3EQR
 
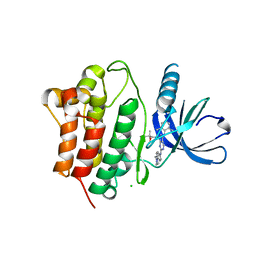 | | Crystal Structure of Ack1 with compound T74 | | Descriptor: | Activated CDC42 kinase 1, CHLORIDE ION, N~3~-(2,6-dimethylphenyl)-1-(3-methoxy-3-methylbutyl)-N~6~-(4-piperazin-1-ylphenyl)-1H-pyrazolo[3,4-d]pyrimidine-3,6-diamine | | Authors: | Liu, J, Wang, Z, Walker, N.P.C. | | Deposit date: | 2008-10-01 | | Release date: | 2008-12-02 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Identification and optimization of N3,N6-diaryl-1H-pyrazolo[3,4-d]pyrimidine-3,6-diamines as a novel class of ACK1 inhibitors.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
5B0X
 
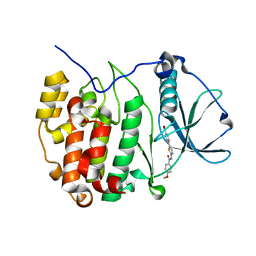 | | Crystal structure of the CK2a/benzoic acid derivative complex | | Descriptor: | 4-[2-[(4-methoxyphenyl)carbonylamino]-1,3-thiazol-5-yl]benzoic acid, Casein kinase II subunit alpha | | Authors: | Kinoshita, T, Nakanishi, I. | | Deposit date: | 2015-11-13 | | Release date: | 2016-03-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-activity relationship study of 4-(thiazol-5-yl)benzoic acid derivatives as potent protein kinase CK2 inhibitors
Bioorg.Med.Chem., 24, 2016
|
|
3O6I
 
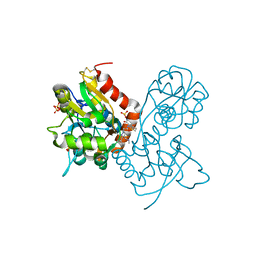 | | Ligand-binding domain of GluA2 (flip) ionotropic glutamate receptor in complex with an allosteric modulator | | Descriptor: | 2-[({3-tert-butyl-4-[(methylamino)methyl]-1H-pyrazol-1-yl}acetyl)amino]-4,5,6,7-tetrahydro-1-benzothiophene-3-carboxamide, GLUTAMIC ACID, Glutamate receptor 2, ... | | Authors: | Maclean, J.K.F, Campbell, R.A, Cumming, I.A, Gillen, K.J, Gillespie, J, Jamieson, C, Kazemier, B, Kiczun, M, Lamont, Y, Lyons, A.J, Martin, F, Moir, E.M, Morrow, J.A, Pantling, J, Rankovic, Z, Smith, L. | | Deposit date: | 2010-07-29 | | Release date: | 2010-09-15 | | Last modified: | 2017-08-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | A novel series of positive modulators of the AMPA receptor: structure-based lead optimization.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
2ZYB
 
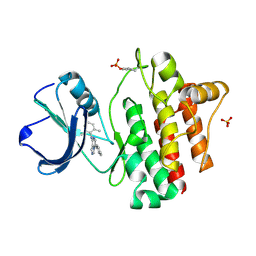 | |
3F7B
 
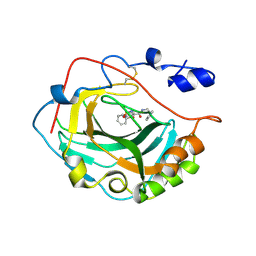 | | Crystal Structure of soluble domain of CA4 in complex with small molecule. | | Descriptor: | Carbonic anhydrase 4, N-(2-phenylethyl)-2-(phenylsulfanyl)-5-sulfamoylpyridine-3-carboxamide, ZINC ION | | Authors: | Greasley, S.E, Ferre, R.A.A, Pauly, T.A, Paz, R. | | Deposit date: | 2008-11-07 | | Release date: | 2009-09-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Thioether benzenesulfonamide inhibitors of carbonic anhydrases II and IV: structure-based drug design, synthesis, and biological evaluation.
Bioorg.Med.Chem., 18, 2010
|
|
3O29
 
