3NYJ
 
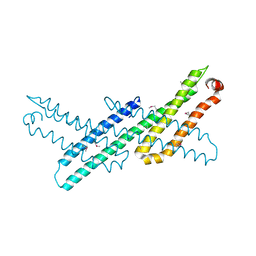 | | Crystal Structure Analysis of APP E2 domain | | Descriptor: | Amyloid beta A4 protein, OSMIUM ION | | Authors: | Ha, Y, Hu, J, Lee, S, Liu, X. | | Deposit date: | 2010-07-15 | | Release date: | 2011-06-01 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The E2 Domains of APP and APLP1 Share a Conserved Mode of Dimerization.
Biochemistry, 50, 2011
|
|
4QCE
 
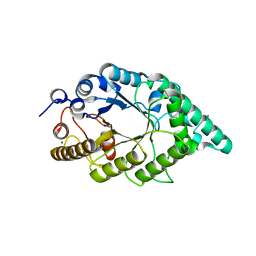 | | Crystal structure of recombinant alkali thermostable GH10 xylanase from Bacillus sp. NG-27 | | Descriptor: | Alkaline thermostable endoxylanase, MAGNESIUM ION, SODIUM ION | | Authors: | Mahanta, P, Bhardwaj, A, Reddy, V.S, Ramakumar, S. | | Deposit date: | 2014-05-11 | | Release date: | 2015-05-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Structural insights into N-terminal to C-terminal interactions and implications for thermostability of a (beta/alpha)8-triosephosphate isomerase barrel enzyme
Febs J., 282, 2015
|
|
6BDJ
 
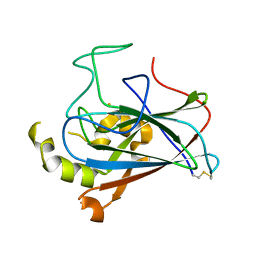 | | Crystal structure of dioxygenase Tetur07g02040 | | Descriptor: | FE (III) ION, Tetur07g02040 | | Authors: | Daneshian, L, Schlachter, C.R, Chruszcz, M. | | Deposit date: | 2017-10-23 | | Release date: | 2018-11-14 | | Last modified: | 2019-05-29 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural and functional characterization of an intradiol ring-cleavage dioxygenase from the polyphagous spider mite herbivore Tetranychus urticae Koch.
Insect Biochem.Mol.Biol., 107, 2019
|
|
3W1K
 
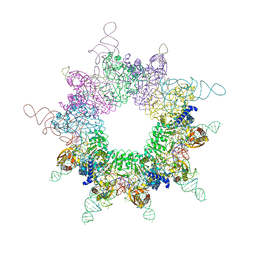 | |
1BC3
 
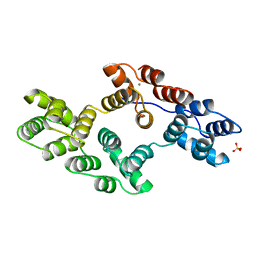 | | RECOMBINANT RAT ANNEXIN V, TRIPLE MUTANT (T72K, S144K, S228K) | | Descriptor: | ANNEXIN V, CALCIUM ION, SULFATE ION | | Authors: | Mo, Y.D, Swairjo, M.A, Li, C.W, Head, J.F, Seaton, B.A. | | Deposit date: | 1998-05-04 | | Release date: | 1998-11-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Mutational and crystallographic analyses of interfacial residues in annexin V suggest direct interactions with phospholipid membrane components.
Biochemistry, 37, 1998
|
|
6AWA
 
