-Search query
-Search result
Showing 1 - 50 of 54 items for (author: wendler & p)

EMDB-50019: 
cryoEM structure of Photosystem II averaged across S2-S3 states at 1.71 Angstrom resolution
Method: single particle / : Hussein R, Graca A, Zouni A, Messinger J, Schroder WP

PDB-9evx: 
cryoEM structure of Photosystem II averaged across S2-S3 states at 1.71 Angstrom resolution
Method: single particle / : Hussein R, Graca A, Zouni A, Messinger J, Schroder WP

EMDB-10557: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Miehlke T, Lisdat F, Zouni A, Wendler P

EMDB-10558: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Mielke T, Lisdat F, Zouni A, Wendler P

EMDB-10559: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Mielke T, Lisdat F, Zouni A, Wendler P

PDB-6tra: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Miehlke T, Lisdat F, Zouni A, Wendler P

PDB-6trc: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Mielke T, Lisdat F, Zouni A, Wendler P

PDB-6trd: 
Cryo- EM structure of the Thermosynechococcus elongatus photosystem I in the presence of cytochrome c6
Method: single particle / : Koelsch A, Radon C, Baumert A, Buerger J, Mielke T, Lisdat F, Zouni A, Wendler P

EMDB-10495: 
Cryo-EM Structure of NADH reduced form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C

EMDB-10496: 
Cryo-EM Structure of as isolated form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C

PDB-6tg9: 
Cryo-EM Structure of NADH reduced form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C, Mittelstaedt G

PDB-6tga: 
Cryo-EM Structure of as isolated form of NAD+-dependent Formate Dehydrogenase from Rhodobacter capsulatus
Method: single particle / : Wendler P, Radon C, Mittelstaedt G

EMDB-3699: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP
Method: single particle / : Bracher A, Milicic G, Ciniawsky S, Wendler P, Hayer-Hartl M, Hartl FU

EMDB-3700: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP
Method: single particle / : Bracher A, Milicic G, Ciniawsky S, Wendler P, Hayer-Hartl M, Hart FU

EMDB-3701: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP and RcaCC
Method: single particle / : Bracher A, Milicic G, Ciniawsky S, Wendler P, Hayer-Hartl M, Hartl FU

EMDB-3702: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP and RcaCC
Method: single particle / : Bracher A, Milicic G, Ciniawsky S, Wendler P, Hayer-Hartl M, Hartl FU

PDB-5nv3: 
Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP
Method: single particle / : Bracher A, Milicic G, Ciniawsky S, Wendler P, Hayer-Hartl M, Hartl FU
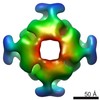
EMDB-3051: 
Structure and mechanism of the Rubisco assembly chaperone Raf1
Method: single particle / : Hauser A, Bhat JY, Milicic G, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M
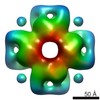
EMDB-3052: 
Structure and mechanism of the Rubisco assembly chaperone Raf1
Method: single particle / : Hauser A, Bhat JY, Milicic G, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

EMDB-3053: 
Structure and mechanism of the Rubisco assembly chaperone Raf1
Method: single particle / : Hauser A, Bhat JY, Milicic G, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

EMDB-2582: 
S. cerevisiae Pex1/6 wild type complex bound to ADP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2583: 
S. cerevisiae Pex1/6 wild type complex bound to gammaS ATP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2584: 
S. cerevisiae Pex1/6 wild type complex bound to ADP-Alfx
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2585: 
S. cerevisiae Pex1/6 wild type complex bound to ATP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2586: 
S. cerevisiae Pex1/6 single Walker B (6WB) complex bound to ATP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2587: 
S. cerevisiae Pex1/6 single Walker B (1WB) complex bound to ATP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2588: 
S. cerevisiae Pex1/6 double Walker B (DWB) complex bound to ATP
Method: single particle / : Ciniawsky S, Grimm I, Saffian D, Girzalsky W, Erdmann R, Wendler P

EMDB-2656: 
The 15S precursor complex of the yeast 20S proteasome
Method: single particle / : Kock M, Nunes MM, Hemann M, Kube S, Dohmen JR, Herzog F, Ramos PC, Wendler P

EMDB-2658: 
20S pre1-1 proteasomal complex from S. cerevisiae carrying the dedicated assembly chaperones Pba1 and Pba2
Method: single particle / : Kock M, Nunes MM, Hemann M, Kube S, Dohmen JR, Herzog F, Ramos PC, Wendler P

EMDB-2524: 
Cryo EM structure of the contractile Type Six Secretion System VipA/B complex
Method: helical / : Kube S, Kapitein N, Zimniak T, Herzog F, Mogk A, Wendler P

EMDB-2525: 
Cryo EM structure of the contractile Type Six Secretion System VipA/B complex
Method: helical / : Kube S, Kapitein N, Zimniak T, Herzog F, Mogk A, Wendler P

PDB-4d2q: 
Negative-stain electron microscopy of E. coli ClpB mutant E432A (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

PDB-4d2u: 
Negative-stain electron microscopy of E. coli ClpB (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

PDB-4d2x: 
Negative-stain electron microscopy of E. coli ClpB of Y503D hyperactive mutant (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2555: 
Negative-stain electron microscopy of E. coli ClpB mutant E432A (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2556: 
Negative-stain electron microscopy of E. coli ClpB mutant E432A (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2557: 
Negative-stain electron microscopy of E. coli ClpB (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2558: 
Negative-stain electron microscopy of E. coli ClpB (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR
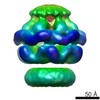
EMDB-2559: 
Negative-stain electron microscopy of E. coli ClpB (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR
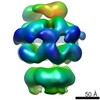
EMDB-2560: 
Negative-stain electron microscopy of E. coli ClpB (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2561: 
Negative-stain electron microscopy of Hsp104 (HAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2562: 
Cryo electron microscopy of E. coli ClpB DWB mutant (BAP form bound to ClpP)
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-2563: 
Cryo electron microscopy of E. coli ClpB
Method: single particle / : Carroni M, Kummer E, Oguchi Y, Clare DK, Wendler P, Sinning I, Kopp J, Mogk A, Bukau B, Saibil HR

EMDB-1940: 
Negative stain EM density of green-type rubisco activase (R294V) from tobacco
Method: single particle / : Stotz M, Mueller-Cajar O, Ciniawsky S, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

PDB-3zw6: 
MODEL OF HEXAMERIC AAA DOMAIN ARRANGEMENT OF GREEN-TYPE RUBISCO ACTIVASE FROM TOBACCO.
Method: single particle / : Stotz M, Mueller-Cajar O, Ciniawsky S, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

PDB-3zuh: 
Negative stain EM Map of the AAA protein CbbX, a red-type Rubisco activase from R. sphaeroides
Method: single particle / : Mueller-Cajar O, Stotz M, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

EMDB-1932: 
Negative stain EM Map of the AAA protein CbbX, a red-type Rubisco activase from R. sphaeroides
Method: single particle / : Mueller-Cajar O, Stotz M, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M

EMDB-1602: 
Motor mechanism for protein threading through Hsp104
Method: single particle / : Wendler P, Shorter J, Snead D, Plisson C, Clare DK, Lindquist S, Saibil H

EMDB-1599: 
Motor mechanism for protein threading through Hsp104
Method: single particle / : Wendler P, Shorter J, Snead D, Plisson C, Clare DK, Lindquist S, Saibil H

EMDB-1600: 
Motor mechanism for protein threading through Hsp104
Method: single particle / : Wendler P, Shorter J, Snead D, Plisson C, Clare DK, Lindquist S, Saibil H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