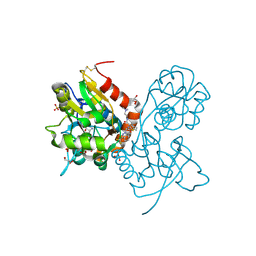 | | Ligand-binding domain of GluA2 (flip) ionotropic glutamate receptor in complex with an allosteric modulator | | Descriptor: | GLUTAMIC ACID, GLYCEROL, Glutamate receptor 2, ... | | Authors: | Maclean, J.K.F, Basten, S, Campbell, R.A, Cumming, I.A, Gillen, K.J, Gillespie, J, Jamieson, C, Kazemier, B, Kiczun, M, Lamont, Y, Lyons, A.J, Moir, E.M, Morrow, J.A, Papakosta, M, Rankovic, Z, Smith, L. | | Deposit date: | 2010-07-22 | | Release date: | 2010-09-15 | | Last modified: | 2017-08-09 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | A novel series of positive modulators of the AMPA receptor: discovery and structure based hit-to-lead studies.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3IG6
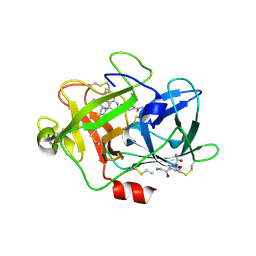 
 | | Low molecular weigth human Urokinase type Plasminogen activator 2-[6-(3'-Aminomethyl-biphenyl-3-yloxy)-4-(3-dimethylamino-pyrrolidin-1-yl)-3,5-difluoro-pyridin-2-yloxy]-4-dimethylamino-benzoic acid complex | | Descriptor: | 2-[(6-{[3'-(aminomethyl)biphenyl-3-yl]oxy}-4-[(3R)-3-(dimethylamino)pyrrolidin-1-yl]-3,5-difluoropyridin-2-yl)oxy]-4-(dimethylamino)benzoic acid, PHOSPHATE ION, Urokinase-type plasminogen activator | | Authors: | Adler, M, Whitlow, M. | | Deposit date: | 2009-07-27 | | Release date: | 2009-10-13 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Identification of orally bioavailable, non-amidine inhibitors of Urokinase Plasminogen Activator (uPA)
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3O28
 
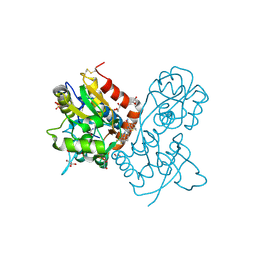 | | Ligand-binding domain of GluA2 (flip) ionotropic glutamate receptor in complex with an allosteric modulator | | Descriptor: | 2-({[3-(trifluoromethyl)-4,5,6,7-tetrahydro-1H-indazol-1-yl]acetyl}amino)-4,5,6,7-tetrahydro-1-benzothiophene-3-carboxamide, DIMETHYL SULFOXIDE, GLUTAMIC ACID, ... | | Authors: | Maclean, J.K.F, Basten, S, Campbell, R.A, Cumming, I.A, Gillen, K.J, Gillespie, J, Jamieson, C, Kazemier, B, Kiczun, M, Lamont, Y, Lyons, A.J, Moir, E.M, Morrow, J.A, Papakosta, M, Rankovic, Z, Smith, L. | | Deposit date: | 2010-07-22 | | Release date: | 2010-09-15 | | Last modified: | 2017-08-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A novel series of positive modulators of the AMPA receptor: discovery and structure based hit-to-lead studies.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
2XRU
 
