-検索条件
-検索結果
検索 (著者・登録者: blum & t)の結果72件中、1から50件目までを表示しています
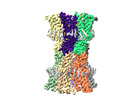
EMDB-18540: 
human connexin-36 gap junction channel in complex with mefloquine
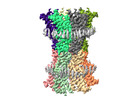
EMDB-18987: 
human connexin-36 gap junction channel
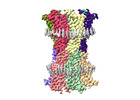
EMDB-18988: 
human connexin-36 gap junction channel in complex with quinine

PDB-8qoj: 
human connexin-36 gap junction channel in complex with mefloquine

PDB-8r7p: 
human connexin-36 gap junction channel

PDB-8r7q: 
human connexin-36 gap junction channel in complex with quinine

EMDB-19163: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

EMDB-19164: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

EMDB-19165: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

EMDB-19166: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

PDB-8rgz: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

PDB-8rh0: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

PDB-8rh1: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.

PDB-8rh2: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.

EMDB-17208: 
CRYO-EM STRUCTURE OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : PARENTAL STRAIN

EMDB-17212: 
CRYO-EM STRUCTURE OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : TB11CS6H1 snoRNA mutant

EMDB-17249: 
CRYO-EM CONSENSUS MAP OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : PARENTAL STRAIN
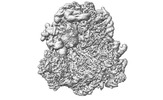
EMDB-17250: 
CRYO-EM FOCUSED REFINEMENT MAPS OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : PARENTAL STRAIN

EMDB-17254: 
CRYO-EM CONSENSUS MAP OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : TB11CS6H1 SKO
EMDB-17255: 
CRYO-EM FOCUSED REFINEMENT OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : TB11CS6H1 snoRNA mutant

PDB-8ova: 
CRYO-EM STRUCTURE OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : PARENTAL STRAIN

PDB-8ove: 
CRYO-EM STRUCTURE OF TRYPANOSOMA BRUCEI PROCYCLIC FORM 80S RIBOSOME : TB11CS6H1 snoRNA mutant

EMDB-28959: 
DNA replication fork binding triggers structural changes in the PriA DNA helicase that regulate the PriA-PriB replication restart pathway in E. coli

PDB-8fak: 
DNA replication fork binding triggers structural changes in the PriA DNA helicase that regulate the PriA-PriB replication restart pathway in E. coli

EMDB-14634: 
Cryo-EM structure of GMPCPP-microtubules in complex with VASH2-SVBP

PDB-7zcw: 
Cryo-EM structure of GMPCPP-microtubules in complex with VASH2-SVBP

EMDB-13642: 
Alpha-latrocrustotoxin monomer

EMDB-13643: 
Delta-latroinsectotoxin dimer

PDB-7ptx: 
Alpha-latrocrustotoxin monomer

PDB-7pty: 
Delta-latroinsectotoxin dimer

EMDB-24318: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C032

EMDB-24319: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C051

EMDB-24320: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C548

PDB-7r8m: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C032

PDB-7r8n: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C051

PDB-7r8o: 
Structure of the SARS-CoV-2 S 6P trimer in complex with neutralizing antibody C548

EMDB-24404: 
Covalent inhibition of hAChE by organophosphates causes homodimer dissociation through long-range allosteric effects

EMDB-23693: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG10-19

EMDB-23694: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG1-22

EMDB-23695: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG7-15

EMDB-23696: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG7-20

EMDB-23697: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG1-24

PDB-7m6e: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG10-19

PDB-7m6f: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG1-22

PDB-7m6g: 
Structure of the SARS-CoV-2 S 6P trimer in complex with the human neutralizing antibody Fab fragment, BG7-15

PDB-7m6h: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG7-20

PDB-7m6i: 
Structure of the SARS-CoV-2 S 2P trimer in complex with the human neutralizing antibody Fab fragment, BG1-24

EMDB-12696: 
Amyloid beta oligomer displayed on the alpha hemolysin scaffold

PDB-7o1q: 
Amyloid beta oligomer displayed on the alpha hemolysin scaffold

EMDB-11022: 
Allostery through DNA drives phenotype switching
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
