcus aureus large ribosomal subunit
+Supramolecule #1: Staphylococcus aureus large ribosomal subunit
+Macromolecule #1: 23S ribosomal RNA
+Macromolecule #2: 5S ribosomal RNA
+Macromolecule #3: 50S ribosomal protein L2
+Macromolecule #4: 50S ribosomal protein L3
+Macromolecule #5: 50S ribosomal protein L4
+Macromolecule #6: 50S ribosomal protein L5
+Macromolecule #7: 50S ribosomal protein L6
+Macromolecule #8: 50S ribosomal protein L13
+Macromolecule #9: 50S ribosomal protein L14
+Macromolecule #10: 50S ribosomal protein L15
+Macromolecule #11: 50S ribosomal protein L16
+Macromolecule #12: 50S ribosomal protein L17
+Macromolecule #13: 50S ribosomal protein L18
+Macromolecule #14: 50S ribosomal protein L19
+Macromolecule #15: 50S ribosomal protein L20
+Macromolecule #16: 50S ribosomal protein L21
+Macromolecule #17: 50S ribosomal protein L22
+Macromolecule #18: 50S ribosomal protein L23
+Macromolecule #19: 50S ribosomal protein L24
+Macromolecule #20: 50S ribosomal protein L25
+Macromolecule #21: 50S ribosomal protein L27
+Macromolecule #22: 50S ribosomal protein L28
+Macromolecule #23: 50S ribosomal protein L29
+Macromolecule #24: 50S ribosomal protein L30
+Macromolecule #25: 50S ribosomal protein L32
+Macromolecule #26: 50S ribosomal protein L33
+Macromolecule #27: 50S ribosomal protein L34
+Macromolecule #28: 50S ribosomal protein L35
+Macromolecule #29: 50S ribosomal protein L36
+Macromolecule #30: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.6 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)