-Search query
-Search result
Showing 1 - 50 of 425 items for (author: chung & i)
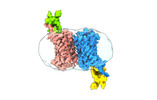
EMDB-36488: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-37212: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Receptor original map)
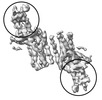
EMDB-37214: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Ligand/CCL7 focused map)

PDB-8jps: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-39126: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly

EMDB-39127: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly

PDB-8ybx: 
Structure of the FADD/Caspase-8/cFLIP death effector domain assembly

EMDB-41837: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K

PDB-8u1x: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K

EMDB-42018: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K

PDB-8u89: 
The structure of the PP2A-B56Delta holoenzyme mutant - E197K
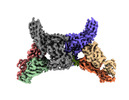
EMDB-34174: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)
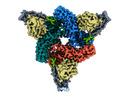
EMDB-36078: 
Structure of beta-arrestin2 in complex with M2Rpp
EMDB-36081: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)
EMDB-36082: 
Structure of beta-arrestin1 in complex with D6Rpp
EMDB-36090: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)
EMDB-36091: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, Cross-linked)
EMDB-36093: 
Muscarinic receptor (M2R) in complex with beta-arrestin1 (Low resolution full map, non-crosslinked)

EMDB-36110: 
Structure of basal beta-arrestin2
EMDB-36124: 
Structure of beta-arrestin1 in complex with C3aRpp
EMDB-36126: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)

PDB-8go9: 
Structure of beta-arrestin2 in complex with a phosphopeptide corresponding to the human Atypical chemokine receptor 2, ACKR2 (D6R)

PDB-8j8r: 
Structure of beta-arrestin2 in complex with M2Rpp

PDB-8j8v: 
Structure of beta-arrestin2 in complex with D6Rpp (Local Refine)

PDB-8j8z: 
Structure of beta-arrestin1 in complex with D6Rpp

PDB-8j97: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local refine, cross-linked)

PDB-8j9k: 
Structure of basal beta-arrestin2

PDB-8ja3: 
Structure of beta-arrestin1 in complex with C3aRpp

PDB-8jaf: 
Structure of Muscarinic receptor (M2R) in complex with beta-arrestin1 (Local Refine, non-cross linked)
EMDB-40985: 
Structure of a group II intron ribonucleoprotein in the pre-ligation (pre-2F) state
EMDB-40986: 
Structure of a group II intron ribonucleoprotein in the pre-branching (pre-1F) state
EMDB-40987: 
Structure of a group II intron ribonucleoprotein in the post-ligation (post-2F) state

PDB-8t2r: 
Structure of a group II intron ribonucleoprotein in the pre-ligation (pre-2F) state

PDB-8t2s: 
Structure of a group II intron ribonucleoprotein in the pre-branching (pre-1F) state

PDB-8t2t: 
Structure of a group II intron ribonucleoprotein in the post-ligation (post-2F) state

EMDB-29013: 
96nm doublet microtubule repeat from wild type mouse sperm

EMDB-28606: 
96nm doublet microtubule repeat from LRRC23-KO mouse sperm

EMDB-34000: 
Open-spiral pentamer of the substrate-free Lon protease with a Y224S mutation

EMDB-34001: 
Spiral hexamer of the substrate-free Lon protease with a Y224S mutation

EMDB-34002: 
Spiral pentamer of the substrate-free Lon protease with a S678A mutation

EMDB-34003: 
Close-ring hexamer of the substrate-bound Lon protease with an S678A mutation

EMDB-34004: 
Spiral hexamer of the substrate-free Lon protease with an S678A mutation

EMDB-34005: 
Open-spiral pentamer of the substrate-free Lon protease with Y397A and S678A mutations

EMDB-34006: 
Spiral hexamer of the substrate-free Lon protease with Y397A and S678A mutations

EMDB-34107: 
MtaLon-Apo for the spiral oligomers of trimer
EMDB-34108: 
MtaLon-Apo for the spiral oligomers of tetramer

EMDB-34109: 
MtaLon-Apo for the spiral oligomers of pentamer

EMDB-34110: 
MtaLon-Apo for the spiral oligomers of hexamer
EMDB-34111: 
MtaLon-ADP for the spiral oligomers of trimer
EMDB-34112: 
MtaLon-ADP for the spiral oligomers of tetramer
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
