1NQC
 
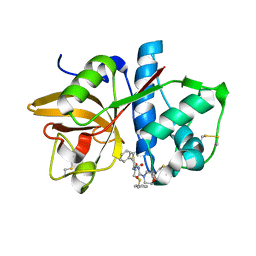 | | Crystal structures of Cathepsin S inhibitor complexes | | Descriptor: | Cathepsin S, N-[(1R)-2-(BENZYLSULFANYL)-1-FORMYLETHYL]-N-(MORPHOLIN-4-YLCARBONYL)-L-PHENYLALANINAMIDE | | Authors: | Pauly, T.A, Sulea, T, Ammirati, M, Sivaraman, J, Danley, D.E, Griffor, M.C, Kamath, A.V, Wang, I.K, Laird, E.R, Menard, R, Cygler, M, Rath, V.L. | | Deposit date: | 2003-01-21 | | Release date: | 2003-04-15 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Specificity determinants of human cathepsin s revealed
by crystal structures of complexes.
Biochemistry, 42, 2003
|
|
1NPV
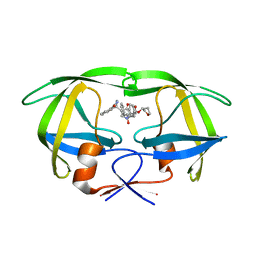 
 | | Crystal structure of HIV-1 protease complexed with LDC271 | | Descriptor: | POL polyprotein, {1-BENZYL-3-[2-BENZYL-3-OXO-4-(1-OXO-1,2,3,4-TETRAHYDRO- ISOQUINOLIN-4-YL)-2,3-DIHYDRO-1H-PYRROL-2-YL]-2- HYDROXY-PROPYL}-CARBAMIC ACID TETRAHYDRO-FURAN-3-YL ESTER | | Authors: | Smith, A.B. | | Deposit date: | 2003-01-20 | | Release date: | 2003-02-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design, synthesis, and biological evaluation of monopyrrolinone-based HIV-1 protease inhibitors.
J.Med.Chem., 46, 2003
|
|
1NQ2
 
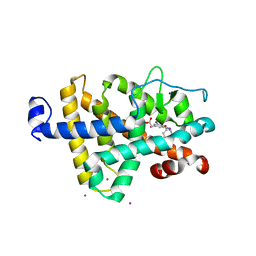 | | Two RTH Mutants with Impaired Hormone Binding | | Descriptor: | ARSENIC, SODIUM ION, Thyroid hormone receptor beta-1, ... | | Authors: | Huber, B.R, Sandler, B, West, B.L, Cunha-Lima, S.T, Nguyen, H.T, Apriletti, J.W, Baxter, J.D, Fletterick, R.J. | | Deposit date: | 2003-01-21 | | Release date: | 2003-04-15 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Two resistance to thyroid hormone mutants with impaired hormone binding
Mol.Endocrinol., 17, 2003
|
|
1NRV
 
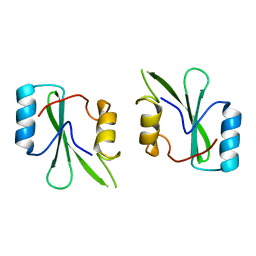 | | Crystal structure of the SH2 domain of Grb10 | | Descriptor: | Growth factor receptor-bound protein 10 | | Authors: | Stein, E.G, Hubbard, S.R. | | Deposit date: | 2003-01-25 | | Release date: | 2003-04-15 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural basis for dimerization of the Grb10 Src homology 2 domain. Implications for ligand specificity.
J.Biol.Chem., 278, 2003
|
|
1NU2
 
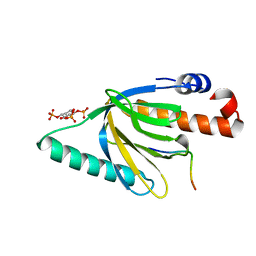 | | Crystal structure of the murine Disabled-1 (Dab1) PTB domain-ApoER2 peptide-PI-4,5P2 ternary complex | | Descriptor: | D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE, Disabled homolog 1, peptide derived from murine Apolipoprotein E Receptor-2 | | Authors: | Stolt, P.C, Jeon, H, Song, H.K, Herz, J, Eck, M.J, Blacklow, S.C. | | Deposit date: | 2003-01-30 | | Release date: | 2003-04-15 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Origins of Peptide Selectivity and Phosphoinositide Binding Revealed by Structures of Disabled-1 PTB Domain Complexes
Structure, 11, 2003
|
|
1PK2
 
