6QYC
 
 | |
8PC4
 
 | | MEMBRANE TARGET COMPLEX 1 | | Descriptor: | (3ALPHA,5BETA,12ALPHA)-3,12-DIHYDROXYCHOLAN-24-OIC ACID, 1,2-Distearoyl-sn-glycerophosphoethanolamine, 6-chloro-3,4-dihydro-2H-1,2,4-benzothiadiazine-7-sulfonamide 1,1-dioxide, ... | | Authors: | Garau, G. | | Deposit date: | 2023-06-09 | | Release date: | 2024-06-26 | | Last modified: | 2025-04-02 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | NAPE-PLD is target of thiazide diuretics.
Cell Chem Biol, 32, 2025
|
|
8SMP
 | | Xenopus laevis hyaluronan synthase 1, UDP-bound, gating loop inserted state | | Descriptor: | Fab15 heavy chain, Fab15 light chain, Hyaluronan synthase 1, ... | | Authors: | Gorniak, I, Zimmer, J. | | Deposit date: | 2023-04-26 | | Release date: | 2024-05-01 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural insights into translocation and tailored synthesis of hyaluronan.
Nat.Struct.Mol.Biol., 32, 2025
|
|
6OT1
 
 | |
8DOC
 
 | | Crystal structure of RPE65 in complex with compound 16e and palmitate | | Descriptor: | (1R)-1-[3-(cyclohexylmethoxy)phenyl]-3-(methylamino)propan-1-ol, 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, FE (II) ION, ... | | Authors: | Bassetto, M, Kiser, P.D. | | Deposit date: | 2022-07-12 | | Release date: | 2023-05-24 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Tuning the Metabolic Stability of Visual Cycle Modulators through Modification of an RPE65 Recognition Motif.
J.Med.Chem., 66, 2023
|
|
9B2P
 | | Structure of the quorum quenching lactonase GcL D122N mutant - bimetallic metal center - C2 space group | | Descriptor: | 1,2-ETHANEDIOL, COBALT (II) ION, FE (III) ION, ... | | Authors: | Corbella, M, Bravo, J.A, Demkiv, A.O, Calixto, A.R, Sompiyachoke, K, Bergonzi, C, Kamerlin, S.C.L, Elias, M. | | Deposit date: | 2024-03-15 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Catalytic Redundancies and Conformational Plasticity Drives Selectivity and Promiscuity in Quorum Quenching Lactonases.
Jacs Au, 4, 2024
|
|
7NBD
 
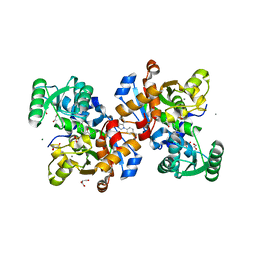 | | Crystal structure of human serine racemase in complex with DSiP fragment Z235449082, XChem fragment screen. | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | Koulouris, C.R, Roe, S.M. | | Deposit date: | 2021-01-26 | | Release date: | 2021-03-03 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.865 Å) | | Cite: | Tyrosine 121 moves revealing a ligandable pocket that couples catalysis to ATP-binding in serine racemase.
Commun Biol, 5, 2022
|
|
3IF6
 
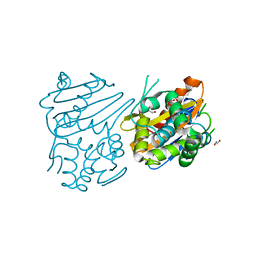 | | Crystal structure of OXA-46 beta-lactamase from P. aeruginosa | | Descriptor: | 1,2-ETHANEDIOL, HEXAETHYLENE GLYCOL, L(+)-TARTARIC ACID, ... | | Authors: | Docquier, J.D, Benvenuti, M, Calderone, V, Giuliani, F, Kapetis, D, De Luca, F, Rossolini, G.M, Mangani, S. | | Deposit date: | 2009-07-24 | | Release date: | 2010-03-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the narrow-spectrum OXA-46 class D beta-lactamase: relationship between active-site lysine carbamylation and inhibition by polycarboxylates
Antimicrob.Agents Chemother., 54, 2010
|
|
7NQ8
 
