4YW7
 
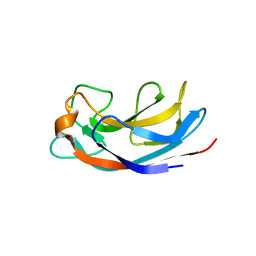 | | Structural Insight into Divalent Galactoside Binding to Pseudomonas aeruginosa lectin LecA | | Descriptor: | (2R,3R,4S,5R,6R,2'R,3'R,4'S,5'R,6'R)-2,2'-([(2R,3R,4S,5S,6S)-3,4-dihydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-2,5-diyl]bis{1H-1,2,3-triazole-1,4-diyl[(2S,3R,4S,5S,6S)-3,4-dihydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-2,5-diyl]-1H-1,2,3-triazole-1,4-diylpropane-3,1-diyloxy})bis[6-(hydroxymethyl)tetrahydro-2H-pyran-3,4,5-triol], CALCIUM ION, PA-I galactophilic lectin, ... | | Authors: | Visini, R, Jin, X, Michaud, G, Bergmann, M, Gillon, E, Imberty, A, Stocker, A, Darbre, T, Pieters, R, Reymond, J.-L. | | Deposit date: | 2015-03-20 | | Release date: | 2015-09-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structural Insight into Multivalent Galactoside Binding to Pseudomonas aeruginosa Lectin LecA.
Acs Chem.Biol., 10, 2015
|
|
4YWA
 
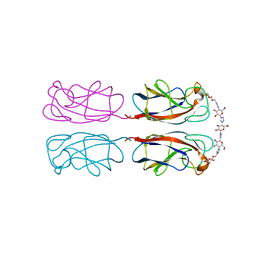 | | Structural Insight into Divalent Galactoside Binding to Pseudomonas aeruginosa lectin LecA | | Descriptor: | (2R,3R,4S,5R,6R,2'R,3'R,4'S,5'R,6'R)-2,2'-([(2R,3R,4S,5S,6S)-3,4-dihydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-2,5-diyl]bis{1H-1,2,3-triazole-1,4-diyl[(2S,3R,4S,5S,6S)-3,4-dihydroxy-6-(hydroxymethyl)tetrahydro-2H-pyran-2,5-diyl]-1H-1,2,3-triazole-1,4-diylmethanediyloxy})bis[6-(hydroxymethyl)tetrahydro-2H-pyran-3,4,5-triol], CALCIUM ION, PA-I galactophilic lectin | | Authors: | Visini, R, Jin, X, Michaud, G, Bergmann, M, Gillon, E, Imberty, A, Stocker, A, Darbre, T, Pieters, R, Reymond, J.-L. | | Deposit date: | 2015-03-20 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.192 Å) | | Cite: | Structural Insight into Multivalent Galactoside Binding to Pseudomonas aeruginosa Lectin LecA.
Acs Chem.Biol., 10, 2015
|
|
6OSJ
 
 | |
6OSL
 
 | |
4ZTS
 
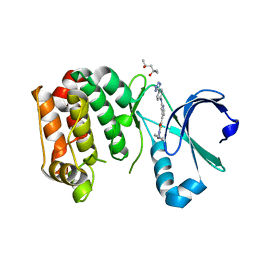 | | Human Aurora A catalytic domain bound to FK1142 | | Descriptor: | (2Z,5Z)-2-[(4-ethylphenyl)imino]-3-methyl-5-[(2-{[4-(1H-tetrazol-5-yl)phenyl]amino}pyridin-4-yl)methylidene]-1,3-thiazolidin-4-one, (4S)-2-METHYL-2,4-PENTANEDIOL, Aurora kinase A | | Authors: | Marcaida, M.J, Kilchmann, F, Schick, T, Reymond, J.L. | | Deposit date: | 2015-05-15 | | Release date: | 2016-07-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Discovery of a Selective Aurora A Kinase Inhibitor by Virtual Screening.
J.Med.Chem., 59, 2016
|
|
1BCS
 
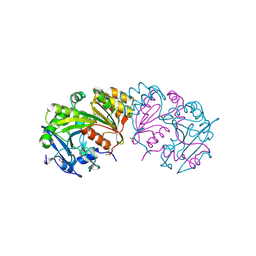 | | COMPLEX OF THE WHEAT SERINE CARBOXYPEPTIDASE, CPDW-II, WITH THE MICROBIAL PEPTIDE ALDEHYDE INHIBITOR, CHYMOSTATIN, AND ARGININE AT 100 DEGREES KELVIN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, ... | | Authors: | Bullock, T.L, Remington, S.J. | | Deposit date: | 1995-11-03 | | Release date: | 1996-03-08 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Peptide aldehyde complexes with wheat serine carboxypeptidase II: implications for the catalytic mechanism and substrate specificity.
J.Mol.Biol., 255, 1996
|
|
5D2A
 