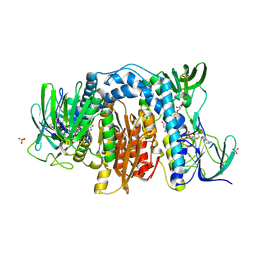 | | 1.83 Angstrom Resolution Crystal Structure of Dihydrolipoyl Dehydrogenase from Pseudomonas putida in Complex with FAD and Adenosine-5'-monophosphate. | | Descriptor: | ADENOSINE MONOPHOSPHATE, Dihydrolipoyl dehydrogenase, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Minasov, G, Shuvalova, L, Kiryukhina, O, Dubrovska, I, Grimshaw, S, Kwon, K, Anderson, W.F, Satchell, K.J.F, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2017-09-05 | | Release date: | 2017-10-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | 1.83 Angstrom Resolution Crystal Structure of Dihydrolipoyl Dehydrogenase from Pseudomonas putida in Complex with FAD and Adenosine-5'-monophosphate.
To Be Published
|
|
1IFS
 
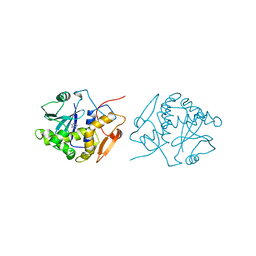 | | RICIN A-CHAIN (RECOMBINANT) COMPLEX WITH ADENOSINE (ADENOSINE BECOMES ADENINE IN THE COMPLEX) | | Descriptor: | ADENINE, RICIN | | Authors: | Weston, S.A, Tucker, A.D, Thatcher, D.R, Derbyshire, D.J, Pauptit, R.A. | | Deposit date: | 1996-07-05 | | Release date: | 1998-01-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | X-ray structure of recombinant ricin A-chain at 1.8 A resolution.
J.Mol.Biol., 244, 1994
|
|
5LFV
 
 | |
6AX6
 
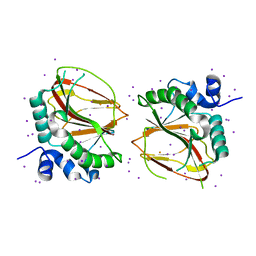 | | The crystal structure of a lysyl hydroxylase from Acanthamoeba polyphaga mimivirus | | Descriptor: | FE (II) ION, IODIDE ION, Procollagen lysyl hydroxylase and glycosyltransferase | | Authors: | Guo, H, Tsai, C, Miller, M.D, Alvarado, S, Tainer, J.A, Phillips Jr, G.N, Kurie, J.M. | | Deposit date: | 2017-09-06 | | Release date: | 2018-02-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.241 Å) | | Cite: | Pro-metastatic collagen lysyl hydroxylase dimer assemblies stabilized by Fe2+-binding.
Nat Commun, 9, 2018
|
|
6BG5
 
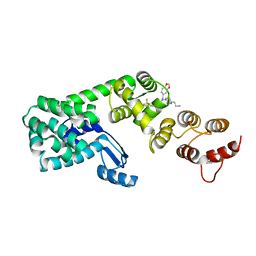 | | Structure of 1-(benzo[d][1,3]dioxol-5-ylmethyl)-1-(1-propylpiperidin-4-yl)-3-(3-(trifluoromethyl)phenyl)urea bound to DCN1 | | Descriptor: | Endolysin, DCN1-like protein 1 chimera, N-[(2H-1,3-benzodioxol-5-yl)methyl]-N-(1-propylpiperidin-4-yl)-N'-[3-(trifluoromethyl)phenyl]urea | | Authors: | Guy, R.K, Schulman, B.A, Scott, D.C, Hammill, J.T. | | Deposit date: | 2017-10-27 | | Release date: | 2018-09-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Piperidinyl Ureas Chemically Control Defective in Cullin Neddylation 1 (DCN1)-Mediated Cullin Neddylation.
J. Med. Chem., 61, 2018
|
|
5LJF
 
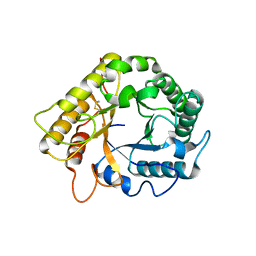 | | Crystal structure of the endo-1,4-glucanase RBcel1 E135A with cellotriose | | Descriptor: | Endoglucanase, beta-D-glucopyranose-(1-4)-beta-D-glucopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Dutoit, R, Collet, L, Galleni, M, Bauvois, C. | | Deposit date: | 2016-07-18 | | Release date: | 2017-08-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.734396 Å) | | Cite: | Glycoside hydrolase family 5: structural snapshots highlighting the involvement of two conserved residues in catalysis.
Acta Crystallogr D Struct Biol, 77, 2021
|
|
5LJX
 
