[English] 日本語
 Yorodumi
Yorodumi- EMDB-39126: Structure of the FADD/Caspase-8/cFLIP death effector domain assembly -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the FADD/Caspase-8/cFLIP death effector domain assembly | |||||||||
 Map data Map data | sharpened map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | FADD / caspase-8 / cellular FLICE-like inhibitory protein / Death effector domain / APOPTOSIS | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of CD8-positive, alpha-beta cytotoxic T cell extravasation / negative regulation of activation-induced cell death of T cells / negative regulation of myoblast fusion / skeletal muscle atrophy / skeletal myofibril assembly / caspase-8 / syncytiotrophoblast cell differentiation involved in labyrinthine layer development / death effector domain binding / tumor necrosis factor receptor superfamily binding / FasL/ CD95L signaling ...positive regulation of CD8-positive, alpha-beta cytotoxic T cell extravasation / negative regulation of activation-induced cell death of T cells / negative regulation of myoblast fusion / skeletal muscle atrophy / skeletal myofibril assembly / caspase-8 / syncytiotrophoblast cell differentiation involved in labyrinthine layer development / death effector domain binding / tumor necrosis factor receptor superfamily binding / FasL/ CD95L signaling / TRAIL signaling / regulation of skeletal muscle satellite cell proliferation / CD95 death-inducing signaling complex / Apoptotic execution phase / death-inducing signaling complex assembly / Activation, myristolyation of BID and translocation to mitochondria / Defective RIPK1-mediated regulated necrosis / TRAIL-activated apoptotic signaling pathway / Microbial modulation of RIPK1-mediated regulated necrosis / ripoptosome / caspase binding / regulation of necroptotic process / TRIF-mediated programmed cell death / Regulation by c-FLIP / CASP8 activity is inhibited / Dimerization of procaspase-8 / positive regulation of extracellular matrix organization / TLR3-mediated TICAM1-dependent programmed cell death / self proteolysis / Caspase activation via Death Receptors in the presence of ligand / positive regulation of macrophage differentiation / positive regulation of adaptive immune response / negative regulation of hepatocyte apoptotic process / response to cobalt ion / positive regulation of glomerular mesangial cell proliferation / necroptotic signaling pathway / NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 / positive regulation of hepatocyte proliferation / skeletal muscle tissue regeneration / CLEC7A/inflammasome pathway / death-inducing signaling complex / negative regulation of necroptotic process / receptor serine/threonine kinase binding / regulation of tumor necrosis factor-mediated signaling pathway / positive regulation of type I interferon-mediated signaling pathway / tumor necrosis factor receptor binding / positive regulation of innate immune response / death receptor binding / natural killer cell activation / motor neuron apoptotic process / positive regulation of activated T cell proliferation / TNFR1-induced proapoptotic signaling / negative regulation of cellular response to transforming growth factor beta stimulus / positive regulation of extrinsic apoptotic signaling pathway / execution phase of apoptosis / RIPK1-mediated regulated necrosis / response to anesthetic / Apoptotic cleavage of cellular proteins / regulation of innate immune response / response to testosterone / T cell homeostasis / positive regulation of proteolysis / macrophage differentiation / B cell activation / negative regulation of cardiac muscle cell apoptotic process / response to tumor necrosis factor / extrinsic apoptotic signaling pathway via death domain receptors / pyroptotic inflammatory response / cellular response