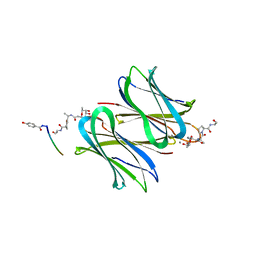 | | Bifunctional dendrimers | | Descriptor: | 3,7-anhydro-2,8-dideoxy-L-glycero-D-gluco-octonic acid, CALCIUM ION, Fucose-binding lectin, ... | | Authors: | Michaud, G, Visini, R, Stocker, A, Darbre, T, Reymond, J.L. | | Deposit date: | 2015-08-05 | | Release date: | 2016-02-10 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.134 Å) | | Cite: | Overcoming antibiotic resistance inPseudomonas aeruginosabiofilms using glycopeptide dendrimers.
Chem Sci, 7, 2016
|
|
4YW6
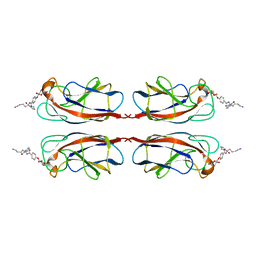 
 | | Structural Insight into Divalent Galactoside Binding to Pseudomonas aeruginosa lectin LecA | | Descriptor: | CALCIUM ION, N-[(2S)-6-amino-1-oxo-1-(pyrrolidin-1-yl)hexan-2-yl]-4-(beta-D-galactopyranosyloxy)benzamide, PA-I galactophilic lectin | | Authors: | Visini, R, Jin, X, Michaud, G, Bergmann, M, Gillon, E, Imberty, A, Stocker, A, Darbre, T, Pieters, R, Reymond, J.-L. | | Deposit date: | 2015-03-20 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural Insight into Multivalent Galactoside Binding to Pseudomonas aeruginosa Lectin LecA.
Acs Chem.Biol., 10, 2015
|
|
8OPH
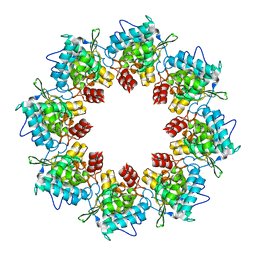 
 | |
1PFZ
 
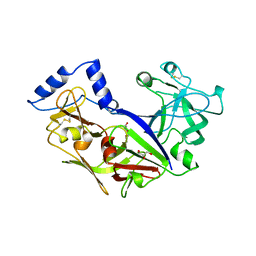 | | PROPLASMEPSIN II FROM PLASMODIUM FALCIPARUM | | Descriptor: | GLYCEROL, PROPLASMEPSIN II | | Authors: | Bernstein, N.K, Cherney, M.M, Loetscher, H, Ridley, R.G, James, M.N.G. | | Deposit date: | 1998-07-07 | | Release date: | 1999-01-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of the novel aspartic proteinase zymogen proplasmepsin II from plasmodium falciparum.
Nat.Struct.Biol., 6, 1999
|
|
9FAB
 
 | | Additional cryo-EM structure of cardiac amyloid AL59 - bent polymorph | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Monoclonal immunoglobulin light chains (LC) | | Authors: | Schulte, T, Speranzini, V, Chaves-Sanjuan, A, Milazzo, M, Ricagno, S. | | Deposit date: | 2024-05-10 | | Release date: | 2024-08-14 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Additional cryo-EM structure of cardiac amyloid AL59 - bent polymorph
To be published
|
|
3VN4
 