 | |
1PR9
 
 | | Human L-Xylulose Reductase Holoenzyme | | Descriptor: | DIHYDROGENPHOSPHATE ION, L-XYLULOSE REDUCTASE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | El-Kabbani, O, Ishikura, S, Darmanin, C, Carbone, V, Chung, R.P.-T, Usami, N, Hara, A. | | Deposit date: | 2003-06-20 | | Release date: | 2004-02-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Crystal structure of human L-xylulose reductase holoenzyme: probing the role of Asn107 with site-directed mutagenesis
Proteins, 55, 2004
|
|
1PT7
 
 | | Crystal structure of the apo-form of the yfdW gene product of E. coli | | Descriptor: | GLYCEROL, Hypothetical protein yfdW, PHOSPHATE ION | | Authors: | Gruez, A, Roig-Zamboni, V, Valencia, C, Campanacci, V, Cambillau, C. | | Deposit date: | 2003-06-23 | | Release date: | 2003-09-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the Escherichia coli yfdW gene product reveals a New fold of two interlaced rings identifying a wide family of CoA transferases.
J.Biol.Chem., 278, 2003
|
|
1Q4X
 
 | | Crystal Structure of Human Thyroid Hormone Receptor beta LBD in complex with specific agonist GC-24 | | Descriptor: | Thyroid hormone receptor beta-1, [4-(3-BENZYL-4-HYDROXYBENZYL)-3,5-DIMETHYLPHENOXY]ACETIC ACID | | Authors: | Borngraeber, S, Budny, M.J, Chiellini, G, Cunha-Lima, S.T, Togashi, M, Webb, P, Baxter, J.D, Scanlan, T.S, Fletterick, R.J. | | Deposit date: | 2003-08-04 | | Release date: | 2004-02-03 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Ligand selectivity by seeking hydrophobicity in thyroid hormone receptor.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1Q60
 
 | | Solution Structure of RSGI RUH-004, a GTF2I domain in Mouse cDNA | | Descriptor: | General transcription factor II-I | | Authors: | Doi-Katayama, Y, Hayashi, F, Hirota, H, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-08-12 | | Release date: | 2004-11-23 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of RSGI RUH-004, a GTF2I domain in Mouse cDNA
To be published
|
|
1PKX
 
 | | Crystal Structure of human ATIC in complex with XMP | | Descriptor: | Bifunctional purine biosynthesis protein PURH, POTASSIUM ION, XANTHOSINE-5'-MONOPHOSPHATE | | Authors: | Wolan, D.W, Cheong, C.G, Greasley, S.E, Wilson, I.A. | | Deposit date: | 2003-06-06 | | Release date: | 2004-02-17 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Insights into the Human and Avian IMP Cyclohydrolase Mechanism via Crystal Structures with the Bound XMP Inhibitor.
Biochemistry, 43, 2004
|
|
1PML
 
 | |
1PSH
 
 | |
1PNH
 
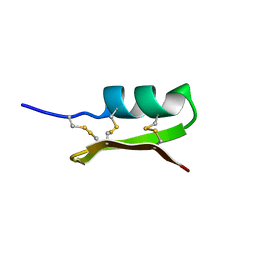 | | SOLUTION STRUCTURE OF PO5-NH2, A SCORPION TOXIN ANALOG WITH HIGH AFFINITY FOR THE APAMIN-SENSITIVE POTASSIUM CHANNEL | | Descriptor: | SCORPION TOXIN | | Authors: | Meunier, S, Bernassau, J.-M, Sabatier, J.-M, Martin-Eauclaire, M.-F, Van Rietschoten, J, Cambillau, C, Darbon, H. | | Deposit date: | 1993-08-25 | | Release date: | 1994-01-31 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of P05-NH2, a scorpion toxin analog with high affinity for the apamin-sensitive potassium channel.
Biochemistry, 32, 1993
|
|
1PT8
 
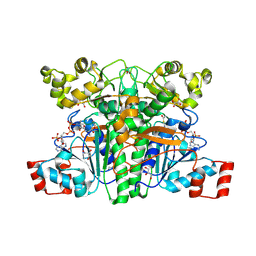 | | Crystal structure of the yfdW gene product of E. coli, in complex with oxalate and acetyl-CoA | | Descriptor: | ACETYL COENZYME *A, GLYCEROL, Hypothetical protein yfdW, ... | | Authors: | Gruez, A, Roig-Zamboni, V, Valencia, C, Campanacci, V, Cambillau, C. | | Deposit date: | 2003-06-23 | | Release date: | 2003-09-09 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure of the Escherichia coli yfdW gene product reveals a new fold of two interlaced rings identifying a wide family of CoA transferases.
J.Biol.Chem., 278, 2003
|
|
1R12
 
