3RIB
 
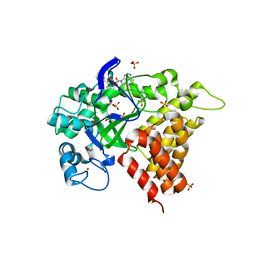 | | Human lysine methyltransferase Smyd2 in complex with AdoHcy | | Descriptor: | N-lysine methyltransferase SMYD2, S-ADENOSYL-L-HOMOCYSTEINE, SULFATE ION, ... | | Authors: | Xu, S, Zhang, T, Zhong, C, Ding, J. | | Deposit date: | 2011-04-13 | | Release date: | 2011-07-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Structure of human lysine methyltransferase Smyd2 reveals insights into the substrate divergence in Smyd proteins
J Mol Cell Biol, 3, 2011
|
|
6XT9
 
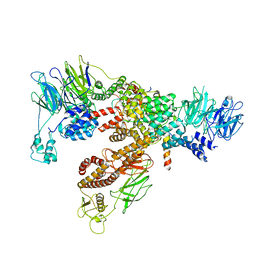 | | Subunits BBS 1,4,8,9,18 of the human BBSome complex | | Descriptor: | BBSome-interacting protein 1, Bardet-Biedl syndrome 1 protein, Bardet-Biedl syndrome 4 protein, ... | | Authors: | Klink, B.U, Raunser, S, Gatsogiannis, C. | | Deposit date: | 2020-01-15 | | Release date: | 2020-01-29 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structure of the human BBSome core complex.
Elife, 9, 2020
|
|
6XYQ
 
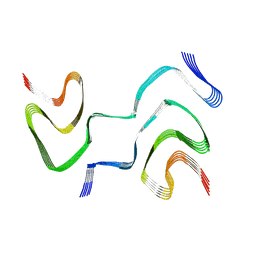 | | Multiple system atrophy Type II-2 alpha-synuclein filament | | Descriptor: | Alpha-synuclein | | Authors: | Schweighauser, M, Shi, Y, Tarutani, A, Kametani, F, Murzin, A.G, Ghetti, B, Matsubara, T, Tomita, T, Ando, T, Hasegawa, K, Murayama, S, Yoshida, M, Hasegawa, M, Scheres, S.H.W, Goedert, M. | | Deposit date: | 2020-01-30 | | Release date: | 2020-02-12 | | Last modified: | 2024-05-22 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Structures of alpha-synuclein filaments from multiple system atrophy.
Nature, 585, 2020
|
|
7UAK
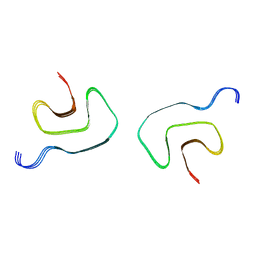 
 | |
7MFE
 
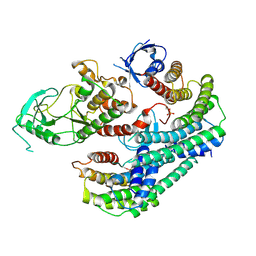 | | Autoinhibited BRAF:(14-3-3)2 complex with the BRAF RBD resolved | | Descriptor: | 14-3-3 protein zeta/delta, Serine/threonine-protein kinase B-raf, ZINC ION | | Authors: | Martinez Fiesco, J.A, Ping, Z, Durrant, D.E, Morrison, D.K. | | Deposit date: | 2021-04-09 | | Release date: | 2022-01-26 | | Last modified: | 2022-02-09 | | Method: | ELECTRON MICROSCOPY (4.07 Å) | | Cite: | Structural insights into the BRAF monomer-to-dimer transition mediated by RAS binding.
Nat Commun, 13, 2022
|
|
7MFF
 
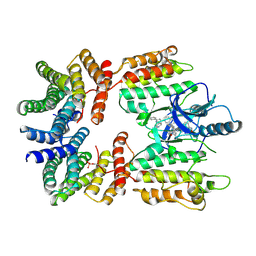 | | Dimeric (BRAF)2:(14-3-3)2 complex bound to SB590885 Inhibitor | | Descriptor: | (1Z)-5-(2-{4-[2-(DIMETHYLAMINO)ETHOXY]PHENYL}-5-PYRIDIN-4-YL-1H-IMIDAZOL-4-YL)INDAN-1-ONE OXIME, 14-3-3 protein zeta/delta, Serine/threonine-protein kinase B-raf | | Authors: | Martinez Fiesco, J.A, Ping, Z, Durrant, D.E, Morrison, D.K. | | Deposit date: | 2021-04-09 | | Release date: | 2022-01-26 | | Last modified: | 2022-02-09 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Structural insights into the BRAF monomer-to-dimer transition mediated by RAS binding.
Nat Commun, 13, 2022
|
|
1VK5
 
