2QTI
 
 | | Crystal structure of the UPF0352 protein SO_2176 from Shewanella oneidensis. NESG target SoR77. | | Descriptor: | UPF0352 protein SO_2176 | | Authors: | Vorobiev, S.M, Su, M, Seetharaman, J, Kuzin, A.P, Wang, D, Cunningham, K, Owens, L, Maglaqui, M, Fang, Y, Xiao, R, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-08-02 | | Release date: | 2007-08-21 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the UPF0352 protein SO_2176 from Shewanella oneidensis.
To be Published
|
|
2QWA
 
 | |
2QWJ
 
 | |
2QFJ
 
 | | Crystal Structure of First Two RRM Domains of FIR Bound to ssDNA from a Portion of FUSE | | Descriptor: | DNA (5'-D(*DTP*DCP*DGP*DGP*DGP*DAP*DTP*DTP*DTP*DTP*DTP*DTP*DAP*DTP*DTP*DTP*DTP*DGP*DTP*DGP*DTP*DTP*DAP*DTP*DT)-3'), FBP-interacting repressor | | Authors: | Crichlow, G.V, Yang, Y, Fan, C, Lolis, E, Braddock, D. | | Deposit date: | 2007-06-27 | | Release date: | 2008-03-04 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Dimerization of FIR upon FUSE DNA binding suggests a mechanism of c-myc inhibition
EMBO J., 27, 2007
|
|
2QGM
 
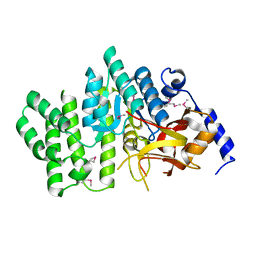 | | Crystal structure of succinoglycan biosynthesis protein at the resolution 1.7 A. Northeast Structural Genomics Consortium target BcR136. | | Descriptor: | Succinoglycan biosynthesis protein | | Authors: | Kuzin, A.P, Abashidze, M, Jayaraman, S, Wang, H, Fang, Y, Maglaqui, M, Ma, L.-C, Xiao, R, Liu, J, Baran, M.C, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-06-29 | | Release date: | 2007-07-24 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of succinoglycan biosynthesis protein at the resolution 1.7 A. Northeast Structural Genomics Consortium target BcR136.
To be Published
|
|
2OVX
 
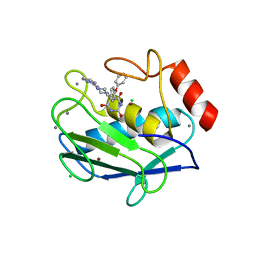 | | MMP-9 active site mutant with barbiturate inhibitor | | Descriptor: | 5-(4-PHENOXYPHENYL)-5-(4-PYRIMIDIN-2-YLPIPERAZIN-1-YL)PYRIMIDINE-2,4,6(2H,3H)-TRIONE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Tochowicz, A, Bode, W, Maskos, K, Goettig, P. | | Deposit date: | 2007-02-15 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of MMP-9 Complexes with Five Inhibitors: Contribution of the Flexible Arg424 Side-chain to Selectivity.
J.Mol.Biol., 371, 2007
|
|
2OP8
 
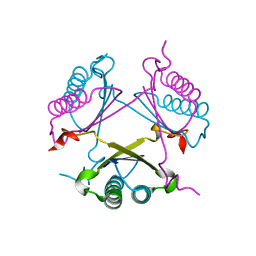 | |
2OL2
 
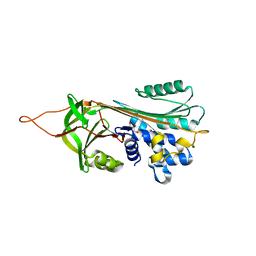 | |
2ORT
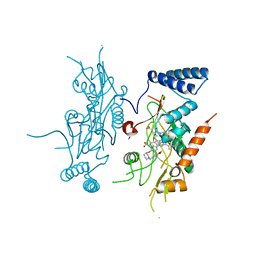 
 | | Murine Inducible Nitric Oxide Synthase Oxygenase Domain (Delta 114) 1-Benzo[1,3]dioxol-5-ylmethyl-3S-(4-imidazol-1-yl-phenoxy)-piperidine Complex | | Descriptor: | (3S)-1-(1,3-BENZODIOXOL-5-YLMETHYL)-3-[4-(1H-IMIDAZOL-1-YL)PHENOXY]PIPERIDINE, Nitric oxide synthase, inducible, ... | | Authors: | Adler, M, Whitlow, M. | | Deposit date: | 2007-02-04 | | Release date: | 2007-04-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Design, Synthesis, and Activity of 2-Imidazol-1-ylpyrimidine Derived Inducible Nitric Oxide Synthase Dimerization Inhibitors
J.Med.Chem., 50, 2007
|
|
2OT8
 
