-検索条件
-検索結果
検索 (著者・登録者: yang & yx)の結果103件中、1から50件目までを表示しています

EMDB-60254: 
Vesamicol-bound VAChT

EMDB-60255: 
Acetylcholine-bound VAChT

EMDB-35618: 
Cryo-EM structure of porcine bc1 complex in isolated state

EMDB-35461: 
Protomer 1 and 2 of the asymmetry trimer of the Cul2-Rbx1-EloBC-FEM1B ubiquitin ligase complex

EMDB-36182: 
An asymmetry dimer of the Cul2-Rbx1-EloBC-FEM1B ubiquitin ligase complexed with BEX2

EMDB-36183: 
Cryo-EM structure of neddylated Cul2-Rbx1-EloBC-FEM1B complexed with FNIP1-FLCN

EMDB-33990: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAbs 3E8 and 5E3 (localized refinement)
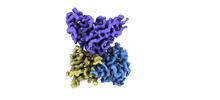
EMDB-33992: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAb 10E4 (localized refinement)
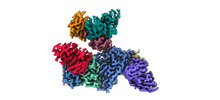
EMDB-33993: 
Cryo-EM density map of EBV gHgL-gp42 in complex with four mAbs 5E3, 3E8, 6H2 and 10E4
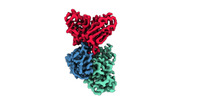
EMDB-33994: 
Cryo-EM structure of EBV gHgL-gp42 in complex with mAb 6H2 (localized refinement)

EMDB-34929: 
Monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex in a DNA binding form

EMDB-34927: 
Cryo-EM structure of monkeypox virus DNA replication holoenzyme F8, A22 and E4 complex without DNA at 2.76 angostram

EMDB-36342: 
Cryo-EM structure of the beta2AR-mBRIL/1b3 Fab/Glue complex with a partial agonist

EMDB-36360: 
cryo-EM structure of the beta2-AR-mBRIL/1b3 Fab/Glue complex with a full agonist

EMDB-36361: 
Cryo-EM structure of the beta2AR-mBRIL/1b3 Fab/Glue complex with an antagonist

EMDB-29903: 
Structure of LARP7 protein p65-telomerase RNA complex in telomerase

EMDB-33310: 
Cryo-EM structure of CopC-CaM-caspase-3 with NAD+

EMDB-33311: 
Cryo-EM structure of CopC-CaM-caspase-3 with ADPR

EMDB-33312: 
Cryo-EM structure of CopC-CaM-caspase-3 with ADPR-deacylization
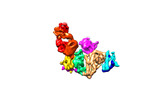
EMDB-33102: 
Cryo-EM structure of EBV glycoprotein complex gHgL-gp42 bound by a neutralizing antibody 6H2

EMDB-33098: 
Structure of somatostatin receptor 2 bound with SST14.

EMDB-33099: 
Structure of somatostatin receptor 2 bound with octreotide.

EMDB-33100: 
Structure of somatostatin receptor 2 bound with lanreotide.

EMDB-32949: 
Structure of Thyrotropin-Releasing Hormone Receptor bound with Taltirelin.

EMDB-32950: 
Structure of Thyrotropin-Releasing Hormone Receptor bound with an Endogenous Peptide Agonist TRH.
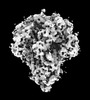
EMDB-32329: 
Cryo-EM map of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32332: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32333: 
Subtomogram averaging of PEDV (Pintung 52) S protein with one protomer in the D0-up conformation and two protomers in the D0-down conformation, determined in situ on intact viral particles

EMDB-32337: 
Subtomogram averaging of PEDV (Pintung 52) S protein with two protomers in the D0-up conformation and one protomer in the D0-down conformation, determined in situ on intact viral particles.

EMDB-32338: 
Cryo-EM map of PEDV S protein with one protomer in the D0-up conformation while the other two in the D0-down conformation
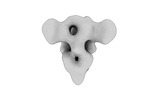
EMDB-32339: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-up conformation determined in situ on intact viral particles.

EMDB-32340: 
Subtomogram averaging of PEDV (Pintung 52) S protein in the postfusion form determined in situ on intact viral particles.

EMDB-33646: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up

EMDB-33647: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein one D0-down and two D0-up

EMDB-33648: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-close conformation
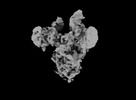
EMDB-33649: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-open conformation

EMDB-33700: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-down
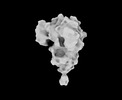
EMDB-33701: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein one D0-up and two D0-down

EMDB-33702: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-up
EMDB-33703: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-down

EMDB-33704: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-up and two D0-down
EMDB-33705: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-down and two D0-up
EMDB-33706: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-up

EMDB-32152: 
Cryo-EM structure of human very long-chain fatty acid ABC transporter ABCD1

EMDB-32171: 
Cryo-EM structure of ATP-bound human very long-chain fatty acid ABC transporter ABCD1

EMDB-32224: 
Cryo-EM structure of C22:0-CoA bound human very long-chain fatty acid ABC transporter ABCD1

EMDB-32526: 
Cryo-EM structure of LY341495/NAM-bound mGlu3

EMDB-32527: 
Cryo-EM structure of inactive mGlu3 bound to LY341495

EMDB-32530: 
Cryo-EM structure of LY2794193-bound mGlu3

EMDB-32557: 
SARS-CoV-2 Omicron S-open
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
