-Search query
-Search result
Showing 1 - 50 of 65 items for (author: smith & tj)

EMDB-42600: 
Murine norovirus in the presence of 1mM calcium

EMDB-42604: 
Murine norovirus + 1 mM MgCl2

EMDB-42623: 
Murine norovirus dialyzed against EDTA

EMDB-27847: 
Mouse norovirus strain CR6, attenuated

EMDB-27849: 
Mouse norovirus strain CR6 at pH 5.0

EMDB-15786: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)
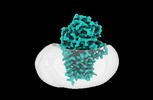

EMDB-15787: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE
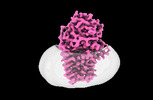
EMDB-15788: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)

EMDB-15789: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE
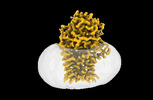
EMDB-15790: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3
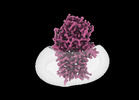
EMDB-15791: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3

PDB-8b0k: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli (Apo form)

PDB-8b0l: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE

PDB-8b0m: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with PE (C387S mutant)

PDB-8b0n: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Lyso-PE

PDB-8b0o: 
Cryo-EM structure apolipoprotein N-acyltransferase Lnt from E.coli in complex with FP3

PDB-8b0p: 
Cryo-EM structure of apolipoprotein N-acyltransferase Lnt from E. coli in complex with Pam3

EMDB-27319: 
Mouse Norovirus strain WU23

EMDB-27321: 
Mouse norovirus strain WU23 + 10mM GCDCA

EMDB-27322: 
Murine norovirus strain WU23 at pH 5

EMDB-26862: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD

PDB-7uxu: 
CryoEM structure of the TIR domain from AbTir in complex with 3AD

EMDB-12590: 
Bacteriophage YerA41 head icosahedral reconstruction

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
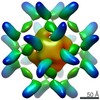
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit

EMDB-24211: 
Mouse norovirus (MNV-1) capsid at pH 5.0

EMDB-24226: 
Mouse norovirus (MNV-1) capsid at pH 7.5

EMDB-24533: 
SARS-CoV-2 spike protein bound to the S2P6 and S2M11 Fab fragments

EMDB-23187: 
Mouse Norovirus Protruding domain complexed with neutralizing Fab fragment from mAb A6.2

EMDB-22491: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody Fab fragment (local refinement of the receptor-binding motif and Fab variable domains)

EMDB-22492: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (one RBD open)

EMDB-22494: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (closed conformation)

EMDB-22497: 
SARS-CoV-2 spike in complex with the S2A4 neutralizing antibody Fab fragment (local refinement of the receptor-binding domain and Fab variable domains)

EMDB-22506: 
SARS-CoV-2 spike in complex with the S2A4 neutralizing antibody Fab fragment

EMDB-22507: 
SARS-CoV-2 spike in complex with the S2H14 neutralizing antibody Fab fragment (two receptor-binding domains open)

EMDB-22508: 
SARS-CoV-2 spike in complex with the S2H14 neutralizing antibody Fab fragment (three receptor-binding domains open)

EMDB-22512: 
SARS-CoV-2 spike in complex with the S304 neutralizing antibody Fab fragment

EMDB-22516: 
SARS-CoV-2 spike in complex with the S2X35 neutralizing antibody Fab fragment

EMDB-22517: 
SARS-CoV-2 spike in complex with the S2X35 neutralizing antibody Fab fragment (local refinement of the receptor-binding motif and Fab variable domains)

PDB-7jv2: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody Fab fragment (local refinement of the receptor-binding motif and Fab variable domains)

PDB-7jv4: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (one RBD open)

PDB-7jv6: 
SARS-CoV-2 spike in complex with the S2H13 neutralizing antibody (closed conformation)

PDB-7jva: 
SARS-CoV-2 spike in complex with the S2A4 neutralizing antibody Fab fragment (local refinement of the receptor-binding domain and Fab variable domains)

PDB-7jvc: 
SARS-CoV-2 spike in complex with the S2A4 neutralizing antibody Fab fragment

PDB-7jw0: 
SARS-CoV-2 spike in complex with the S304 neutralizing antibody Fab fragment

EMDB-11041: 
The structure of the dimeric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

EMDB-11042: 
The structure of the tetrameric HDAC1/MIDEAS/DNTTIP1 MiDAC deacetylase complex

EMDB-9204: 
Cucumber Leaf Spot Virus
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