 | |
2OPQ
 
 | | Crystal Structure of L100I Mutant HIV-1 Reverse Transcriptase in Complex with GW420867X. | | Descriptor: | ISOPROPYL (2S)-2-ETHYL-7-FLUORO-3-OXO-3,4-DIHYDROQUINOXALINE-1(2H)-CARBOXYLATE, PHOSPHATE ION, Reverse transcriptase/ribonuclease H, ... | | Authors: | Ren, J, Nichols, C.E, Chamberlain, P.P, Weaver, K.L, Short, S.A, Chan, J.H, Kleim, J, Stammers, D.K. | | Deposit date: | 2007-01-30 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Relationship of Potency and Resilience to Drug Resistant Mutations for GW420867X Revealed by Crystal Structures of Inhibitor Complexes for Wild-Type, Leu100Ile, Lys101Glu, and Tyr188Cys Mutant HIV-1 Reverse Transcriptases.
J.Med.Chem., 50, 2007
|
|
2OUB
 
 | | Crystal structure of the complex formed between phospholipase A2 and atenolol at 2.75 A resolution | | Descriptor: | 2-(4-(2-HYDROXY-3-(ISOPROPYLAMINO)PROPOXY)PHENYL)ETHANAMIDE, Phospholipase A2 VRV-PL-VIIIa | | Authors: | Kumar, S, Singh, N, Sharma, S, Kaur, P, Singh, T.P. | | Deposit date: | 2007-02-10 | | Release date: | 2007-02-27 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Crystal structure of the complex formed between phospholipase A2 and atenolol at 2.75 A resolution
To be Published
|
|
2OVO
 
 | |
2OW0
 
 | | MMP-9 active site mutant with iodine-labeled carboxylate inhibitor | | Descriptor: | CALCIUM ION, CHLORIDE ION, Matrix metalloproteinase-9 (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB), ... | | Authors: | Tochowicz, A, Bode, W, Maskos, K, Goettig, P. | | Deposit date: | 2007-02-15 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of MMP-9 Complexes with Five Inhibitors: Contribution of the Flexible Arg424 Side-chain to Selectivity.
J.Mol.Biol., 371, 2007
|
|
2OTF
 
 | | Crystal structure of the complex formed between phospholipase A2 and atenolol at 1.95 A resolution | | Descriptor: | 2-(4-(2-HYDROXY-3-(ISOPROPYLAMINO)PROPOXY)PHENYL)ETHANAMIDE, Phospholipase A2 VRV-PL-VIIIa | | Authors: | Kumar, S, Singh, N, Sharma, S, Bhushan, A, Kaur, P, Singh, T.P. | | Deposit date: | 2007-02-08 | | Release date: | 2007-02-20 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of the complex formed between phospholipase A2 and atenolol at 1.95 A resolution
To be Published
|
|
2OW2
 
