4IBQ
 
 | | Human p53 core domain with hot spot mutation R273C | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Cellular tumor antigen p53, ... | | Authors: | Rozenberg, H, Eldar, A, Diskin-Posner, Y, Shakked, Z. | | Deposit date: | 2012-12-09 | | Release date: | 2013-08-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural studies of p53 inactivation by DNA-contact mutations and its rescue by suppressor mutations via alternative protein-DNA interactions.
Nucleic Acids Res., 41, 2013
|
|
4IBS
 
 | | Human p53 core domain with hot spot mutation R273H (form I) | | Descriptor: | 1,2-ETHANEDIOL, Cellular tumor antigen p53, ZINC ION | | Authors: | Rozenberg, H, Eldar, A, Diskin-Posner, Y, Shakked, Z. | | Deposit date: | 2012-12-09 | | Release date: | 2013-08-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural studies of p53 inactivation by DNA-contact mutations and its rescue by suppressor mutations via alternative protein-DNA interactions.
Nucleic Acids Res., 41, 2013
|
|
4IJT
 
 | | Human p53 core domain with hot spot mutation R273H (form II) | | Descriptor: | 1,2-ETHANEDIOL, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, Cellular tumor antigen p53, ... | | Authors: | Rozenberg, H, Eldar, A, Diskin-Posner, Y, Shakked, Z. | | Deposit date: | 2012-12-23 | | Release date: | 2013-08-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural studies of p53 inactivation by DNA-contact mutations and its rescue by suppressor mutations via alternative protein-DNA interactions.
Nucleic Acids Res., 41, 2013
|
|
6FJ5
 | | New Insights into the Role of DNA Shape on Its Recognition by p53 Proteins (complex p53DBD-AGG-HG) | | Descriptor: | 1,2-ETHANEDIOL, Cellular tumor antigen p53, DNA, ... | | Authors: | Golovenko, D, Rozenberg, H, Shakked, Z. | | Deposit date: | 2018-01-20 | | Release date: | 2018-06-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.051 Å) | | Cite: | New Insights into the Role of DNA Shape on Its Recognition by p53 Proteins.
Structure, 26, 2018
|
|
4LOF
 
 | | Human p53 Core Domain Mutant V157F/N235K/N239Y | | Descriptor: | Cellular tumor antigen p53, ZINC ION | | Authors: | Wallentine, B.D, Wang, Y, Luecke, H. | | Deposit date: | 2013-07-12 | | Release date: | 2013-07-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of oncogenic, suppressor and rescued p53 core-domain variants: mechanisms of mutant p53 rescue.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4LOE
 
 | | Human p53 Core Domain Mutant N239Y | | Descriptor: | Cellular tumor antigen p53, ZINC ION | | Authors: | Wallentine, B.D, Wang, Y, Luecke, H. | | Deposit date: | 2013-07-12 | | Release date: | 2013-07-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structures of oncogenic, suppressor and rescued p53 core-domain variants: mechanisms of mutant p53 rescue.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
7XZZ
 
 | | Cryo-EM structure of the nucleosome in complex with p53 | | Descriptor: | Cellular tumor antigen p53, DNA (169-MER), Histone H2A type 1-B/E, ... | | Authors: | Nishimura, M, Nozawa, K, Takizawa, Y, Kurumizaka, H. | | Deposit date: | 2022-06-03 | | Release date: | 2022-10-12 | | Last modified: | 2023-02-15 | | Method: | ELECTRON MICROSCOPY (4.07 Å) | | Cite: | Structural basis for p53 binding to its nucleosomal target DNA sequence.
Pnas Nexus, 1, 2022
|
|
2J1W
 
 | |
8GCR
 
 | | HPV16 E6-E6AP-p53 complex | | Descriptor: | Cellular tumor antigen p53, Maltose/maltodextrin-binding periplasmic protein,Protein E6, Ubiquitin-protein ligase E3A, ... | | Authors: | Bratkowski, M.A, Wang, J.C.K, Hao, Q, Nile, A.H. | | Deposit date: | 2023-03-02 | | Release date: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.38 Å) | | Cite: | Structure of the p53 degradation complex from HPV16.
Nat Commun, 15, 2024
|
|
8DC4
 