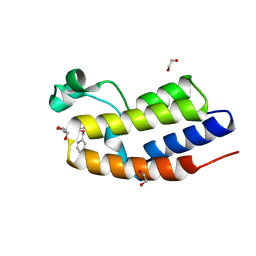 | | C-TERMINAL BROMODOMAIN OF HUMAN BRD2 WITH N-ethyl-3-(1-methyl-1H-1,2,3-triazol-4-yl)-4-(pyridin-2-ylmethoxy)benzamide | | Descriptor: | 1,2-ETHANEDIOL, Bromodomain-containing protein 2, N-ethyl-3-(1-methyl-1H-1,2,3-triazol-4-yl)-4-(pyridin-2-ylmethoxy)benzamide | | Authors: | Chung, C.W. | | Deposit date: | 2021-03-01 | | Release date: | 2021-03-24 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.602 Å) | | Cite: | Template-Hopping Approach Leads to Potent, Selective, and Highly Soluble Bromo and Extraterminal Domain (BET) Second Bromodomain (BD2) Inhibitors.
J.Med.Chem., 64, 2021
|
|
7RJP
 
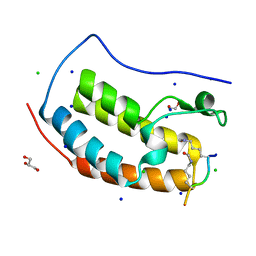 | | Crystal structure of human Bromodomain containing protein 4 (BRD4) in complex with SHMT | | Descriptor: | 1,2-ETHANEDIOL, Bromodomain-containing protein 4, CHLORIDE ION, ... | | Authors: | Fedorov, E, Islam, K, Ghosh, A. | | Deposit date: | 2021-07-21 | | Release date: | 2022-08-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Uncovering the Bromodomain Interactome using Site-Specific Azide-Acetyllysine Photochemistry, Proteomic Profiling and Structural Characterization
To Be Published
|
|
7NQI
 
 | | C-TERMINAL BROMODOMAIN OF HUMAN BRD2 WITH 2-benzyl-N-cyclopropyl-6-(1-methyl-1H-1,2,3-triazol-4-yl)isonicotinamide | | Descriptor: | 1,2-ETHANEDIOL, 2-benzyl-N-cyclopropyl-6-(1-methyl-1H-1,2,3-triazol-4-yl)isonicotinamide, Bromodomain-containing protein 2 | | Authors: | Chung, C.W. | | Deposit date: | 2021-03-01 | | Release date: | 2021-03-24 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.603 Å) | | Cite: | Template-Hopping Approach Leads to Potent, Selective, and Highly Soluble Bromo and Extraterminal Domain (BET) Second Bromodomain (BD2) Inhibitors.
J.Med.Chem., 64, 2021
|
|
8IMR
 | | Structure of ligand-free human macrophage migration inhibitory factor | | Descriptor: | 1,2-ETHANEDIOL, ISOPROPYL ALCOHOL, Macrophage migration inhibitory factor, ... | | Authors: | Sugishima, K, Noguchi, K, Yohda, M, Odaka, M, Matsumura, H. | | Deposit date: | 2023-03-07 | | Release date: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Identification of methotrexate as an inhibitor of macrophage migration inhibitory factor by high-resolution crystal structure analysis
To Be Published
|
|
6B1Z
 
 | |
5NHS
 
 | | The crystal structure of Xanthomonas albilineans N5, N10-methylenetetrahydrofolate dehydrogenase-cyclohydrolase (FolD) | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional protein FolD | | Authors: | Bueno, R.V, Guido, R.V.C, Maluf, F.V, Lima, G.M.A, Oliveira, A.A. | | Deposit date: | 2017-03-22 | | Release date: | 2018-05-16 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | The crystal structure of Xanthomonas albilineans N5, N10-methylenetetrahydrofolate dehydrogenase-cyclohydrolase (FolD)
To Be Published
|
|
8IYM
 
 | | Crystal structure of a protein acetyltransferase, HP0935 | | Descriptor: | 1,2-ETHANEDIOL, N-acetyltransferase domain-containing protein, POTASSIUM ION, ... | | Authors: | Dadireddy, V, Mahanta, P, Kumar, A, Desirazu, R.N, Ramakumar, S. | | Deposit date: | 2023-04-05 | | Release date: | 2024-04-10 | | Last modified: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Biochemical and structural characterization of a GNAT superfamily protein acetyltransferase from Helicobacter pylori.
J.Biol.Chem., 301, 2025
|
|
5A61
 
 | | Crystal structure of full-length E. coli ygiF in complex with tripolyphosphate and two manganese ions. | | Descriptor: | 1,2-ETHANEDIOL, INORGANIC TRIPHOSPHATASE, MANGANESE (II) ION, ... | | Authors: | Martinez, J, Truffault, V, Hothorn, M. | | Deposit date: | 2015-06-23 | | Release date: | 2015-08-05 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural Determinants for Substrate Binding and Catalysis in Triphosphate Tunnel Metalloenzymes.
J.Biol.Chem., 290, 2015
|
|
8AVT
 
