4L4M
 
 | | Structural Analysis of a Phosphoribosylated Inhibitor in Complex with Human Nicotinamide Phosphoribosyltransferase | | Descriptor: | 1,2-ETHANEDIOL, N-{4-[(3,5-difluorophenyl)sulfonyl]benzyl}imidazo[1,2-a]pyridine-7-carboxamide, Nicotinamide phosphoribosyltransferase, ... | | Authors: | Oh, A, Ho, Y, Zak, M, Liu, Y, Yuen, P, Zheng, X, Dragovich, S.P, Wang, W. | | Deposit date: | 2013-06-08 | | Release date: | 2014-06-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.445 Å) | | Cite: | Structural and biochemical analyses of the catalysis and potency impact of inhibitor phosphoribosylation by human nicotinamide phosphoribosyltransferase.
Chembiochem, 15, 2014
|
|
4O12
 
 | |
4L4L
 
 | | Structural Analysis of a Phosphoribosylated Inhibitor in Complex with Human Nicotinamide Phosphoribosyltransferase | | Descriptor: | 1,2-ETHANEDIOL, 6-({4-[(3,5-difluorophenyl)sulfonyl]benzyl}carbamoyl)-1-(5-O-phosphono-beta-D-ribofuranosyl)imidazo[1,2-a]pyridin-1-ium, Nicotinamide phosphoribosyltransferase, ... | | Authors: | Oh, A, Ho, Y, Zak, M, Liu, Y, Yuen, P, Zheng, X, Dragovich, S.P, Wang, W. | | Deposit date: | 2013-06-08 | | Release date: | 2014-06-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.122 Å) | | Cite: | Structural and biochemical analyses of the catalysis and potency impact of inhibitor phosphoribosylation by human nicotinamide phosphoribosyltransferase.
Chembiochem, 15, 2014
|
|
3BFT
 
 | | Structure of the ligand-binding core of GluR2 in complex with the agonist (S)-TDPA at 2.25 A resolution | | Descriptor: | (2S)-2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propanoic acid, CACODYLATE ION, CHLORIDE ION, ... | | Authors: | Beich-Frandsen, M, Mirza, O, Vestergaard, B, Gajhede, M, Kastrup, J.S. | | Deposit date: | 2007-11-23 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Structures of the ligand-binding core of iGluR2 in complex with the agonists (R)- and (S)-2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propionic acid explain their unusual equipotency.
J.Med.Chem., 51, 2008
|
|
1XX2
 
 | | Refinement of P99 beta-lactamase from Enterobacter cloacae | | Descriptor: | Beta-lactamase | | Authors: | Knox, J.R, Sun, T. | | Deposit date: | 2004-11-03 | | Release date: | 2004-11-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystallographic structure of a phosphonate derivative of the Enterobacter cloacae P99 cephalosporinase: mechanistic interpretation of a beta-lactamase transition-state analog.
Biochemistry, 33, 1994
|
|
3BFU
 
 | | Structure of the ligand-binding core of GluR2 in complex with the agonist (R)-TDPA at 1.95 A resolution | | Descriptor: | (2R)-2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propanoic acid, Glutamate receptor 2 | | Authors: | Beich-Frandsen, M, Mirza, O, Vestergaard, B, Gajhede, M, Kastrup, J.S. | | Deposit date: | 2007-11-23 | | Release date: | 2008-10-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structures of the ligand-binding core of iGluR2 in complex with the agonists (R)- and (S)-2-amino-3-(4-hydroxy-1,2,5-thiadiazol-3-yl)propionic acid explain their unusual equipotency.
J.Med.Chem., 51, 2008
|
|
3CP4
 
 | |
2ZAX
 
 | | Crystal Structure of Ferric Cytochrome P450cam | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CAMPHOR, Cytochrome P450-cam, ... | | Authors: | Sakurai, K, Shimada, H, Harada, K, Hayashi, T, Tsukihara, T. | | Deposit date: | 2007-10-11 | | Release date: | 2008-01-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Evaluation of the functional role of the heme-6-propionate side chain in cytochrome P450cam
J.Am.Chem.Soc., 130, 2008
|
|
2ZAW
 