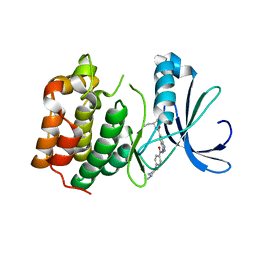 | | AURORA-A T288E COMPLEXED WITH PHA-828300 | | Descriptor: | 3-({[4-(4-METHYLPIPERAZIN-1-YL)PHENYL]CARBONYL}AMINO)-N-[(1R)-1-PHENYLPROPYL]-1H-THIENO[3,2-C]PYRAZOLE-5-CARBOXAMIDE, SERINE/THREONINE-PROTEIN KINASE 6 | | Authors: | Bindi, S, Fancelli, D, Alli, C, Berta, D, Bertrand, J.A, Cameron, A.D, Cappella, P, Carpinelli, P, Cervi, G, Croci, W, D'Anello, M, Forte, B, LauraGiorgini, M, Marsiglio, A, Moll, J, Pesenti, E, Pittala, V, Pulici, M, Riccardi-Sirtori, F, Roletto, F, Soncini, C, Storici, P, Varasi, M, Volpi, D, Zugnoni, P, Vianello, P. | | Deposit date: | 2010-09-22 | | Release date: | 2010-09-29 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Thieno[3,2-C]Pyrazoles: A Novel Class of Aurora Inhibitors with Favorable Antitumor Activity.
Bioorg.Med.Chem., 18, 2010
|
|
4HY0
 
 | | Crystal structure of XIAP BIR3 with T3256336 | | Descriptor: | (3S,7R,8aR)-2-{(2S)-2-(4,4-difluorocyclohexyl)-2-[(N-methyl-L-alanyl)amino]acetyl}-N-[(4R)-3,4-dihydro-2H-chromen-4-yl]-7-ethoxyoctahydropyrrolo[1,2-a]pyrazine-3-carboxamide, E3 ubiquitin-protein ligase XIAP, ZINC ION | | Authors: | Snell, G.S, Dougan, D.R. | | Deposit date: | 2012-11-12 | | Release date: | 2013-01-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Design and Synthesis of Potent Inhibitor of Apoptosis (IAP) Proteins Antagonists Bearing an Octahydropyrrolo[1,2-a]pyrazine Scaffold as a Novel Proline Mimetic.
J.Med.Chem., 56, 2013
|
|
3R16
 
 | | Human CAII bound to N-(4-sulfamoylphenyl)-2-(thiophen-2-yl) acetamide | | Descriptor: | Carbonic anhydrase 2, DIMETHYL SULFOXIDE, GLYCEROL, ... | | Authors: | Biswas, S, McKenna, R, Supuran, C.T. | | Deposit date: | 2011-03-09 | | Release date: | 2011-05-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Conformational variability of different sulfonamide inhibitors with thienyl-acetamido moieties attributes to differential binding in the active site of cytosolic human carbonic anhydrase isoforms.
Bioorg.Med.Chem., 19, 2011
|
|
2GD8
 
 | | Crystal structure analysis of the human carbonic anhydrase II in complex with a 2-substituted estradiol bis-sulfamate | | Descriptor: | (9BETA,13ALPHA,14BETA,17ALPHA)-2-METHOXYESTRA-1,3,5(10)-TRIENE-3,17-DIYL DISULFAMATE, CHLORIDE ION, Carbonic anhydrase 2, ... | | Authors: | De Simone, G, Di Fiore, A. | | Deposit date: | 2006-03-15 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | 2-substituted estradiol bis-sulfamates, multitargeted antitumor agents: synthesis, in vitro SAR, protein crystallography, and in vivo activity.
J.Med.Chem., 49, 2006
|
|
4HY2
 
 | |
2IT4
 
 | | X ray structure of the complex between Carbonic Anhydrase I and the phosphonate antiviral drug foscarnet | | Descriptor: | Carbonic anhydrase 1, PHOSPHONOFORMIC ACID, ZINC ION | | Authors: | Temperini, C, Innocenti, A, Guerri, A, Scozzafava, A, Supuran, C.T. | | Deposit date: | 2006-10-19 | | Release date: | 2007-09-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Phosph(on)ate as a zinc-binding group in metalloenzyme inhibitors: X-ray crystal structure of the antiviral drug foscarnet complexed to human carbonic anhydrase I.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
4I9H
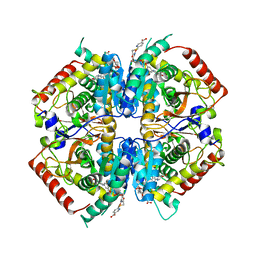 
 | | Crystal structure of rabbit LDHA in complex with AP28669 | | Descriptor: | 1-O-[3-(5-carboxypyridin-2-yl)-5-fluorophenyl]-6-O-[4-({[(5-carboxypyridin-2-yl)sulfanyl]acetyl}amino)-2-chloro-5-methoxyphenyl]-D-mannitol, L-lactate dehydrogenase A chain | | Authors: | Zhou, T, Stephan, Z.G, Kohlmann, A, Li, F, Commodore, L, Greenfield, M.T, Shakespeare, W.C, Zhu, X, Dalgarno, D.C. | | Deposit date: | 2012-12-05 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Fragment growing and linking lead to novel nanomolar lactate dehydrogenase inhibitors.
J.Med.Chem., 56, 2013
|
|
3FCO
 