 | |
8DC6
 
 | | Crystal structure of p53 Y220C covalently bound to indole KG6 | | Descriptor: | 1-(2-methylprop-2-enoyl)-1H-indole-3-carbaldehyde, bound form, Cellular tumor antigen p53, ... | | Authors: | Guiley, K.Z, Shokat, K.M. | | Deposit date: | 2022-06-15 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.60000908 Å) | | Cite: | A Small Molecule Reacts with the p53 Somatic Mutant Y220C to Rescue Wild-type Thermal Stability.
Cancer Discov, 13, 2023
|
|
8DC7
 
 | | Crystal structure of p53 Y220C covalently bound to indole KG10 | | Descriptor: | 4-[4-(4-methylpiperazin-1-yl)phenyl]-1-(2-methylprop-2-enoyl)-1H-indole-3-carbaldehyde, bound form, Cellular tumor antigen p53, ... | | Authors: | Guiley, K.Z, Shokat, K.M. | | Deposit date: | 2022-06-15 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9870069 Å) | | Cite: | A Small Molecule Reacts with the p53 Somatic Mutant Y220C to Rescue Wild-type Thermal Stability.
Cancer Discov, 13, 2023
|
|
8DC8
 
 | | Crystal structure of p53 Y220C covalently bound to azaindole KG13 | | Descriptor: | 2-methyl-1-[(4P)-3-methyl-4-(2-methyl-1,2,3,4-tetrahydroisoquinolin-6-yl)-1H-pyrrolo[2,3-c]pyridin-1-yl]prop-2-en-1-one, bound form, Cellular tumor antigen p53, ... | | Authors: | Guiley, K.Z, Shokat, K.M. | | Deposit date: | 2022-06-15 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7200973 Å) | | Cite: | A Small Molecule Reacts with the p53 Somatic Mutant Y220C to Rescue Wild-type Thermal Stability.
Cancer Discov, 13, 2023
|
|
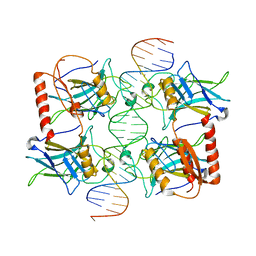
3Q06
 
 | |
7XZX
 
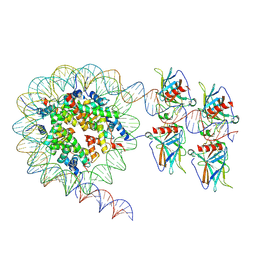 | | Cryo-EM structure of the nucleosome in complex with p53 DNA-binding domain | | Descriptor: | Cellular tumor antigen p53, DNA (193-MER), Histone H2A type 1-B/E, ... | | Authors: | Nishimura, M, Nozawa, K, Takizawa, Y, Kurumizaka, H. | | Deposit date: | 2022-06-03 | | Release date: | 2022-10-12 | | Last modified: | 2023-02-15 | | Method: | ELECTRON MICROSCOPY (4.53 Å) | | Cite: | Structural basis for p53 binding to its nucleosomal target DNA sequence.
Pnas Nexus, 1, 2022
|
|
3Q01
 
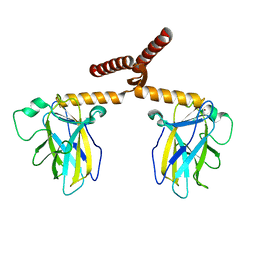 | |
4CRI
 
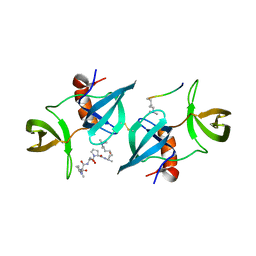 | | Crystal Structure of 53BP1 tandem tudor domains in complex with methylated K810 Rb peptide | | Descriptor: | RB1 PROTEIN, TUMOR SUPPRESSOR P53-BINDING PROTEIN 1 | | Authors: | Krojer, T, Johansson, C, Gileadi, C, Fedorov, O, Carr, S, La Thangue, N.B, Vollmar, M, Crawley, L, von Delft, F, Bountra, C, Arrowsmith, C.H, Edwards, A, Oppermann, U. | | Deposit date: | 2014-02-26 | | Release date: | 2014-08-06 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Lysine Methylation-Dependent Binding of 53BP1 to the Prb Tumor Suppressor.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
3Q05
 
