-Search query
-Search result
Showing 1 - 50 of 111 items for (author: yvon & m)

EMDB-52631: 
Structure of the Chaetomium thermophilum Pmt4 homodimer (C2 symmetry)
Method: single particle / : McDowell MA, Wild K, Sinning I

EMDB-52632: 
Structure of the Chaetomium thermophilum Pmt4 homodimer (C1 symmetry)
Method: single particle / : McDowell MA, Wild K, Sinning I

PDB-9i5k: 
Structure of the Chaetomium thermophilum Pmt4 homodimer (C2 symmetry)
Method: single particle / : McDowell MA, Wild K, Sinning I

PDB-9i5l: 
Structure of the Chaetomium thermophilum Pmt4 homodimer (C1 symmetry)
Method: single particle / : McDowell MA, Wild K, Sinning I

EMDB-50850: 
TRPC4 in complex with E-AzPico
Method: single particle / : Vinayagam D, Raunser S

EMDB-50851: 
TRPC4 in complex with Z-AzPico
Method: single particle / : Vinayagam D, Raunser S

PDB-9fxl: 
TRPC4 in complex with E-AzPico
Method: single particle / : Vinayagam D, Raunser S

PDB-9fxm: 
TRPC4 in complex with Z-AzPico
Method: single particle / : Vinayagam D, Raunser S

EMDB-52168: 
LysR Type Transcriptional Regulator LsrB from Agrobacterium tumefaciens
Method: single particle / : Elders H, Schmidt JJ, Fiedler R, Hofmann E, Narberhaus F

PDB-9hh1: 
LysR Type Transcriptional Regulator LsrB from Agrobacterium tumefaciens
Method: single particle / : Elders H, Schmidt JJ, Fiedler R, Hofmann E, Narberhaus F

EMDB-42524: 
Cryo-EM Structure of Full-Length Spike Protein of Omicron XBB.1.5
Method: single particle / : Huynh KW, Chang JS, Fennell KF, Che Y, Wu H

PDB-8usz: 
Cryo-EM Structure of Full-Length Spike Protein of Omicron XBB.1.5
Method: single particle / : Huynh KW, Chang JS, Fennell KF, Che Y, Wu H

EMDB-43813: 
VIR-7229 Fab fragment bound the SARS-CoV-2 BA.2.86 spike trimer (local refinement of the BA 2.86 RBD/VIR-7229 VHVL)
Method: single particle / : Park YJ, Tortorici MA, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-43842: 
VIR-7229 Fab fragment bound the BA.2.86 spike trimer (global refinement)
Method: single particle / : Tortorici MA, Park YJ, Veelser D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-9asd: 
VIR-7229 Fab fragment bound the SARS-CoV-2 BA.2.86 spike trimer (local refinement of the BA 2.86 RBD/VIR-7229 VHVL)
Method: single particle / : Park YJ, Tortorici MA, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

PDB-9au2: 
VIR-7229 Fab fragment bound the BA.2.86 spike trimer (global refinement)
Method: single particle / : Tortorici MA, Park YJ, Veelser D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-18680: 
FZD3 in complex with nanobody 9
Method: single particle / : Zhao Y, Jones EY

EMDB-50071: 
Cryo-EM structure of the icosahedral lumazine synthase from Vicia faba.
Method: single particle / : Chee M, Trapani S, Hoh F, Lai Kee Him J, Yvon M, Blanc S, Bron P

PDB-9ez8: 
Cryo-EM structure of the icosahedral lumazine synthase from Vicia faba.
Method: single particle / : Chee M, Trapani S, Hoh F, Lai Kee Him J, Yvon M, Blanc S, Bron P

EMDB-15903: 
Automated simulation-based refinement of maltoporin into a cryo-EM density
Method: single particle / : Yvonnesdotter L, Rovsnik U, Blau C, Lycksell M, Howard RJ, Lindahl E

