+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 96nm doublet microtubule repeat from wild type mouse sperm | |||||||||
 Map data Map data | 96nm doublet microtubule repeat from WT mouse sperm | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | 96 nm doublet microtubule from mouse sperm / STRUCTURAL PROTEIN | |||||||||
| Biological species |  | |||||||||
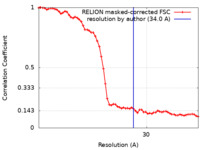
| Method | subtomogram averaging / cryo EM / Resolution: 34.0 Å | |||||||||
 Authors Authors | Hwang JY / Chai P / Nawaz S | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2023 Journal: Elife / Year: 2023Title: LRRC23 truncation impairs radial spoke 3 head assembly and sperm motility underlying male infertility. Authors: Jae Yeon Hwang / Pengxin Chai / Shoaib Nawaz / Jungmin Choi / Francesc Lopez-Giraldez / Shabir Hussain / Kaya Bilguvar / Shrikant Mane / Richard P Lifton / Wasim Ahmad / Kai Zhang / Jean-Ju Chung /    Abstract: Radial spokes (RS) are T-shaped multiprotein complexes on the axonemal microtubules. Repeated RS1, RS2, and RS3 couple the central pair to modulate ciliary and flagellar motility. Despite the cell ...Radial spokes (RS) are T-shaped multiprotein complexes on the axonemal microtubules. Repeated RS1, RS2, and RS3 couple the central pair to modulate ciliary and flagellar motility. Despite the cell type specificity of RS3 substructures, their molecular components remain largely unknown. Here, we report that a leucine-rich repeat-containing protein, LRRC23, is an RS3 head component essential for its head assembly and flagellar motility in mammalian spermatozoa. From infertile male patients with defective sperm motility, we identified a splice site variant of . A mutant mouse model mimicking this variant produces a truncated LRRC23 at the C-terminus that fails to localize to the sperm tail, causing male infertility due to defective sperm motility. LRRC23 was previously proposed to be an ortholog of the RS stalk protein RSP15. However, we found that purified recombinant LRRC23 interacts with an RS head protein RSPH9, which is abolished by the C-terminal truncation. Evolutionary and structural comparison also shows that LRRC34, not LRRC23, is the RSP15 ortholog. Cryo-electron tomography clearly revealed that the absence of the RS3 head and the sperm-specific RS2-RS3 bridge structure in LRRC23 mutant spermatozoa. Our study provides new insights into the structure and function of RS3 in mammalian spermatozoa and the molecular pathogenicity of LRRC23 underlying reduced sperm motility in infertile human males. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_29013.map.gz emd_29013.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-29013-v30.xml emd-29013-v30.xml emd-29013.xml emd-29013.xml | 16.6 KB 16.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_29013_fsc.xml emd_29013_fsc.xml | 5.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_29013.png emd_29013.png | 58.8 KB | ||
| Filedesc metadata |  emd-29013.cif.gz emd-29013.cif.gz | 4.3 KB | ||
| Others |  emd_29013_additional_1.map.gz emd_29013_additional_1.map.gz emd_29013_half_map_1.map.gz emd_29013_half_map_1.map.gz emd_29013_half_map_2.map.gz emd_29013_half_map_2.map.gz | 14.7 MB 7.7 MB 7.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-29013 http://ftp.pdbj.org/pub/emdb/structures/EMD-29013 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29013 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-29013 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_29013.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_29013.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 96nm doublet microtubule repeat from WT mouse sperm | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 10.2 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: unmasked 96nm doublet microtubule repeat from WT mouse sperm
| File | emd_29013_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unmasked 96nm doublet microtubule repeat from WT mouse sperm | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2 of 96nm doublet microtubule repeat from WT mouse sperm
| File | emd_29013_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 of 96nm doublet microtubule repeat from WT mouse sperm | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1 of 96nm doublet microtubule repeat from WT mouse sperm
| File | emd_29013_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 of 96nm doublet microtubule repeat from WT mouse sperm | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Mouse sperm
| Entire | Name: Mouse sperm |
|---|---|
| Components |
|
-Supramolecule #1: Mouse sperm
| Supramolecule | Name: Mouse sperm / type: cell / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 5.5 |
|---|---|
| Grid | Model: Quantifoil R2/2 / Material: GOLD / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 30 % / Chamber temperature: 298 K / Instrument: HOMEMADE PLUNGER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Phase plate: VOLTA PHASE PLATE |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 0.5 µm / Nominal defocus min: 0.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)