 | | Crystal Structure of Ferric Cytochrome P450cam Reconstituted with 6-Methyl-6-depropionated Hemin | | Descriptor: | 6-METHY-6-DEPROPIONATEHEMIN, CAMPHOR, CHLORIDE ION, ... | | Authors: | Harada, K, Sakurai, K, Shimada, H, Tsukihara, T, Hayashi, T. | | Deposit date: | 2007-10-11 | | Release date: | 2008-01-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Evaluation of the functional role of the heme-6-propionate side chain in cytochrome P450cam
J.Am.Chem.Soc., 130, 2008
|
|
4OLG
 
 | | Crystal structure of AmpC beta-lactamase in complex with covalently bound N-formyl 7-aminocephalosporanic acid | | Descriptor: | (2R,5Z)-5-[(acetyloxy)methylidene]-2-[(1R)-1-(formylamino)-2-oxoethyl]-5,6-dihydro-2H-1,3-thiazine-4-carboxylic acid, Beta-lactamase, PHOSPHATE ION | | Authors: | Shoichet, B.K, Barelier, S. | | Deposit date: | 2014-01-23 | | Release date: | 2014-05-28 | | Last modified: | 2014-06-18 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Substrate deconstruction and the nonadditivity of enzyme recognition.
J.Am.Chem.Soc., 136, 2014
|
|
4OLD
 
 | | Crystal structure of AmpC beta-lactamase in complex with the product form of (6R,7R)-7-amino-8-oxo-5-thia-1-azabicyclo[4.2.0]oct-2-ene-2-carboxylic acid | | Descriptor: | (2R)-2-[(R)-amino(carboxy)methyl]-3,6-dihydro-2H-1,3-thiazine-4-carboxylic acid, Beta-lactamase, PHOSPHATE ION | | Authors: | Shoichet, B.K, Barelier, S. | | Deposit date: | 2014-01-23 | | Release date: | 2014-05-28 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Substrate deconstruction and the nonadditivity of enzyme recognition.
J.Am.Chem.Soc., 136, 2014
|
|
4XUX
 
 | | Structure of ampC bound to RPX-7009 at 1.75 A | | Descriptor: | 1,2-ETHANEDIOL, 2-(2-METHOXYETHOXY)ETHANOL, Beta-lactamase, ... | | Authors: | Clifton, M.C, Abendroth, J. | | Deposit date: | 2015-01-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Discovery of a Cyclic Boronic Acid beta-Lactamase Inhibitor (RPX7009) with Utility vs Class A Serine Carbapenemases.
J.Med.Chem., 58, 2015
|
|
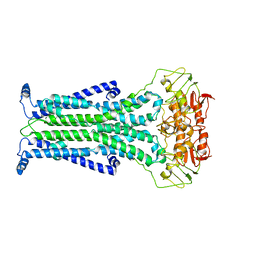
5C73
 