 | | Crystal Structure of 11beta-Hydroxysteroid Dehydrogenase 1 (11b-HSD1) in Complex with Benzamide Inhibitor | | Descriptor: | Corticosteroid 11-beta-dehydrogenase isozyme 1, N-cyclopropyl-N-(cis-4-cyclopropyl-4-hydroxycyclohexyl)-4-[(1S)-2,2,2-trifluoro-1-hydroxy-1-methylethyl]benzamide, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Wang, Z, Sudom, A, Walker, N.P. | | Deposit date: | 2008-11-21 | | Release date: | 2009-12-01 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Optimization of novel di-substituted cyclohexylbenzamide derivatives as potent 11 beta-HSD1 inhibitors.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3RSV
 
 | |
4I8X
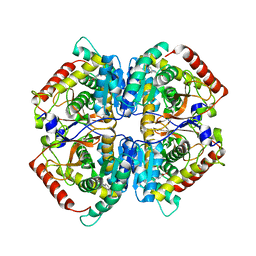 
 | | Crystal structure of rabbit LDHA in complex with AP27460 | | Descriptor: | 6-phenylpyridine-3-carboxylic acid, L-lactate dehydrogenase A chain | | Authors: | Zhou, T, Stephan, Z.G, Kohlmann, A, Li, F, Commodore, L, Greenfield, M.T, Zhu, X, Dalgarno, D.C. | | Deposit date: | 2012-12-04 | | Release date: | 2013-01-23 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Fragment growing and linking lead to novel nanomolar lactate dehydrogenase inhibitors.
J.Med.Chem., 56, 2013
|
|
3RTH
 
 | |
3L8V
 
 | |
3RSX
 
 | |
3D0B
 
 | | Crystal Structure of Benzamide Tetrahydro-4H-carbazol-4-one bound to Hsp90 | | Descriptor: | 2-[(2-methoxyethyl)amino]-4-(4-oxo-1,2,3,4-tetrahydro-9H-carbazol-9-yl)benzamide, Heat shock protein HSP 90-alpha | | Authors: | Barta, T.E, Veal, J.M, Huang, K.H, Hall, S.H. | | Deposit date: | 2008-05-01 | | Release date: | 2008-06-17 | | Last modified: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Discovery of benzamide tetrahydro-4H-carbazol-4-ones as novel small molecule inhibitors of Hsp90
BIOORG.MED.CHEM.LETT., 18, 2008
|
|
3RTM
 
 | |
3RU1
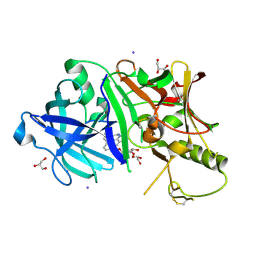 
 | |
5AL5
 
 | | Crystal structure of TNKS2 in complex with 2-(4-((pyridin-4-yl)methyl) piperazin-1-yl)-3,4,5,6,7,8-hexahydroquinazolin-4-one | | Descriptor: | 2-(4-((pyridin-4-yl)methyl)piperazin-1-yl)-3,4,5,6,7,8-hexahydroquinazolin-4-one, GLYCEROL, SULFATE ION, ... | | Authors: | Nkizinkiko, Y, Lehtio, L. | | Deposit date: | 2015-03-06 | | Release date: | 2015-07-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Discovery of Potent and Selective Nonplanar Tankyrase Inhibiting Nicotinamide Mimics.
Bioorg.Med.Chem., 23, 2015
|
|