PDB-8b7v: 
Automated simulation-based refinement of maltoporin into a cryo-EM density
Method: single particle / : Yvonnesdotter L, Rovsnik U, Blau C, Lycksell M, Howard RJ, Lindahl E

EMDB-16376: 
CryoEM structure of a tungsten-containing aldehyde oxidoreductase from Aromatoleum aromaticum
Method: single particle / : Winiarska A, Ramirez-Amador F, Hege D, Gemmecker Y, Prinz S, Hochberg G, Heider J, Szaleniec M, Schuller JM

PDB-8c0z: 
CryoEM structure of a tungsten-containing aldehyde oxidoreductase from Aromatoleum aromaticum
Method: single particle / : Winiarska A, Ramirez-Amador F, Hege D, Gemmecker Y, Prinz S, Hochberg G, Heider J, Szaleniec M, Schuller JM

EMDB-15894: 
KimA from B. subtilis with nucleotide second-messenger c-di-AMP bound
Method: single particle / : Vonck J, Wieferig JP

EMDB-15895: 
Upright KimA dimer with bound c-di-AMP from B. subtilis
Method: single particle / : Vonck J, Wieferig JP

PDB-8b70: 
KimA from B. subtilis with nucleotide second-messenger c-di-AMP bound
Method: single particle / : Vonck J, Wieferig JP

PDB-8b71: 
Upright KimA dimer with bound c-di-AMP from B. subtilis
Method: single particle / : Vonck J, Wieferig JP

EMDB-11953: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Composite Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-11954: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 2)
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14810: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Consensus Map
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-14811: 
SARS-CoV-2 S 2P trimer in complex with monovalent DARPin R2 (State 1) - Focused Refinement
Method: single particle / : Hurdiss DL, Drulyte I

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
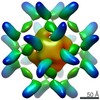
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-22995: 
Spike protein trimer
Method: single particle / : Asarnow D, Faust B, Bohn M, Bulkley D, Manglik A, Craik CS, Cheng Y

EMDB-13428: 
Pfu sArlG (PF0333, N-d31-ArlG)
Method: single particle / : Umrekar TR, Winterborn YB, Sivabalasarma S, Brantl J, Albers SV, Beeby M

EMDB-12675: 
GLIC, pentameric ligand gated ion channel, pH 3 State 2
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

EMDB-12677: 
GLIC, pentameric ligand-gated ion channel, pH 5, state 2
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

EMDB-12678: 
GLIC, pentameric ligand-gated ion channel, pH 7 state 2
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

EMDB-11202: 
GLIC pentameric ligand-gated ion channel, pH 7
Method: single particle / : Rovsnik U, Zhuang Y

EMDB-11208: 
GLIC pentameric ligand-gated ion channel, pH 5
Method: single particle / : Rovsnik U, Zhuang Y

EMDB-11209: 
GLIC pentameric ligand-gated ion channel, pH 3
Method: single particle / : Rovsnik U, Zhuang Y

PDB-6zgd: 
GLIC pentameric ligand-gated ion channel, pH 7
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

PDB-6zgj: 
GLIC pentameric ligand-gated ion channel, pH 5
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

PDB-6zgk: 
GLIC pentameric ligand-gated ion channel, pH 3
Method: single particle / : Rovsnik U, Zhuang Y, Forsberg BO, Carroni M, Yvonnesdotter L, Howard RJ, Lindahl E

EMDB-22993: 
SARS-CoV-2 spike glycoprotein:Fab 5A6 complex I
Method: single particle / : Asarnow D, Charles C, Cheng Y

EMDB-22994: 
SARS-CoV-2 Spike protein in complex with Fab 2H4
Method: single particle / : Asarnow D, Charles C, Cheng Y

EMDB-22997: 
SARS-CoV-2 spike glycoprotein:Fab 3D11 complex
Method: single particle / : Asarnow D, Charles C, Cheng Y
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