 | | Racemic protein crystal structure of aureocin A53 from Staphylococcus aureus in the presence of glycerol 3-phosphate | | Descriptor: | 1,2-ETHANEDIOL, Bacteriocin aureocin A53, D-Aureocin A53, ... | | Authors: | Lander, A.J, Baumann, P, Rizkallah, P, Jin, Y, Luk, L.Y.P. | | Deposit date: | 2022-08-26 | | Release date: | 2023-07-26 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Roles of inter- and intramolecular tryptophan interactions in membrane-active proteins revealed by racemic protein crystallography.
Commun Chem, 6, 2023
|
|
9EWT
 
 | | Optimisation of Potent, Efficacious, Selective and Blood-Brain Barrier Penetrating Inhibitors Targeting EGFR Exon20 Insertion Mutations | | Descriptor: | 1-[2-[3-[(E)-2-(5-cyclopropyl-1,3-oxazol-2-yl)ethenyl]phenyl]-3-pyridin-4-yl-4,6-dihydropyrrolo[3,4-d]imidazol-5-yl]propan-1-one, Epidermal growth factor receptor | | Authors: | Hargreaves, D. | | Deposit date: | 2024-04-04 | | Release date: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.019 Å) | | Cite: | Optimization of Potent, Efficacious, Selective and Blood-Brain Barrier Penetrating Inhibitors Targeting EGFR Exon20 Insertion Mutations.
J.Med.Chem., 2024
|
|
8AVU
 
 | | Racemic protein crystal structure of aureocin A53 from Staphylococcus aureus in the dimeric state | | Descriptor: | 1,2-ETHANEDIOL, Bacteriocin aureocin A53, D-Aureocin A53, ... | | Authors: | Lander, A.J, Baumann, P, Rizkallah, P, Jin, Y, Luk, L.Y.P. | | Deposit date: | 2022-08-26 | | Release date: | 2023-07-26 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (0.89 Å) | | Cite: | Roles of inter- and intramolecular tryptophan interactions in membrane-active proteins revealed by racemic protein crystallography.
Commun Chem, 6, 2023
|
|
9EVO
 
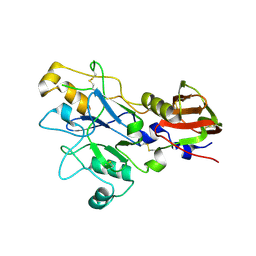 | |
7NNG
 
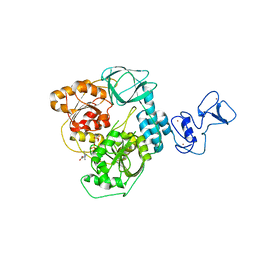 | | Crystal structure of the SARS-CoV-2 helicase in complex with Z2327226104 | | Descriptor: | 1-(2-methylphenyl)-1,2,3-triazole-4-carboxylic acid, PHOSPHATE ION, SARS-CoV-2 helicase NSP13, ... | | Authors: | Newman, J.A, Yosaatmadja, Y, Douangamath, A, Bountra, C, Gileadi, O. | | Deposit date: | 2021-02-24 | | Release date: | 2021-04-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Structure, mechanism and crystallographic fragment screening of the SARS-CoV-2 NSP13 helicase.
Nat Commun, 12, 2021
|
|
9IQW
 
 | |
8IOG
 
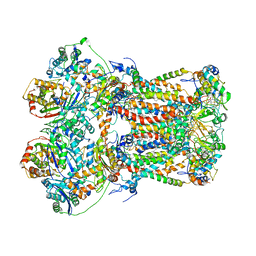 | |
6QF4
 
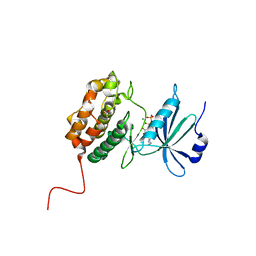 | | X-Ray structure of human Serine/Threonine Kinase 17B (STK17B) aka DRAK2 in complex with ADP obtained by on-chip soaking | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Lieske, J, Cerv, M, Kreida, S, Barthelmess, M, Fischer, P, Pakendorf, T, Yefanov, O, Mariani, V, Seine, T, Ross, B.H, Crosas, E, Lorbeer, O, Burkhardt, A, Lane, T.J, Guenther, S, Bergtholdt, J, Schoen, S, Tornroth-Horsefield, S, Chapman, H.N, Meents, A. | | Deposit date: | 2019-01-09 | | Release date: | 2019-07-10 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | On-chip crystallization for serial crystallography experiments and on-chip ligand-binding studies.
Iucrj, 6, 2019
|
|
1ETT
 
 | |