 | | MMP-9 active site mutant with difluoro butanoic acid inhibitor | | Descriptor: | (3R)-4,4-DIFLUORO-3-[(4-METHOXYPHENYL)SULFONYL]BUTANOIC ACID, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Tochowicz, A, Bode, W, Maskos, K, Goettig, P. | | Deposit date: | 2007-02-15 | | Release date: | 2007-06-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal Structures of MMP-9 Complexes with Five Inhibitors: Contribution of the Flexible Arg424 Side-chain to Selectivity.
J.Mol.Biol., 371, 2007
|
|
2R4L
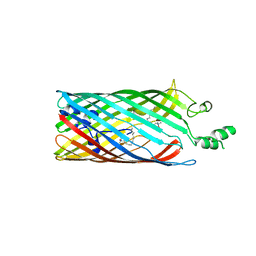 
 | | Crystal structure of the long-chain fatty acid transporter FadL mutant P34A | | Descriptor: | LAURYL DIMETHYLAMINE-N-OXIDE, Long-chain fatty acid transport protein | | Authors: | Hearn, E.M, Patel, D.R, Lepore, B.W, Indic, M, van den Berg, B. | | Deposit date: | 2007-08-31 | | Release date: | 2008-09-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Transmembrane passage of hydrophobic compounds through a protein channel wall
Nature, 458, 2009
|
|
2QXH
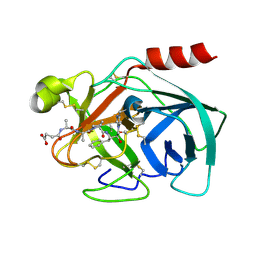 
 | | Crystal Structure of Human Kallikrein 7 in Complex with Suc-Ala-Ala-Pro-Phe-chloromethylketone | | Descriptor: | Kallikrein-7, N-(3-carboxypropanoyl)-L-alanyl-L-alanyl-N-[(2S,3S)-4-chloro-3-hydroxy-1-phenylbutan-2-yl]-L-prolinamide | | Authors: | Debela, M, Hess, P, Magdolen, V, Schechter, N.M, Bode, W, Goettig, P. | | Deposit date: | 2007-08-11 | | Release date: | 2008-01-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Chymotryptic specificity determinants in the 1.0 A structure of the zinc-inhibited human tissue kallikrein 7.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2QE2
 
 | | Structure of HCV NS5B Bound to an Anthranilic Acid Inhibitor | | Descriptor: | 2-{[N-(2-ACETYL-5-CHLORO-4-FLUOROPHENYL)GLYCYL]AMINO}BENZOIC ACID, RNA-directed RNA polymerase | | Authors: | Chopra, R, Svenson, K, Bard, J. | | Deposit date: | 2007-06-22 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Identification of Anthranilic Acid Derivatives as a Novel Class of Allosteric Inhibitors of Hepatitis C NS5B Polymerase
J.Med.Chem., 50, 2007
|
|
2QF0
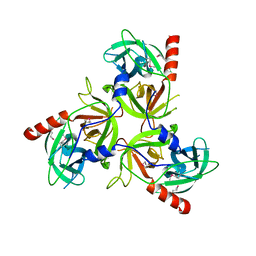 
 | |
2QGO
 
 | |
2QGS
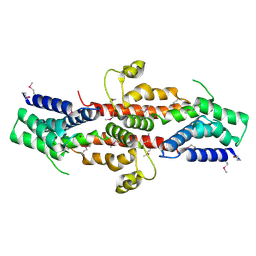 
 | | Crystal structure of SE1688 protein from Staphylococcus epidermidis. Northeast Structural Genomics Consortium target SeR89 | | Descriptor: | Protein SE1688, ZINC ION | | Authors: | Forouhar, F, Su, M, Benach, J, Seetharaman, J, Chen, C.X, Fang, Y, Cunningham, K, Owen, L, Ma, L.-C, Xiao, R, Liu, J, Baran, M.C, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-06-29 | | Release date: | 2007-07-17 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of SE1688 protein from Staphylococcus epidermidis.
To be Published
|
|
2QNC
 
 | | Crystal structure of T4 Endonuclease VII N62D mutant in complex with a DNA Holliday junction | | Descriptor: | 1,2-ETHANEDIOL, DNA (5'-D(*DAP*DGP*DGP*DCP*DCP*DTP*DAP*DGP*DCP*DGP*DTP*DCP*DCP*DGP*DGP*DAP*DAP*DTP*DTP*DCP*DTP*DTP*DCP*DG)-3'), DNA (5'-D(*DCP*DAP*DCP*DAP*DTP*DCP*DGP*DAP*DTP*DGP*DGP*DAP*DGP*DCP*DCP*DGP*DCP*DTP*DAP*DGP*DGP*DCP*DCP*DT)-3'), ... | | Authors: | Biertumpfel, C, Yang, W, Suck, D. | | Deposit date: | 2007-07-18 | | Release date: | 2008-01-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of T4 endonuclease VII resolving a Holliday junction.
Nature, 449, 2007
|
|
2NNQ
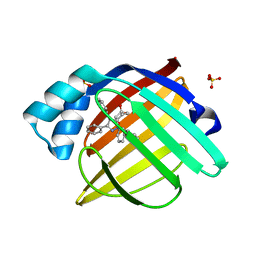 
 | |
2NM2
 
 | |