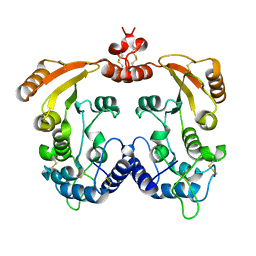 | | Native Aplysia ADP ribosyl cyclase | | Descriptor: | ADP-ribosyl cyclase | | Authors: | Love, M.L, Szebenyi, D.M.E, Kriksunov, I.A, Thiel, D.J, Munshi, C, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2003-09-23 | | Release date: | 2004-03-09 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | ADP-ribosyl cyclase; crystal structures reveal a covalent intermediate.
Structure, 12, 2004
|
|
1QEW
 
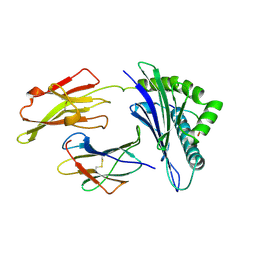 | | HUMAN CLASS I HISTOCOMPATIBILITY ANTIGEN (HLA-A 0201) COMPLEX WITH A NONAMERIC PEPTIDE FROM MELANOMA-ASSOCIATED ANTIGEN 3 (RESIDUES 271-279) | | Descriptor: | PROTEIN (BETA-2-MICROGLOBULIN), PROTEIN (HLA CLASS I HISTOCOMPATIBILITY ANTIGEN, B-35 B* 3501 ALPHA CHAIN), ... | | Authors: | Orth, P, Alings, C, Saenger, W, Ziegler, A. | | Deposit date: | 1999-04-02 | | Release date: | 2003-11-18 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | HUMAN CLASS I HISTOCOMPATIBILITY ANTIGEN (HLA-A 0201)
COMPLEX WITH A NONAMERIC PEPTIDE FROM MELANOMA-ASSOCIATED
ANTIGEN 3 (RESIDUES 271-279)
To be Published
|
|
1QX5
 
 | |
3UJZ
 
 | |
1R0S
 
 | | Crystal structure of ADP-ribosyl cyclase Glu179Ala mutant | | Descriptor: | ADP-ribosyl cyclase | | Authors: | Love, M.L, Szebenyi, D.M.E, Kriksunov, I.A, Thiel, D.J, Munshi, C, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2003-09-22 | | Release date: | 2004-03-09 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | ADP-ribosyl cyclase; crystal structures reveal a covalent intermediate.
Structure, 12, 2004
|
|
1QMN
 
 | |
3SKB
 
 | | Structural characterization of a GII.4 2004 norovirus variant (TCH05) | | Descriptor: | Capsid | | Authors: | Shanker, S, Choi, J.-M, Sankaran, B, Atmar, R.L, Estes, M.K, Prasad, B.V.V. | | Deposit date: | 2011-06-22 | | Release date: | 2011-07-13 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.22 Å) | | Cite: | Structural Analysis of Histo-Blood Group Antigen Binding Specificity in a Norovirus GII.4 Epidemic Variant: Implications for Epochal Evolution.
J.Virol., 85, 2011
|
|
1QRD
 
 | | QUINONE REDUCTASE/FAD/CIBACRON BLUE/DUROQUINONE COMPLEX | | Descriptor: | CIBACRON BLUE, DUROQUINONE, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Li, R, Bianchet, M.A, Talalay, P, Amzel, L.M. | | Deposit date: | 1995-07-28 | | Release date: | 1996-10-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The three-dimensional structure of NAD(P)H:quinone reductase, a flavoprotein involved in cancer chemoprotection and chemotherapy: mechanism of the two-electron reduction.
Proc.Natl.Acad.Sci.USA, 92, 1995
|
|
3SJP
 
 | | Structural characterization of a GII.4 2004 norovirus variant (TCH05) | | Descriptor: | Capsid, ZINC ION | | Authors: | Shanker, S, Choi, J.-M, Sankaran, B, Atmar, R.L, Estes, M.K, Prasad, B.V.V. | | Deposit date: | 2011-06-21 | | Release date: | 2011-07-13 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.004 Å) | | Cite: | Structural Analysis of Histo-Blood Group Antigen Binding Specificity in a Norovirus GII.4 Epidemic Variant: Implications for Epochal Evolution.
J.Virol., 85, 2011
|
|
3V0C
 
 | | 4.3 angstrom crystal structure of an inactive BoNT/A (E224Q/R363A/Y366F) | | Descriptor: | BoNT/A, ZINC ION | | Authors: | Gu, S, Rumpel, S, Zhou, J, Strotmeier, J, Bigalke, H, Perry, K, Shoemaker, C.B, Rummel, A, Jin, R. | | Deposit date: | 2011-12-07 | | Release date: | 2012-03-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (4.3 Å) | | Cite: | Botulinum neurotoxin is shielded by NTNHA in an interlocked complex.
Science, 335, 2012
|
|