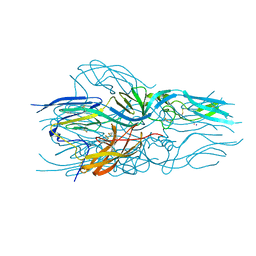 | |
4PXH
 
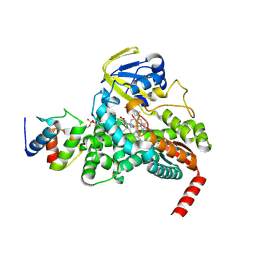 | |
4HP2
 
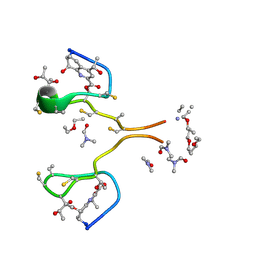 | | Invariom refinement of a new dimeric monoclinic 2 solvate of thiostrepton at 0.64 angstrom resolution | | Descriptor: | DIMETHYLFORMAMIDE, Thiostrepton, diethyl ether | | Authors: | Proepper, K, Holstein, J.J, Huebschle, C.B, Bond, C.S, Dittrich, B. | | Deposit date: | 2012-10-23 | | Release date: | 2013-10-02 | | Last modified: | 2018-04-04 | | Method: | X-RAY DIFFRACTION (0.64 Å) | | Cite: | Invariom refinement of a new monoclinic solvate of thiostrepton at 0.64 angstrom resolution.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
8JBZ
 
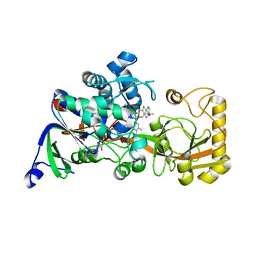 | | Crystal structure of 3-ketosteroid delta1-dehydrogenase from Rhodococcus erythropolis SQ1 in complex with 4-androstadiene-3,17- dione | | Descriptor: | 3-ketosteroid dehydrogenase, 4-ANDROSTENE-3-17-DIONE, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Hu, Y.L, Li, X, Cheng, X.Y, Song, S.K, Su, Z.D. | | Deposit date: | 2023-05-10 | | Release date: | 2023-05-31 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.079 Å) | | Cite: | Crystal structure of 3-ketosteroid delta1-dehydrogenase from Rhodococcus erythropolis SQ1 in complex with 4-androstadiene-3,17- dione
To Be Published
|
|
1BCI
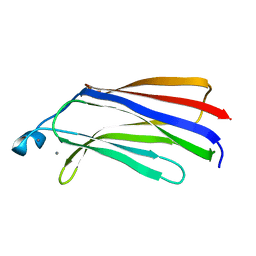 
 | | C2 DOMAIN OF CYTOSOLIC PHOSPHOLIPASE A2, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | CALCIUM ION, CYTOSOLIC PHOSPHOLIPASE A2 | | Authors: | Xu, G.Y, Mcdonagh, T, Yu, H.A, Nalefski, E.A, Clark, J.D, Cumming, D.A. | | Deposit date: | 1998-04-30 | | Release date: | 1998-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure and membrane interactions of the C2 domain of cytosolic phospholipase A2.
J.Mol.Biol., 280, 1998
|
|
4PZD
 