 | | Crystal structure of the exosite-containing fragment of human ADAMTS13 (P475S mutant) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, A disintegrin and metalloproteinase with thrombospondin motifs 13, ... | | Authors: | Nakayama, D, Akiyama, M, Takeda, S, Kokame, K, Takagi, J, Miyata, T. | | Deposit date: | 2011-12-21 | | Release date: | 2012-12-26 | | Last modified: | 2024-07-31 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure and enzymatic activity of an ADAMTS-13 mutant with the East Asian-specific P475S polymorphism.
J.Thromb.Haemost., 11, 2013
|
|
4Q2K
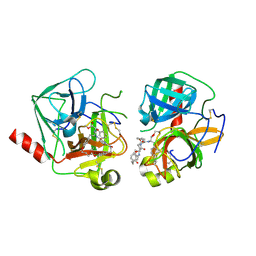 
 | | Bovine alpha chymotrypsin bound to a cyclic peptide inhibitor, 5b | | Descriptor: | (11S)-4,9-dioxo-N-[(2S)-1-oxo-3-phenylpropan-2-yl]-17,22-dioxa-10,30-diazatetracyclo[21.2.2.2~13,16~.1~5,8~]triaconta-1(25),5,7,13,15,23,26,28-octaene-11-carboxamide, Chymotrypsinogen A | | Authors: | Chan, H.Y, Bruning, J.B, Abell, A.D. | | Deposit date: | 2014-04-09 | | Release date: | 2014-07-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Macrocyclic protease inhibitors with reduced peptide character.
Angew.Chem.Int.Ed.Engl., 53, 2014
|
|
7SZR
 
 | | NIK bound to inhibitor G02792917 | | Descriptor: | 1-(3-{[(1R,4R,5S)-4-hydroxy-2-methyl-3-oxo-2-azabicyclo[3.1.0]hexan-4-yl]ethynyl}phenyl)-1H-pyrazolo[3,4-b]pyridine-3-carboxamide, Mitogen-activated protein kinase kinase kinase 14, SULFATE ION | | Authors: | Liau, N.P.D, Hymowitz, S.G. | | Deposit date: | 2021-11-29 | | Release date: | 2023-06-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Filling a nick in NIK: Extending the half-life of a NIK inhibitor through structure-based drug design.
Bioorg.Med.Chem.Lett., 89, 2023
|
|
8A96
 
 | | SARS Cov2 Spike RBD in complex with Fab47 | | Descriptor: | Fab47 Heavy chain (variable domain), Fab47 Light chain (variable domain), Spike protein S1 | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2022-06-27 | | Release date: | 2023-05-03 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Immunoglobulin germline gene polymorphisms influence the function of SARS-CoV-2 neutralizing antibodies.
Immunity, 56, 2023
|
|
8A99
 
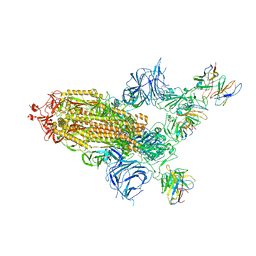 | | SARS Cov2 Spike in 1-up conformation complex with Fab47 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fab47 (Heavy chain variable domain), ... | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2022-06-28 | | Release date: | 2023-05-10 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Immunoglobulin germline gene polymorphisms influence the function of SARS-CoV-2 neutralizing antibodies.
Immunity, 56, 2023
|
|
8A94
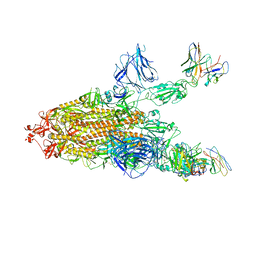 
 | | SARS CoV2 Spike in the 2-up state in complex with Fab47. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fab47 (Heavy chain Variable domain), ... | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2022-06-27 | | Release date: | 2023-05-03 | | Last modified: | 2023-07-26 | | Method: | ELECTRON MICROSCOPY (2.4 Å) | | Cite: | Immunoglobulin germline gene polymorphisms influence the function of SARS-CoV-2 neutralizing antibodies.
Immunity, 56, 2023
|
|
8A95
 
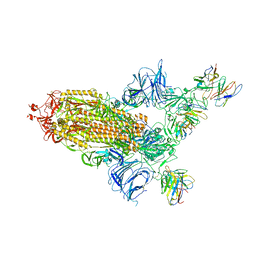 | | SARS Cov2 Spike RBD in complex with Fab47 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Fab47 (Heavy chain variable domain), ... | | Authors: | Hallberg, B.M, Das, H. | | Deposit date: | 2022-06-27 | | Release date: | 2023-08-16 | | Method: | ELECTRON MICROSCOPY (2.64 Å) | | Cite: | Immunoglobulin germline gene polymorphisms influence the function of SARS-CoV-2 neutralizing antibodies.
Immunity, 56, 2023
|
|
8AWT
 
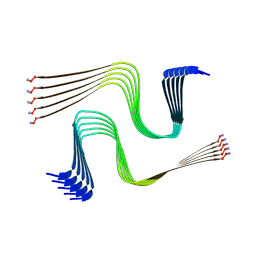 | | IAPP S20G lag-phase fibril polymorph 2PF-P | | Descriptor: | Islet amyloid polypeptide | | Authors: | Wilkinson, M, Xu, Y, Gallardo, R, Radford, S.E, Ranson, N.A. | | Deposit date: | 2022-08-30 | | Release date: | 2024-01-10 | | Last modified: | 2024-10-16 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural evolution of fibril polymorphs during amyloid assembly.
Cell, 186, 2023
|
|
7V4B
 
