8VMN
 
 | | H3K4me3 nucleosome bound to PRC2_AJ1-450 | | Descriptor: | DNA (157-MER), Histone H2A, Histone H2B, ... | | Authors: | Cookis, T, Nogales, E. | | Deposit date: | 2024-01-13 | | Release date: | 2025-01-22 | | Last modified: | 2025-03-05 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural basis for the inhibition of PRC2 by active transcription histone posttranslational modifications.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8VLR
 
 | | Cryo-EM structure of native H2AK119bu nucleosome at 2.6 | | Descriptor: | DNA (136-MER), Histone H2A type 1-B/E, Histone H2B type 1-A, ... | | Authors: | Wang, Y, Zhang, K. | | Deposit date: | 2024-01-12 | | Release date: | 2025-01-15 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Role of histone variants H2BC1 and H2AZ.2 in H2AK119ub nucleosome organization and Polycomb gene silencing
To Be Published
|
|
8VO0
 
 | |
8VOB
 
 | | H3K36me3-modified nucleosome bound to PRC2_AJ1-450 | | Descriptor: | DNA (157-MER), Histone H2A type 1, Histone H2B 1.1, ... | | Authors: | Cookis, T, Nogales, E. | | Deposit date: | 2024-01-14 | | Release date: | 2025-01-15 | | Last modified: | 2025-03-05 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structural basis for the inhibition of PRC2 by active transcription histone posttranslational modifications.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8VVN
 
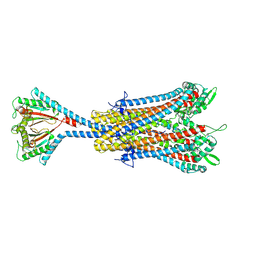 | |
4MQH
 
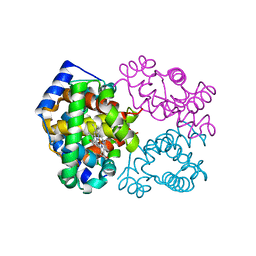 | |
3GXI
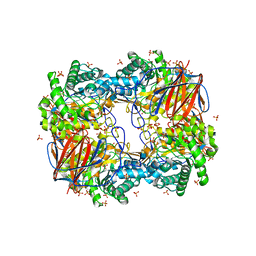 
 | | Crystal structure of acid-beta-glucosidase at pH 5.5 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Glucosylceramidase, PHOSPHATE ION | | Authors: | Lieberman, R.L. | | Deposit date: | 2009-04-02 | | Release date: | 2009-05-05 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Effects of pH and iminosugar pharmacological chaperones on lysosomal glycosidase structure and stability.
Biochemistry, 48, 2009
|
|
8VW9
 
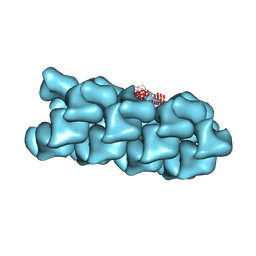 | | CryoEM Structure of a FtsH Helical Assembly in the Presence of ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, ATP-dependent zinc metalloprotease FtsH, MAGNESIUM ION, ... | | Authors: | Li, Y, Zhu, J, Zhang, Z, Wang, F, Egelman, E.H, Tezcan, F.A. | | Deposit date: | 2024-01-31 | | Release date: | 2025-01-15 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Transforming an ATP-dependent enzyme into a dissipative, self-assembling system.
Nat.Chem.Biol., 21, 2025
|
|
4MUV
 
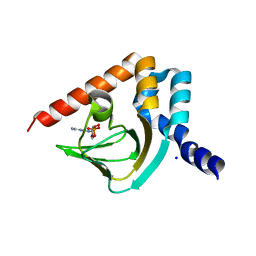 | | M. loti cyclic-nucleotide binding domain mutant displaying inverted ligand selectivity, cyclic-GMP bound | | Descriptor: | CYCLIC GUANOSINE MONOPHOSPHATE, Cyclic nucleotide-gated potassium channel mll3241, SODIUM ION | | Authors: | Fonseca, F, Pessoa, J, Morais-Cabral, J.H. | | Deposit date: | 2013-09-23 | | Release date: | 2014-06-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Determinants of ligand selectivity in a cyclic nucleotide-regulated potassium channel.
J.Gen.Physiol., 144, 2014
|
|
8VWC
 