to dexamethasone stimulus / Caspase-mediated cleavage of cytoskeletal proteins / lymph node development / positive regulation of execution phase of apoptosis / negative regulation of extrinsic apoptotic signaling pathway via death domain receptors / skeletal muscle tissue development / spleen development / negative regulation of reactive oxygen species biosynthetic process / extrinsic apoptotic signaling pathway / extrinsic apoptotic signaling pathway in absence of ligand / behavioral response to cocaine / cellular response to nitric oxide / cysteine-type peptidase activity / signaling adaptor activity / regulation of cytokine production / : / cellular response to epidermal growth factor stimulus / thymus development / T cell activation / negative regulation of extrinsic apoptotic signaling pathway / positive regulation of interleukin-1 beta production / positive regulation of interleukin-8 production / negative regulation of canonical NF-kappaB signal transduction / Regulation of NF-kappa B signaling / cellular response to estradiol stimulus / kidney development / apoptotic signaling pathway / Regulation of TNFR1 signaling / cellular response to mechanical stimulus / protein maturation / NOD1/2 Signaling Pathway / protein processing Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
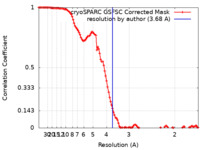
| Method | single particle reconstruction / cryo EM / Resolution: 3.68 Å | |||||||||
 Authors Authors | Lin S-C / Yang C-Y | |||||||||
| Funding support |  Taiwan, 1 items Taiwan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Deciphering DED assembly mechanisms in FADD-procaspase-8-cFLIP complexes regulating apoptosis. Authors: Chao-Yu Yang / Chia-I Lien / Yi-Chun Tseng / Yi-Fan Tu / Arkadiusz W Kulczyk / Yen-Chen Lu / Yin-Ting Wang / Tsung-Wei Su / Li-Chung Hsu / Yu-Chih Lo / Su-Chang Lin /   Abstract: Fas-associated protein with death domain (FADD), procaspase-8, and cellular FLICE-inhibitory proteins (cFLIP) assemble through death-effector domains (DEDs), directing death receptor signaling ...Fas-associated protein with death domain (FADD), procaspase-8, and cellular FLICE-inhibitory proteins (cFLIP) assemble through death-effector domains (DEDs), directing death receptor signaling towards cell survival or apoptosis. Understanding their three-dimensional regulatory mechanism has been limited by the absence of atomic coordinates for their ternary DED complex. By employing X-ray crystallography and cryogenic electron microscopy (cryo-EM), we present the atomic coordinates of human FADD-procaspase-8-cFLIP complexes, revealing structural insights into these critical interactions. These structures illustrate how FADD and cFLIP orchestrate the assembly of caspase-8-containing complexes and offer mechanistic explanations for their role in promoting or inhibiting apoptotic and necroptotic signaling. A helical procaspase-8-cFLIP hetero-double layer in the complex appears to promote limited caspase-8 activation for cell survival. Our structure-guided mutagenesis supports the role of the triple-FADD complex in caspase-8 activation and in regulating receptor-interacting protein kinase 1 (RIPK1). These results propose a unified mechanism for DED assembly and procaspase-8 activation in the regulation of apoptotic and necroptotic signaling across various cellular pathways involved in development, innate immunity, and disease. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39126.map.gz emd_39126.map.gz | 70.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39126-v30.xml emd-39126-v30.xml emd-39126.xml emd-39126.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39126_fsc.xml emd_39126_fsc.xml | 11 KB | Display |  FSC data file FSC data file |
| Images |  emd_39126.png emd_39126.png | 37.3 KB | ||