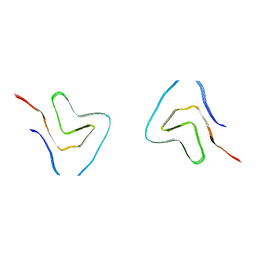 | |
1MD8
 
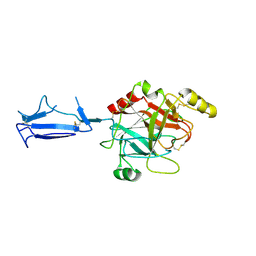 | | Monomeric structure of the active catalytic domain of complement protease C1r | | Descriptor: | C1R COMPLEMENT SERINE PROTEASE | | Authors: | Budayova-Spano, M, Grabarse, W, Thielens, N.M, Hillen, H, Lacroix, M, Schmidt, M, Fontecilla-Camps, J, Arlaud, G.J, Gaboriaud, C. | | Deposit date: | 2002-08-07 | | Release date: | 2003-08-07 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Monomeric structures of the zymogen and active catalytic domain of complement protease c1r: further insights into the c1 activation mechanism
Structure, 10, 2002
|
|
7JI2
 
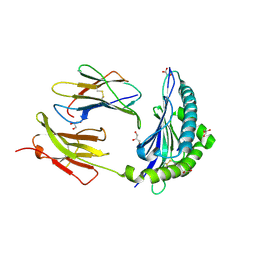 | | Crystal Structure of H2-Kb in complex with a OVA mutant peptide | | Descriptor: | Beta-2-microglobulin, GLYCEROL, H-2 class I histocompatibility antigen, ... | | Authors: | Li, X, Mallis, R.J, Mizsei, R, Tan, K, Reinherz, E.L, Wang, J. | | Deposit date: | 2020-07-22 | | Release date: | 2020-12-23 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Pre-T cell receptors topologically sample self-ligands during thymocyte beta-selection.
Science, 371, 2021
|
|
8BXW
 
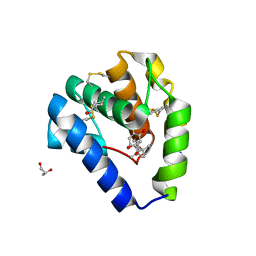 | | Crystal structure of Odorant Binding Protein 5 from Anopheles gambiae (AgamOBP5) with Carvacrol | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1-BUTANOL, 2-methyl-5-propan-2-yl-phenol, ... | | Authors: | Liggri, P.G.V, Tsitsanou, K.E, Zographos, S.E. | | Deposit date: | 2022-12-10 | | Release date: | 2023-03-22 | | Last modified: | 2023-04-12 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The structure of AgamOBP5 in complex with the natural insect repellents Carvacrol and Thymol: Crystallographic, fluorescence and thermodynamic binding studies.
Int.J.Biol.Macromol., 237, 2023
|
|
7JYZ
 
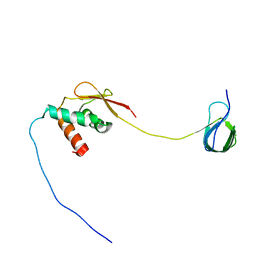 | | Solution NMR structure and dynamics of human Brd3 ET in complex with MLV IN CTD | | Descriptor: | Bromodomain-containing protein 3, Integrase | | Authors: | Aiyer, S, Liu, G, Swapna, G.V.T, Hao, J, Ma, L.C, Roth, M.J, Montelione, G.T. | | Deposit date: | 2020-09-01 | | Release date: | 2021-06-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A common binding motif in the ET domain of BRD3 forms polymorphic structural interfaces with host and viral proteins.
Structure, 29, 2021
|
|
7JQ8
 
 | | Solution NMR structure of human Brd3 ET domain | | Descriptor: | Bromodomain-containing protein 3, Integrase peptide | | Authors: | Aiyer, S, Swapna, G.V.T, Roth, J.M, Montelione, G.T. | | Deposit date: | 2020-08-10 | | Release date: | 2021-06-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | A common binding motif in the ET domain of BRD3 forms polymorphic structural interfaces with host and viral proteins.
Structure, 29, 2021
|
|
