-検索条件
-検索結果
検索 (著者・登録者: guenther & e)の結果全35件を表示しています

EMDB-13867: 
Structure of the bacterial type VI secretion system effector RhsA.

PDB-7q97: 
Structure of the bacterial type VI secretion system effector RhsA.

EMDB-13843: 
Structure of VgrG1 from Pseudomonas protegens.

PDB-7q5p: 
Structure of VgrG1 from Pseudomonas protegens.

PDB-6y6k: 
Cryo-EM structure of a Phenuiviridae L protein

EMDB-9204: 
Cucumber Leaf Spot Virus

EMDB-9205: 
Red Clover Necrotic Mosaic Virus

PDB-6mrl: 
Cucumber Leaf Spot Virus

PDB-6mrm: 
Red Clover Necrotic Mosaic Virus

EMDB-0535: 
Cryo-EM Reconstruction of Protease-Activateable Adeno-Associated Virus 9 (AAV9-L001)

PDB-6nxe: 
Cryo-EM Reconstruction of Protease-Activateable Adeno-Associated Virus 9 (AAV9-L001)

PDB-6hij: 
Cryo-EM structure of the human ABCG2-MZ29-Fab complex with cholesterol and PE lipids docked

EMDB-7466: 
Segment NFGTFS, with familial mutation A315T and phosphorylated threonine, from the low complexity domain of TDP-43, residues 312-317

EMDB-7467: 
SWGMMGMLASQ segment from the low complexity domain of TDP-43, residues 333-343

EMDB-8857: 
MicroED structure of the segment, NFGEFS, from the A315E familial variant of the low complexity domain of TDP-43, residues 312-317

PDB-5wkb: 
MicroED structure of the segment, NFGEFS, from the A315E familial variant of the low complexity domain of TDP-43, residues 312-317

PDB-6cf4: 
Segment NFGTFS, with familial mutation A315T and phosphorylated threonine, from the low complexity domain of TDP-43, residues 312-317

PDB-6cfh: 
SWGMMGMLASQ segment from the low complexity domain of TDP-43

EMDB-3953: 
Structure of inhibitor-bound ABCG2

EMDB-4246: 
Structure of inhibitor-bound ABCG2

EMDB-4256: 
Structure of an inhibitor-bound ABC transporter

PDB-6eti: 
Structure of inhibitor-bound ABCG2

PDB-6feq: 
Structure of inhibitor-bound ABCG2

PDB-6ffc: 
Structure of an inhibitor-bound ABC transporter

EMDB-8781: 
CryoEM structure of the segment, DLIIKGISVHI, assembled into a triple-helical fibril

PDB-5w7v: 
CryoEM structure of the segment, DLIIKGISVHI, assembled into a triple-helical fibril

EMDB-8765: 
MicroED structure of the segment, DLIIKGISVHI, from the RRM2 of TDP-43, residues 247-257

PDB-5w52: 
MicroED structure of the segment, DLIIKGISVHI, from the RRM2 of TDP-43, residues 247-257

EMDB-3001: 
MicroED structure of the segment, GVVHGVTTVA, from the A53T familial mutant of Parkinson's disease protein, alpha-synuclein, residues 47-56
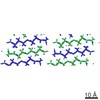
EMDB-3028: 
MicroED structure of the toxic core segment, GAVVTGVTAVA, from Parkinson's disease protein, alpha-synuclein, residues 69-78.

PDB-4znn: 
MicroED structure of the segment, GVVHGVTTVA, from the A53T familial mutant of Parkinson's disease protein, alpha-synuclein residues 47-56

PDB-4ril: 
Structure of the amyloid forming segment, GAVVTGVTAVA, from the NAC domain of Parkinson's disease protein alpha-synuclein, residues 68-78, determined by electron diffraction

EMDB-5369: 
Structural transitions in RCNMV revealing potential mechanism of RNA release (native map)

EMDB-5372: 
Structural transitions in RCNMV revealing potential mechanism of RNA release (EGTA map)

EMDB-5373: 
Structural transitions in RCNMV revealing potential mechanism of RNA release (EDTA map)
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