 | |
5G4N
 
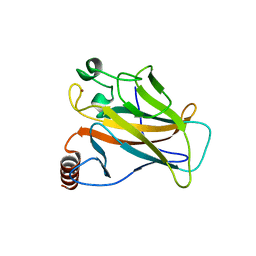 | | Crystal structure of the p53 cancer mutant Y220C in complex with a difluorinated derivative of the small molecule stabilizer Phikan083 | | Descriptor: | 1-[9-(2,2-difluoroethyl)-9H-carbazol-3-yl]-N-methylmethanamine, CELLULAR TUMOR ANTIGEN P53, GLYCEROL, ... | | Authors: | Joerger, A.C, Bauer, M, Jones, R.N, Spencer, J. | | Deposit date: | 2016-05-13 | | Release date: | 2016-06-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Harnessing Fluorine-Sulfur Contacts and Multipolar Interactions for the Design of P53 Mutant Y220C Rescue Drugs.
Acs Chem.Biol., 11, 2016
|
|
5G4O
 
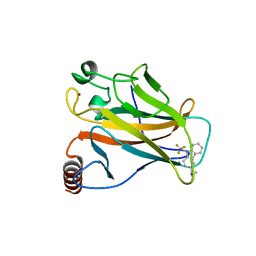 | | Crystal structure of the p53 cancer mutant Y220C in complex with a trifluorinated derivative of the small molecule stabilizer Phikan083 | | Descriptor: | CELLULAR TUMOR ANTIGEN P53, N,N-dimethyl-1-[9-(2,2,2-trifluoroethyl)-9H-carbazol-3-yl]methanamine, ZINC ION | | Authors: | Joerger, A.C, Bauer, M, Baud, M.G.J, Spencer, J. | | Deposit date: | 2016-05-13 | | Release date: | 2016-06-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Harnessing Fluorine-Sulfur Contacts and Multipolar Interactions for the Design of P53 Mutant Y220C Rescue Drugs.
Acs Chem.Biol., 11, 2016
|
|
5G4M
 
 | |
7DVD
 
 | | The crystal structure of p53 DNA binding domain and PUMA complex | | Descriptor: | Bcl-2-binding component 3, isoforms 1/2, Cellular tumor antigen p53, ... | | Authors: | Han, C.W, Lee, H.N, Jeong, M.S, Jang, S.B. | | Deposit date: | 2021-01-13 | | Release date: | 2021-08-04 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Structural basis of the p53 DNA binding domain and PUMA complex.
Biochem.Biophys.Res.Commun., 548, 2021
|
|
5BUA
 
 | |
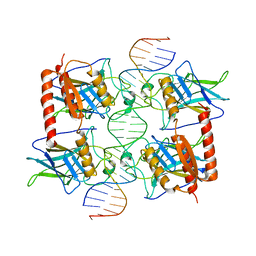
2GEQ
 
 | | Crystal Structure of a p53 Core Dimer Bound to DNA | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5'-D(*GP*CP*GP*TP*GP*AP*GP*CP*AP*TP*GP*CP*TP*CP*AP*C)-3', Cellular tumor antigen p53, ... | | Authors: | Ho, W.C, Fitzgerald, M.X, Marmorstein, R. | | Deposit date: | 2006-03-20 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the p53 Core Domain Dimer Bound to DNA.
J.Biol.Chem., 281, 2006
|
|
7EAX
 | | Crystal complex of p53-V272M and antimony ion | | Descriptor: | ANTIMONY (III) ION, Cellular tumor antigen p53, ZINC ION | | Authors: | Lu, M, Tang, Y. | | Deposit date: | 2021-03-08 | | Release date: | 2022-02-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Repurposing antiparasitic antimonials to noncovalently rescue temperature-sensitive p53 mutations.
Cell Rep, 39, 2022
|
|