 | |
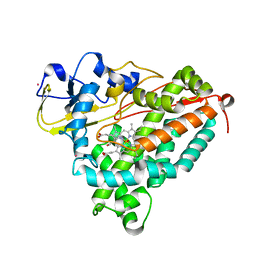
2A1M
 
 | |
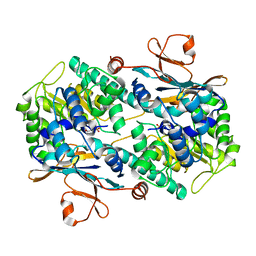
2G96
 
 | |
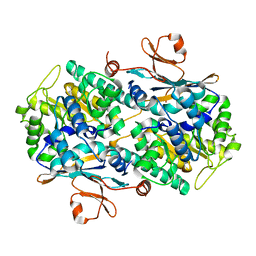
2G95
 
 | |
4OKP
 
 | |
2A1O
 
 | |
2G97
 | |
2A1N
 
 | |
2X1Z
 
 | | Structure of Peridinin-Chlorophyll-Protein reconstituted with Chl-d | | Descriptor: | CADMIUM ION, CHLORIDE ION, CHLOROPHYLL D, ... | | Authors: | Schulte, T, Hiller, R.G, Hofmann, E. | | Deposit date: | 2010-01-09 | | Release date: | 2010-02-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-Ray Structures of the Peridinin-Chlorophyll-Protein Reconstituted with Different Chlorophylls.
FEBS Lett., 584, 2010
|
|
2FEU
 
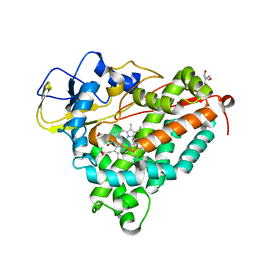 | | P450CAM from Pseudomonas putida reconstituted with manganic protoporphyrin IX | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CAMPHOR, Cytochrome P450-cam, ... | | Authors: | von Koenig, K, Makris, T.M, Sligar, S.G, Schlichting, I. | | Deposit date: | 2005-12-16 | | Release date: | 2006-03-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The status of high-valent metal oxo complexes in the P450 cytochromes.
J.Inorg.Biochem., 100, 2006
|
|
2X21
 
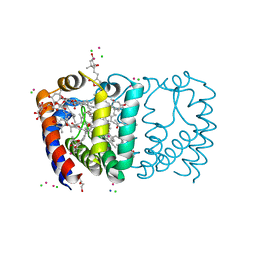 | | Structure of Peridinin-Chlorophyll-Protein reconstituted with BChl-a | | Descriptor: | BACTERIOCHLOROPHYLL A, CADMIUM ION, CHLORIDE ION, ... | | Authors: | Schulte, T, Hiller, R.G, Hofmann, E. | | Deposit date: | 2010-01-09 | | Release date: | 2010-02-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | X-Ray Structures of the Peridinin-Chlorophyll-Protein Reconstituted with Different Chlorophylls.
FEBS Lett., 584, 2010
|
|
2X20
 
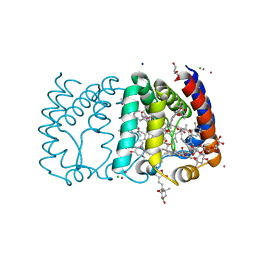 | | Structure of Peridinin-Chlorophyll-Protein reconstituted with Chl-b | | Descriptor: | CADMIUM ION, CHLORIDE ION, CHLOROPHYLL B, ... | | Authors: | Schulte, T, Hiller, R.G, Hofmann, E. | | Deposit date: | 2010-01-09 | | Release date: | 2010-02-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | X-Ray Structures of the Peridinin-Chlorophyll-Protein Reconstituted with Different Chlorophylls.
FEBS Lett., 584, 2010
|
|
5ZOI
 
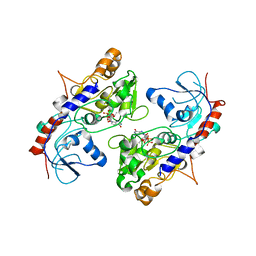 | | Crystal Structure of alpha1,3-Fucosyltransferase | | Descriptor: | Alpha-(1,3)-fucosyltransferase FucT, [[(2S,3R,4S,5R)-5-(2-azanyl-6-oxidanylidene-1H-purin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] [(2S,3S,4S,5R,6R)-6-methyl-3,4,5-tris(oxidanyl)oxan-2-yl] hydrogen phosphate | | Authors: | Tan, Y, Yang, G. | | Deposit date: | 2018-04-13 | | Release date: | 2019-06-26 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Directed evolution of an alpha 1,3-fucosyltransferase using a single-cell ultrahigh-throughput screening method.
Sci Adv, 5, 2019
|
|
