-Search query
-Search result
Showing all 47 items for (author: wells & ja)

EMDB-40589: 
hPAD4 bound to Activating Fab hA362

EMDB-40590: 
hPAD4 bound to inhibitory Fab hI365

PDB-8smk: 
hPAD4 bound to Activating Fab hA362

PDB-8sml: 
hPAD4 bound to inhibitory Fab hI365

EMDB-17958: 
Structure of DPS determined by cryoEM at 100 keV

EMDB-17959: 
Structure of bacterial ribosome determined by cryoEM at 100 keV

EMDB-17960: 
Structure of GABAAR determined by cryoEM at 100 keV

EMDB-17961: 
Structure of mouse heavy-chain apoferritin determined by cryoEM at 100 keV

EMDB-17962: 
Structure of catalase determined by cryoEM at 100 keV

EMDB-17963: 
Structure of AHIR determined by cryoEM at 100 keV

EMDB-17964: 
Structure of GAPDH determined by cryoEM at 100 keV

EMDB-17965: 
Structure of E. coli glutamine synthetase determined by cryoEM at 100 keV

EMDB-17966: 
Structure of human apo ALDH1A1 determined by cryoEM at 100 keV

EMDB-17967: 
Structure of PaaZ determined by cryoEM at 100 keV

EMDB-17968: 
Structure of lumazine synthase determined by cryoEM at 100 keV

PDB-8pv9: 
Structure of DPS determined by cryoEM at 100 keV

PDB-8pva: 
Structure of bacterial ribosome determined by cryoEM at 100 keV

PDB-8pvb: 
Structure of GABAAR determined by cryoEM at 100 keV

PDB-8pvc: 
Structure of mouse heavy-chain apoferritin determined by cryoEM at 100 keV

PDB-8pvd: 
Structure of catalase determined by cryoEM at 100 keV

PDB-8pve: 
Structure of AHIR determined by cryoEM at 100 keV

PDB-8pvf: 
Structure of GAPDH determined by cryoEM at 100 keV

PDB-8pvg: 
Structure of E. coli glutamine synthetase determined by cryoEM at 100 keV

PDB-8pvh: 
Structure of human apo ALDH1A1 determined by cryoEM at 100 keV

PDB-8pvi: 
Structure of PaaZ determined by cryoEM at 100 keV

PDB-8pvj: 
Structure of lumazine synthase determined by cryoEM at 100 keV
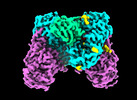
EMDB-40047: 
Structure of human ENPP1 in complex with variable heavy domain VH27.2

PDB-8ghr: 
Structure of human ENPP1 in complex with variable heavy domain VH27.2

EMDB-27730: 
SARS-CoV-2 Wuhan-hu-1-Spike-RBD bound to linker variant of affinity matured ACE2 mimetic CVD432

EMDB-27731: 
SARS-CoV-2 Wuhan-hu-1-Spike-RBD bound to computationally engineered ACE2 mimetic CVD293

PDB-8dv1: 
SARS-CoV-2 Wuhan-hu-1-Spike-RBD bound to linker variant of affinity matured ACE2 mimetic CVD432

PDB-8dv2: 
SARS-CoV-2 Wuhan-hu-1-Spike-RBD bound to computationally engineered ACE2 mimetic CVD293

EMDB-10793: 
Cryo-EM structure of the Full-length disease type human Huntingtin

PDB-6yej: 
Cryo-EM structure of the Full-length disease type human Huntingtin

EMDB-22514: 
SARS CoV2 Spike ectodomain with engineered trimerized VH binder

PDB-7jwb: 
SARS CoV2 Spike ectodomain with engineered trimerized VH binder

EMDB-4937: 
Cryo-EM 3D map of normal Huntingtin

EMDB-4944: 
Cryo-EM 3D map of the N-terminal GFP tagged normal type Huntingtin

PDB-6rmh: 
The Rigid-body refined model of the normal Huntingtin.

EMDB-20175: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch bNAb 10-1074

EMDB-20176: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab874NHP

EMDB-20177: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab897NHP

EMDB-20178: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab275MUR

PDB-6orn: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch bNAb 10-1074

PDB-6oro: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab874NHP

PDB-6orp: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab897NHP

PDB-6orq: 
Modified BG505 SOSIP-based immunogen RC1 in complex with the elicited V3-glycan patch antibody Ab275MUR
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