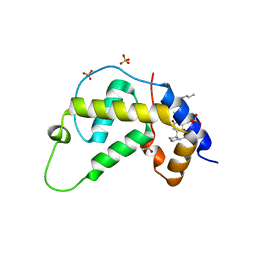 | | X-ray Structure of Gene Product from Arabidopsis Thaliana At3g22680 | | Descriptor: | 1,2-ETHANEDIOL, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, SULFATE ION, ... | | Authors: | Wesenberg, G.E, Smith, D.W, Phillips Jr, G.N, Johnson, K.A, Bingman, C.A, Allard, S.T.M, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2004-05-06 | | Release date: | 2004-05-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.604 Å) | | Cite: | Structure at 1.6 A resolution of the protein from gene locus At3g22680 from Arabidopsis thaliana.
Acta Crystallogr.,Sect.F, 61, 2005
|
|
1VRL
 
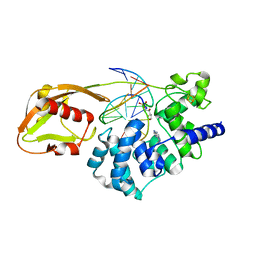 | | MutY adenine glycosylase in complex with DNA and soaked adenine free base | | Descriptor: | 5'-D(*AP*AP*GP*AP*CP*(8OG)P*TP*GP*GP*AP*C)-3', 5'-D(*TP*GP*TP*CP*CP*AP*(HPD)P*GP*TP*CP*T)-3', ADENINE, ... | | Authors: | Fromme, J.C, Banerjee, A, Huang, S.J, Verdine, G.L. | | Deposit date: | 2005-03-08 | | Release date: | 2005-03-22 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis for removal of adenine mispaired with 8-oxoguanine by MutY adenine DNA glycosylase
Nature, 427, 2004
|
|
3PSL
 
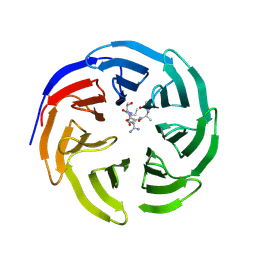 | | Fine-tuning the stimulation of MLL1 methyltransferase activity by a histone H3 based peptide mimetic | | Descriptor: | N-alpha acetylated form of histone H3, WD repeat-containing protein 5 | | Authors: | Avdic, V, Zhang, P, Lanouette, S, Voronova, A, Skerjanc, I, Couture, J.-F. | | Deposit date: | 2010-12-01 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Fine-tuning the stimulation of MLL1 methyltransferase activity by a histone H3-based peptide mimetic.
Faseb J., 25, 2011
|
|
3EIK
 
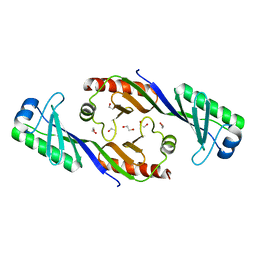 | |
4MD8
 
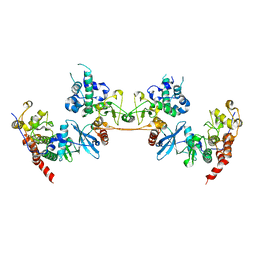 | |
3EG6
 
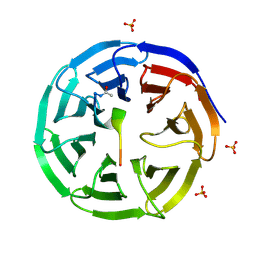 | |
7WMM
 
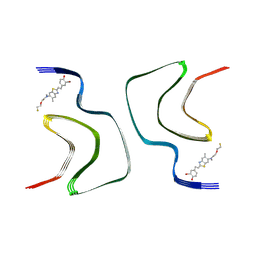 | | alpha-synuclein fibril-F0502B complex | | Descriptor: | 2-bromanyl-4-[(~{E})-2-[6-[2-(2-fluoranylethoxy)ethyl-methyl-amino]-5-methyl-1,3-benzothiazol-2-yl]ethenyl]phenol, Alpha-synuclein | | Authors: | Tao, Y.Q, Zhao, Q.Y, Liu, C, Li, D. | | Deposit date: | 2022-01-15 | | Release date: | 2023-01-18 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | alpha-synuclein fibril-F0502B complex
To Be Published
|
|
7MFD
 