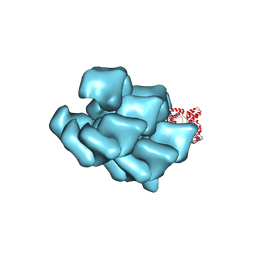 | | CryoEM Structure of a FtsH Helical Assembly in the Aged State | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, ATP-dependent zinc metalloprotease FtsH, MAGNESIUM ION, ... | | Authors: | Li, Y, Zhu, J, Zhang, Z, Wang, F, Egelman, E.H, Tezcan, F.A. | | Deposit date: | 2024-01-31 | | Release date: | 2025-01-15 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Transforming an ATP-dependent enzyme into a dissipative, self-assembling system.
Nat.Chem.Biol., 21, 2025
|
|
8VCF
 
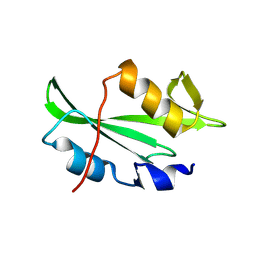 | |
8W3T
 
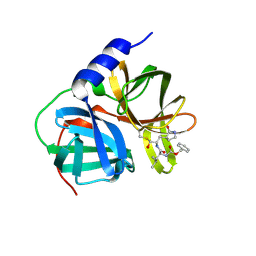 | |
4MXA
 
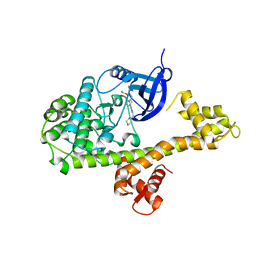 | | CDPK1 from Neospora caninum in complex with inhibitor RM-1-132 | | Descriptor: | 3-(6-ethoxynaphthalen-2-yl)-1-(piperidin-4-ylmethyl)-1H-pyrazolo[3,4-d]pyrimidin-4-amine, Calmodulin-like domain protein kinase isoenzyme gamma, related | | Authors: | Merritt, E.A. | | Deposit date: | 2013-09-26 | | Release date: | 2013-10-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Neospora caninum Calcium-Dependent Protein Kinase 1 Is an Effective Drug Target for Neosporosis Therapy.
Plos One, 9, 2014
|
|
8VS0
 
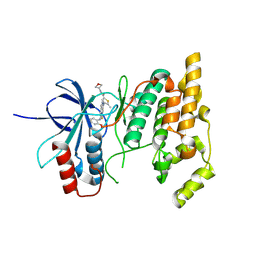 | |
8VVY
 
 | | Human Cullin-1 in complex with CAND2 | | Descriptor: | Cullin-1, Cullin-associated NEDD8-dissociated protein 2 | | Authors: | Kenny, S, Liu, X, Das, C. | | Deposit date: | 2024-01-31 | | Release date: | 2025-01-29 | | Last modified: | 2025-08-13 | | Method: | ELECTRON MICROSCOPY (3.49 Å) | | Cite: | Molecular mechanisms of CAND2 in regulating SCF ubiquitin ligases.
Nat Commun, 16, 2025
|
|
8VWY
 
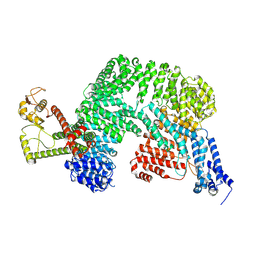
 | | Complex structure of mouse C1ql3 with BAI3 | | Descriptor: | Adhesion G protein-coupled receptor B3, CALCIUM ION, Complement C1q-like protein 3 | | Authors: | Miao, Y, Sudhof, T.C. | | Deposit date: | 2024-02-02 | | Release date: | 2025-02-05 | | Last modified: | 2025-08-20 | | Method: | ELECTRON MICROSCOPY (2.78 Å) | | Cite: | Structure of the complex of C1q-like 3 protein with adhesion-GPCR BAI3.
Commun Biol, 8, 2025
|
|
4V6B
 
