-検索条件
-検索結果
検索 (著者・登録者: chipman & pr)の結果52件中、1から50件目までを表示しています

EMDB-6168: 
Structure of Enteric Pathogen Bovine Parvovirus

EMDB-5296: 
cryo-EM reconstruction of West Nile virus

EMDB-5378: 
The structure of PBCV-1 (icosahedrally averaged map)

EMDB-5384: 
The structure of PBCV-1 (five-fold averaged map)

EMDB-5385: 
The structure of PBCV-1 (local symmetry averaged map of major capsid protein)
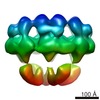
EMDB-2056: 
Electron cryo-microscopy of the Hantaan virus glycoprotein spike

EMDB-5190: 
West Nile Virus in complex with Fab fragments of the neutralizing monoclonal antibody CR4354

EMDB-5179: 
The Interaction of Decay-accelerating Factor with Echovirus 7

EMDB-5115: 
West Nile virus in complex with a single-chain antibody derivative of the neutralizing monoclonal antibody E16

EMDB-5117: 
Structure of furin-cleaved immature dengue virus at low pH

EMDB-5106: 
Fab from MAb B interacting with feline panleukopenia virus

EMDB-5107: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 6

EMDB-5108: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 8

EMDB-5109: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 15

EMDB-5110: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb 16

EMDB-5111: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb E

EMDB-5112: 
Feline panleukopenia virus in complex with FAb from neutralizing antibody MAb F

EMDB-5102: 
Immature Dengue Virus (DENV) in complex with Fab fragments of the anti-fusion loop antibody E53

EMDB-5103: 
Immature West Nile Virus (WNV) in complex with Fab fragments of the anti-fusion loop antibody E53

EMDB-1597: 
5-fold averaged density map of Paramecium bursaria Chlorella virus-1 (PBCV-1).
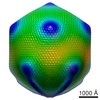
EMDB-5039: 
Cryo-EM reconstruction of the giant Mimivirus using C5 symmetry

EMDB-5006: 
Structure of immature Dengue virus at low pH

EMDB-5105: 
Canine parvovirus in complex with FAb from neutralizing antibody MAb 14

EMDB-1580: 
CryoEM and 3D image reconstruction of Chilo Iridescent virus at 13 angstrom resolution

EMDB-1466: 
Human Parvovirus B19 (DNA-containing wildtype)

EMDB-1467: 
Human Parvovirus B19 (empty wildtype particle)

EMDB-1468: 
Virus-like particle of Human Parvovirus B19 (VLP consisting of VP2 only)

EMDB-1418: 
Binding of a neutralizing antibody to dengue virus alters the arrangement of surface glycoproteins

EMDB-1326: 
Minute virus of mice, a parvovirus, in complex with the Fab fragment of a neutralizing monoclonal antibody.

EMDB-1280: 
Cryo-electron microscopy study of bacteriophage T4 displaying anthrax toxin proteins.

EMDB-1393: 
Cryo-EM study of the Pseudomonas bacteriophage phiKZ.

EMDB-1392: 
A three-dimensional cryo-electron microscopy structure of the bacteriophage phiKZ head.

EMDB-1415: 
Cryo-EM study of the Pseudomonas bacteriophage phiKZ.

EMDB-1414: 
Molecular architecture of the prolate head of bacteriophage T4.

EMDB-1411: 
Interaction of decay-accelerating factor with coxsackievirus B3.

EMDB-1412: 
Interaction of decay-accelerating factor with coxsackievirus B3.

EMDB-1314: 
Structure of immature West Nile virus.

EMDB-1287: 
Asymmetric binding of transferrin receptor to parvovirus capsids.

EMDB-1288: 
Asymmetric binding of transferrin receptor to parvovirus capsids.

EMDB-1234: 
West Nile virus in complex with the Fab fragment of a neutralizing monoclonal antibody.

EMDB-1166: 
Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN.
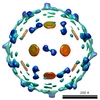
EMDB-1167: 
Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN.

EMDB-1121: 
Mapping the structure and function of the E1 and E2 glycoproteins in alphaviruses.

EMDB-1126: 
The tail structure of bacteriophage T4 and its mechanism of contraction.

EMDB-1114: 
The crystal structure of coxsackievirus A21 and its interaction with ICAM-1.

EMDB-1116: 
Conservation of the capsid structure in tailed dsDNA bacteriophages: the pseudoatomic structure of phi29.

EMDB-1117: 
Conservation of the capsid structure in tailed dsDNA bacteriophages: the pseudoatomic structure of phi29.

EMDB-1120: 
Conservation of the capsid structure in tailed dsDNA bacteriophages: the pseudoatomic structure of phi29.

EMDB-1086: 
Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host.

EMDB-1089: 
Three-dimensional rearrangement of proteins in the tail of bacteriophage T4 on infection of its host.
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
