6KR4
 
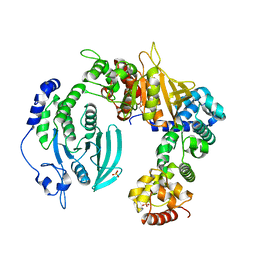 | | Crystal structure of the liprin-alpha3_SAM123/LAR_D1D2 complex | | Descriptor: | HEXAETHYLENE GLYCOL, Liprin-alpha-3, PHOSPHATE ION, ... | | Authors: | Xie, X, Liang, M, Luo, L, Wei, Z. | | Deposit date: | 2019-08-20 | | Release date: | 2020-01-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural basis of liprin-alpha-promoted LAR-RPTP clustering for modulation of phosphatase activity.
Nat Commun, 11, 2020
|
|
3FL5
 
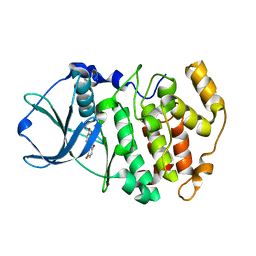 | | Protein kinase CK2 in complex with the inhibitor Quinalizarin | | Descriptor: | 1,2,5,8-tetrahydroxyanthracene-9,10-dione, Casein kinase II subunit alpha, DI(HYDROXYETHYL)ETHER | | Authors: | Mazzorana, M, Franchin, C, Battistutta, R. | | Deposit date: | 2008-12-18 | | Release date: | 2009-08-18 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Quinalizarin as a potent, selective and cell-permeable inhibitor of protein kinase CK2
Biochem.J., 421, 2009
|
|
6L54
 
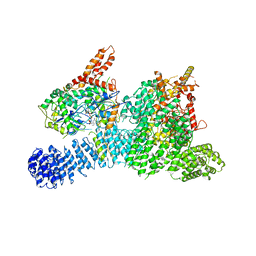 | | Structure of SMG189 | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Protein SMG8, ... | | Authors: | Xu, Y, Qi, Y. | | Deposit date: | 2019-10-22 | | Release date: | 2020-04-15 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.43 Å) | | Cite: | Cryo-EM structure of SMG1-SMG8-SMG9 complex.
Cell Res., 29, 2019
|
|
3GD2
 
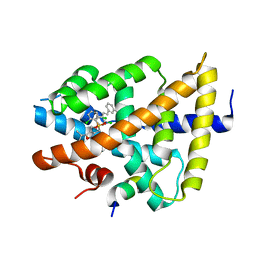 | | isoxazole ligand bound to farnesoid X receptor (FXR) | | Descriptor: | 3-[(E)-2-(2-chloro-4-{[3-{[(R)-(2,6-dichlorophenyl)(hydroxy)-lambda~4~-sulfanyl]methyl}-5-(1-methylethyl)isoxazol-4-yl]methoxy}phenyl)ethenyl]benzoic acid, Bile acid receptor, activator peptide | | Authors: | Madauss, K.P, Williams, S.P, Deaton, D.N, Wisely, G.B, Mcfadyen, R.B. | | Deposit date: | 2009-02-23 | | Release date: | 2009-07-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Substituted isoxazole analogs of farnesoid X receptor (FXR) agonist GW4064.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3GDH
 
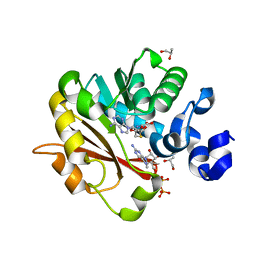 | | Methyltransferase domain of human Trimethylguanosine Synthase 1 (TGS1) bound to m7GTP and adenosyl-homocysteine (active form) | | Descriptor: | 7-METHYL-GUANOSINE-5'-TRIPHOSPHATE, R-1,2-PROPANEDIOL, S-1,2-PROPANEDIOL, ... | | Authors: | Monecke, T, Dickmanns, A, Ficner, R. | | Deposit date: | 2009-02-24 | | Release date: | 2009-06-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for m7G-cap hypermethylation of small nuclear, small nucleolar and telomerase RNA by the dimethyltransferase TGS1.
Nucleic Acids Res., 37, 2009
|
|
5MJY
 