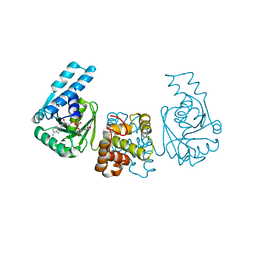 | |
8IVJ
 
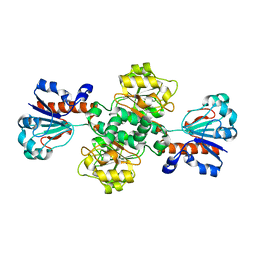 | |
8FSQ
 
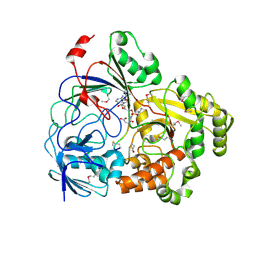 | | Complex Structure of YejA with Microcin C7 | | Descriptor: | 5'-O-[(R)-amino(3-aminopropoxy)phosphoryl]adenosine, Microcin C7 peptide portion, YejA | | Authors: | Naik, S.K, Dong, S.-H. | | Deposit date: | 2023-01-11 | | Release date: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Trojan Horse Peptide Conjugates Remodel the Activity Spectrum of Clinical Antibiotics
To Be Published
|
|
4Q2B
 
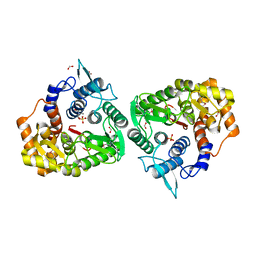 | | The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Endo-1,4-beta-D-glucanase, FORMIC ACID, ... | | Authors: | Tan, K, Joachimiak, G, Endres, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2014-04-07 | | Release date: | 2014-06-25 | | Last modified: | 2015-04-29 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | The crystal structure of an endo-1,4-D-glucanase from Pseudomonas putida KT2440
To be Published
|
|
8CZI
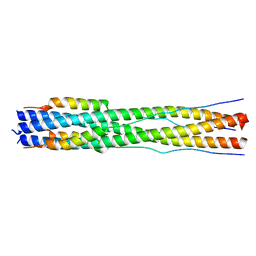 
 | |
5LGE
 
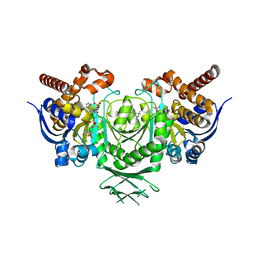 | | Crystal Structure of human IDH1 mutant (R132H) in complex with NADP+ and an Inhibitor related to BAY 1436032 | | Descriptor: | 1,2-ETHANEDIOL, 2-[(4-propan-2-ylphenyl)amino]-1-[(1~{S},5~{S})-3,3,5-trimethylcyclohexyl]benzimidazole-5-carboxylic acid, ACETATE ION, ... | | Authors: | Hillig, R.C, Hars, U, Korndoerfer, I.P. | | Deposit date: | 2016-07-07 | | Release date: | 2017-02-08 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Pan-mutant IDH1 inhibitor BAY 1436032 for effective treatment of IDH1 mutant astrocytoma in vivo.
Acta Neuropathol., 133, 2017
|
|
1AWJ
 
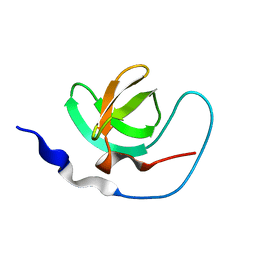 | | INTRAMOLECULAR ITK-PROLINE COMPLEX, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | ITK | | Authors: | Andreotti, A.H, Bunnell, S.C, Feng, S, Berg, L.J, Schreiber, S.L. | | Deposit date: | 1997-10-02 | | Release date: | 1998-01-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Regulatory intramolecular association in a tyrosine kinase of the Tec family.
Nature, 385, 1997
|
|
4Q3Q
 
 | | Crystal structure of Schistosoma mansoni arginase in complex with inhibitor ABH | | Descriptor: | 2(S)-AMINO-6-BORONOHEXANOIC ACID, Arginase, GLYCEROL, ... | | Authors: | Hai, Y, Edwards, J.E, Van Zandt, M.C, Hoffmann, K.F, Christianson, D.W. | | Deposit date: | 2014-04-12 | | Release date: | 2014-07-16 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Crystal Structure of Schistosoma mansoni Arginase, a Potential Drug Target for the Treatment of Schistosomiasis.
Biochemistry, 53, 2014
|
|
4Q3Z
 
 | |