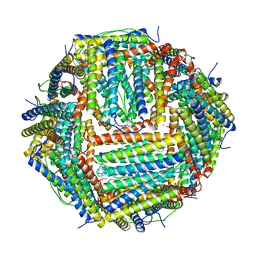 | | Crystal structure of human ferritin Phe167SerfsX26 mutant. | | Descriptor: | CALCIUM ION, Ferritin | | Authors: | Hurley, T.D, Vidal, R. | | Deposit date: | 2009-06-19 | | Release date: | 2014-07-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Unraveling of the E-helices and disruption of 4-fold pores are associated with iron mishandling in a mutant ferritin causing neurodegeneration
J.Biol.Chem., 285, 2010
|
|
8VO9
 
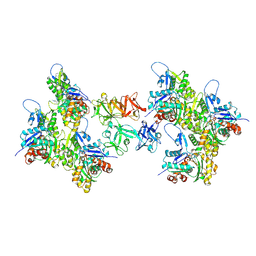 | | Cryo-EM structure of fascin crosslinked F-actin (Eigen_middle) | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin, alpha skeletal muscle, ... | | Authors: | Gong, R, Reynolds, M.J, Alushin, G.M. | | Deposit date: | 2024-01-14 | | Release date: | 2025-01-15 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Fascin structural plasticity mediates flexible actin bundle construction.
Nat.Struct.Mol.Biol., 32, 2025
|
|
4MT6
 
 | |
8VV5
 
 | |
8VW6
 
 | | Crystal structure of SsuD | | Descriptor: | Alkanesulfonate monooxygenase | | Authors: | Caputo, A.T, Hu, M, Scott, C. | | Deposit date: | 2024-01-31 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.606 Å) | | Cite: | Crystral structure of SsuD, a desulfonase protein from Rhodococcus jostii RHA
TBD
|
|
8VYB
 
 | | Cryo-EM structure of human core Rab3GAP1/2 complex | | Descriptor: | Isoform 2 of Rab3 GTPase-activating protein catalytic subunit, Rab3 GTPase-activating protein non-catalytic subunit | | Authors: | Nguyen, K.M, Yip, C.K. | | Deposit date: | 2024-02-07 | | Release date: | 2025-01-15 | | Method: | ELECTRON MICROSCOPY (3.37 Å) | | Cite: | Biochemical and structural characterization of Rab3GAP reveals insights into Rab18 nucleotide exchange activity.
Nat Commun, 16, 2025
|
|
8W2S
 
 | |
8VX3
 
 | | Structure of an exported small alarmone synthetase from Streptomyces albidoflavus | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, SODIUM ION, ... | | Authors: | Ahmad, S, Tsang, K.K, Schiefer, V, Kim, Y, Whitney, J.C. | | Deposit date: | 2024-02-03 | | Release date: | 2025-02-05 | | Last modified: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of a (p)ppApp synthetase from Streptomyces albidoflavus
To Be Published
|
|
4V6G
 
 | | Initiation complex of 70S ribosome with two tRNAs and mRNA. | | Descriptor: | 16S RRNA (E.COLI NUMBERING), 23S ribosomal RNA, 30S RIBOSOMAL PROTEIN S10, ... | | Authors: | Jenner, L.B, Yusupova, G, Yusupov, M. | | Deposit date: | 2009-07-10 | | Release date: | 2014-07-09 | | Last modified: | 2024-12-25 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural aspects of messenger RNA reading frame maintenance by the ribosome.
Nat.Struct.Mol.Biol., 17, 2010
|
|