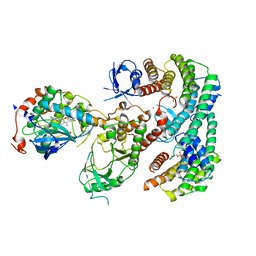 | | Autoinhibited BRAF:(14-3-3)2:MEK complex with the BRAF RBD resolved | | Descriptor: | 14-3-3 protein zeta/delta, Dual specificity mitogen-activated protein kinase kinase 1, N-(3-fluoro-4-{[4-methyl-2-oxo-7-(pyrimidin-2-yloxy)-2H-chromen-3-yl]methyl}pyridin-2-yl)-N'-methylsulfuric diamide, ... | | Authors: | Martinez Fiesco, J.A, Ping, Z, Durrant, D.E, Morrison, D.K. | | Deposit date: | 2021-04-09 | | Release date: | 2022-01-26 | | Last modified: | 2022-02-09 | | Method: | ELECTRON MICROSCOPY (3.66 Å) | | Cite: | Structural insights into the BRAF monomer-to-dimer transition mediated by RAS binding.
Nat Commun, 13, 2022
|
|
7ZIT
 
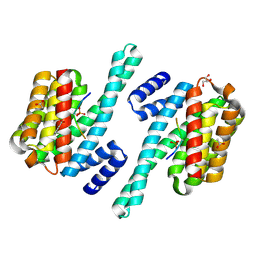 | | 14-3-3 in complex with SARS-COV2 N phospho-peptide | | Descriptor: | 14-3-3 protein zeta/delta, ACETATE ION, BENZOIC ACID, ... | | Authors: | Eisenreichova, A, Boura, E. | | Deposit date: | 2022-04-08 | | Release date: | 2022-10-26 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structural basis for SARS-CoV-2 nucleocapsid (N) protein recognition by 14-3-3 proteins.
J.Struct.Biol., 214, 2022
|
|
6IDF
 
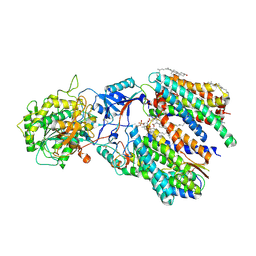 | | Cryo-EM structure of gamma secretase in complex with a Notch fragment | | Descriptor: | 1,2-DIACYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Yang, G, Zhou, R, Zhou, Q, Guo, X, Yan, C, Ke, M, Lei, J, Shi, Y. | | Deposit date: | 2018-09-09 | | Release date: | 2018-12-26 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Structural basis of Notch recognition by human gamma-secretase
Nature, 565, 2019
|
|
6GES
 
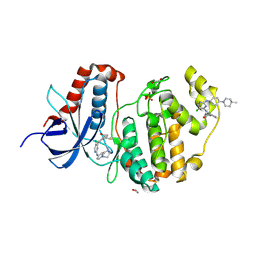 | | Crystal structure of ERK1 covalently bound to SM1-71 | | Descriptor: | 1,2-ETHANEDIOL, Mitogen-activated protein kinase 3, N-{2-[(5-chloro-2-{[4-(4-methylpiperazin-1-yl)phenyl]amino}pyrimidin-4-yl)amino]phenyl}propanamide, ... | | Authors: | Chaikuad, A, Suman, R, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Gray, N.S, Knapp, S, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-04-27 | | Release date: | 2019-02-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Leveraging Compound Promiscuity to Identify Targetable Cysteines within the Kinome.
Cell Chem Biol, 26, 2019
|
|
4MD9
 
 | |
4MD7
 
 | |
8AH2
 
 | | Crystal structure of human 14-3-3 zeta fused to the NPM1 peptide including phosphoserine-48 | | Descriptor: | 14-3-3 protein zeta/delta,Nucleophosmin | | Authors: | Boyko, K.M, Kapitonova, A.A, Tugaeva, K.V, Varfolomeeva, L.A, Sluchanko, N.N. | | Deposit date: | 2022-07-20 | | Release date: | 2022-09-14 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis for the recognition by 14-3-3 proteins of a conditional binding site within the oligomerization domain of human nucleophosmin.
Biochem.Biophys.Res.Commun., 627, 2022
|
|
7XO2
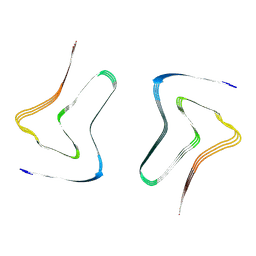 
 | |
7XO0
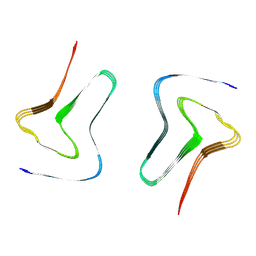 
 | |
7XO1
 
 | |
7XO3
 
 | |
7XJX
 
 | | The cryo-EM structure of Fe3+ induced alpha-syn fibril. | | Descriptor: | Alpha-synuclein | | Authors: | Zhao, Q.Y, Tao, Y.Q, Zhao, K, Tao, Y.Q, Li, D. | | Deposit date: | 2022-04-18 | | Release date: | 2023-01-18 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | Structural Insights of Fe3+ Induced alpha-synuclein Fibrillation in Parkinson' Disease
J.Mol.Biol., 435, 2023
|
|