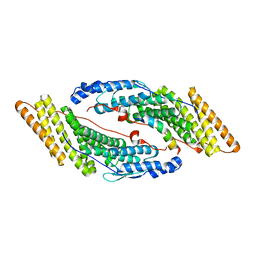 | |
6L53
 
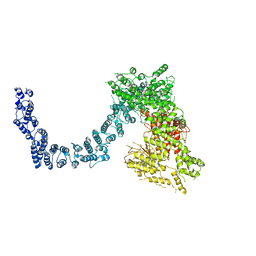 | | Structure of SMG1 | | Descriptor: | Serine/threonine-protein kinase SMG1 | | Authors: | Xu, Y, Qi, Y. | | Deposit date: | 2019-10-22 | | Release date: | 2020-04-15 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.63 Å) | | Cite: | Cryo-EM structure of SMG1-SMG8-SMG9 complex.
Cell Res., 29, 2019
|
|
3VOQ
 
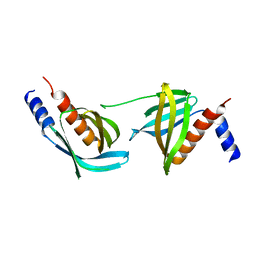 | |
3TAC
 
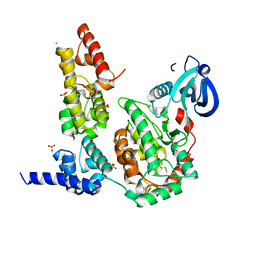 | | Crystal Structure of the Liprin-alpha/CASK complex | | Descriptor: | CHLORIDE ION, Liprin-alpha-2, Peripheral plasma membrane protein CASK, ... | | Authors: | Wei, Z, Zheng, S, Yu, C, Zhang, M. | | Deposit date: | 2011-08-03 | | Release date: | 2011-10-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Liprin-mediated large signaling complex organization revealed by the liprin-alpha/CASK and liprin-alpha/liprin-beta complex structures
Mol.Cell, 43, 2011
|
|
6L99
 
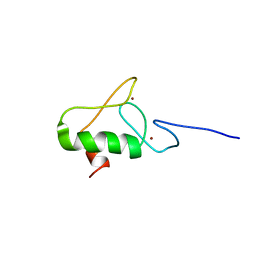 | | Zinc finger of RING finger protein 144A | | Descriptor: | E3 ubiquitin-protein ligase RNF144A, ZINC ION | | Authors: | Miyamoto, K. | | Deposit date: | 2019-11-08 | | Release date: | 2020-11-11 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Zinc finger of RING finger protein 144A
To Be Published
|
|
6I7L
 
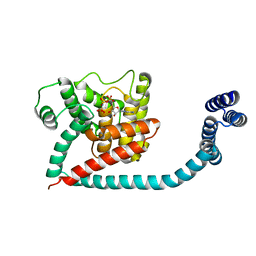 | | Crystal structure of monomeric FICD mutant L258D complexed with MgAMP-PNP | | Descriptor: | Adenosine monophosphate-protein transferase FICD, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
6I7G
 
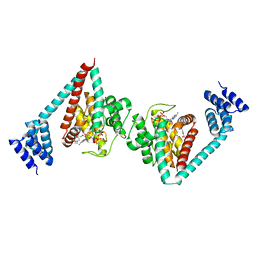 | | Crystal structure of dimeric wild type FICD complexed with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Adenosine monophosphate-protein transferase FICD, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Perera, L.A, Yan, Y, Read, R.J, Ron, D. | | Deposit date: | 2018-11-16 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | An oligomeric state-dependent switch in the ER enzyme FICD regulates AMPylation and deAMPylation of BiP.
Embo J., 38, 2019
|
|
3UWT
 
 | |
2GHQ
 
 | |
6EON
 
 | | Galactanase BT0290 | | Descriptor: | Beta-galactosidase, CALCIUM ION, alpha-D-galactopyranose | | Authors: | Basle, A, Munoz, J, Gilbert, H. | | Deposit date: | 2017-10-10 | | Release date: | 2017-11-29 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | A surface endogalactanase in Bacteroides thetaiotaomicron confers keystone status for arabinogalactan degradation.
Nat Microbiol, 3, 2018
|
|
6KIP
 
 | |
3US5
 
 | |
6LUR
 
 | |
3AL3
 
 | | Crystal Structure of TopBP1 BRCT7/8-BACH1 peptide complex | | Descriptor: | DNA topoisomerase 2-binding protein 1, FORMIC ACID, Peptide of Fanconi anemia group J protein | | Authors: | Leung, C.C, Glover, J.N. | | Deposit date: | 2010-07-22 | | Release date: | 2010-12-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Molecular basis of BACH1/FANCJ recognition by TopBP1 in DNA replication checkpoint control
J.Biol.Chem., 286, 2011
|
|
2YPD
 