| Filedesc metadata |  emd-39126.cif.gz emd-39126.cif.gz | 6.4 KB | ||
| Others |  emd_39126_half_map_1.map.gz emd_39126_half_map_1.map.gz emd_39126_half_map_2.map.gz emd_39126_half_map_2.map.gz | 127.1 MB 127.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39126 http://ftp.pdbj.org/pub/emdb/structures/EMD-39126 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39126 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39126 | HTTPS FTP |
-Related structure data
| Related structure data |  8ybxMC  8yd7C  8yd8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_39126.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39126.map.gz / Format: CCP4 / Size: 137.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.84 Å | ||||||||||||||||||||||||||||||||||||
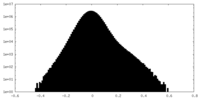
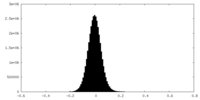
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_39126_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
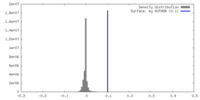
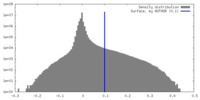
| Density Histograms |
-Half map: #1
| File | emd_39126_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : FADD/Caspase-8/cFLIP death effector domain assembly
| Entire | Name: FADD/Caspase-8/cFLIP death effector domain assembly |
|---|---|
| Components |
|
-Supramolecule #1: FADD/Caspase-8/cFLIP death effector domain assembly
| Supramolecule | Name: FADD/Caspase-8/cFLIP death effector domain assembly / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Caspase-8 subunit p10
| Macromolecule | Name: Caspase-8 subunit p10 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.191648 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDFSRNLYDI GEQLDSEDLA SLKFLSLDYI PQRKQEPIKD ALMLFQRLQE KRMLEESNLS FLKELLFRIN RLDLLITYLN TRKEEMERE LQTPGRAQIS AYRVMLYQIS EEVSRSELRS FKGGLQEEIS KCKLDDDMNL LDIFIEMEKR VILGEGKLDI L KRVCAQIN ...String: MDFSRNLYDI GEQLDSEDLA SLKFLSLDYI PQRKQEPIKD ALMLFQRLQE KRMLEESNLS FLKELLFRIN RLDLLITYLN TRKEEMERE LQTPGRAQIS AYRVMLYQIS EEVSRSELRS FKGGLQEEIS KCKLDDDMNL LDIFIEMEKR VILGEGKLDI L KRVCAQIN KSLLKIINDY EEFSKERSSS LEGSPDEFSN GEELCGVMTI SDSPREQDSE SQTLDKVYQM KSKPRGYCLI IN NHNFAKA REKVPKLHSI RDRNGTHLDA GALTTTFEEL HFEIKPHDDC TVEQIYEILK IYQLMDHSNM DCFICCILSH GDK GIIYGT DGQEAPIYEL TSQFTGLKCP SLAGKPKVFF IQAAQGDNYQ KGIPVETASE EQPYLEMALS SPQTRYIPDE ADFL LGMAT VNNCVSYRNP AEGTWYIQSL CQSLRERCPR GDDILTILTE VNYEVSNKDD KKNMGKQMPQ PTFTLRKKLV FPSD UniProtKB: Caspase-8 |
-Macromolecule #2: CASP8 and FADD-like apoptosis regulator subunit p43
| Macromolecule | Name: CASP8 and FADD-like apoptosis regulator subunit p43 / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 20.878479 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSAEVIHQVE EALDTDEKEM LLFLCRDVAI DVVPPNVRDL LDILRERGKL SVGDLAELLY RVRRFDLLKR ILKMDRKAVE THLLRNPHL VSDYRVLMAE IGEDLDKSDV SSLIFLMKDY MGRGKISKEK SFLDLVVELE KLNLVAPDQL DLLEKCLKNI H RIDLKTKI QKYKQSVQGA GTS UniProtKB: CASP8 and FADD-like apoptosis regulator |
-Macromolecule #3: FAS-associated death domain protein
| Macromolecule | Name: FAS-associated death domain protein / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.381533 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDPFLVLLHS VSSSLSSSEL TELKFLCLGR VGKRKLERVQ SGLDLFSMLL EQNDLEPGHT ELLRELLASL RRHDLLRRVD DFEAGAAAG AAPGEEDLCA AFNVICDNVG KDWRRLARQL KVSDTKIDSI EDRYPRNLTE RVRESLRIWK NTEKENATVA H LVGALRSC ...String: MDPFLVLLHS VSSSLSSSEL TELKFLCLGR VGKRKLERVQ SGLDLFSMLL EQNDLEPGHT ELLRELLASL RRHDLLRRVD DFEAGAAAG AAPGEEDLCA AFNVICDNVG KDWRRLARQL KVSDTKIDSI EDRYPRNLTE RVRESLRIWK NTEKENATVA H LVGALRSC QMNLVADLVQ EVQQARDLQN RSGAMSPMSW NSDASTSEAS LEHHHHHH UniProtKB: FAS-associated death domain protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 72.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 165000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)