 | | Crystal structure of the Jumonji domain of human Jumonji domain containing 1C protein | | Descriptor: | 2-(ETHYLMERCURI-THIO)-BENZOIC ACID, PROBABLE JMJC DOMAIN-CONTAINING HISTONE DEMETHYLATION PROT EIN 2C, SODIUM ION | | Authors: | Vollmar, M, Johansson, C, Krojer, T, Berridge, G, Burgess-Brown, N, Strain-Damerell, C, Froese, S, Williams, E, Goubin, S, Coutandin, D, von Delft, F, Arrowsmith, C.H, Bountra, C, Edwards, A, Oppermann, U. | | Deposit date: | 2012-10-30 | | Release date: | 2012-12-12 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of the Jumonji Domain of Human Jumonji Domain Containing 1C Protein
To be Published
|
|
6EUI
 
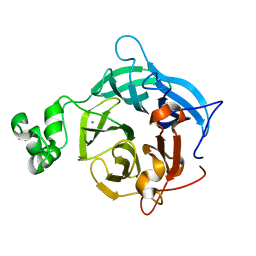 | | The GH43, Beta 1,3 Galactosidase, BT3683 with galactose | | Descriptor: | Beta-glucanase, CALCIUM ION, beta-D-galactopyranose | | Authors: | Cartmell, A, Gilbert, H.J. | | Deposit date: | 2017-10-30 | | Release date: | 2018-10-17 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | A surface endogalactanase in Bacteroides thetaiotaomicron confers keystone status for arabinogalactan degradation.
Nat Microbiol, 3, 2018
|
|
8XT3
 
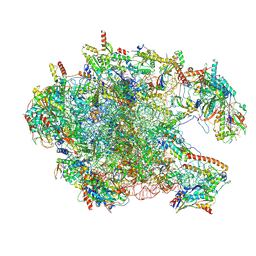 | | Cryo-EM structure of the human 39S mitoribosome with 10uM Tigecycline | | Descriptor: | 16s rRNA, 39S ribosomal protein L22, mitochondrial, ... | | Authors: | Li, X, Wang, M, Cheng, J. | | Deposit date: | 2024-01-10 | | Release date: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural basis for differential inhibition of eukaryotic ribosomes by tigecycline.
Nat Commun, 15, 2024
|
|
3HC6
 
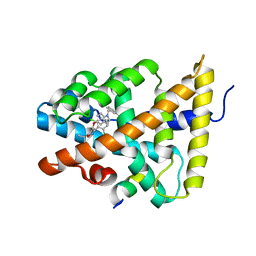 | | FXR with SRC1 and GSK088 | | Descriptor: | 3-[(5-{[3-(2,6-dichlorophenyl)-5-(1-methylethyl)isoxazol-4-yl]methoxy}-1H-indol-1-yl)methyl]benzoic acid, Bile acid receptor, Nuclear receptor coactivator 1, ... | | Authors: | Williams, S.P, Madauss, K.P. | | Deposit date: | 2009-05-05 | | Release date: | 2009-07-21 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | FXR agonist activity of conformationally constrained analogs of GW 4064.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3B0A
 
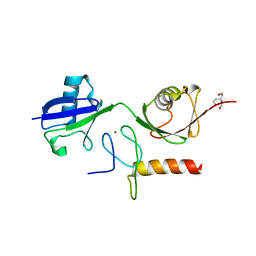 | | Crystal structure of the mouse HOIL1-L-NZF in complex with linear di-ubiquitin | | Descriptor: | Polyubiquitin-C, RanBP-type and C3HC4-type zinc finger-containing protein 1, TRIS(HYDROXYETHYL)AMINOMETHANE, ... | | Authors: | Sato, Y, Fujita, H, Yoshikawa, A, Yamashita, M, Yamagata, A, Kaiser, S.E, Iwai, K, Fukai, S. | | Deposit date: | 2011-06-07 | | Release date: | 2011-12-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Specific recognition of linear ubiquitin chains by the Npl4 zinc finger (NZF) domain of the HOIL-1L subunit of the linear ubiquitin chain assembly complex
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
6EUH
 
 | | The GH43, Beta 1,3 Galactosidase, BT3683 with galactodeoxynojirimycin | | Descriptor: | (2R,3S,4R,5S)-2-(hydroxymethyl)piperidine-3,4,5-triol, Beta-glucanase, CALCIUM ION | | Authors: | Cartmell, A, Gilbert, H.J. | | Deposit date: | 2017-10-30 | | Release date: | 2018-10-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A surface endogalactanase in Bacteroides thetaiotaomicron confers keystone status for arabinogalactan degradation.
Nat Microbiol, 3, 2018
|
|
