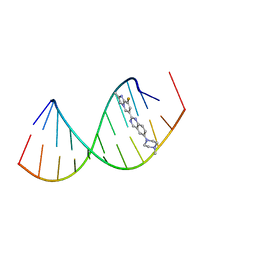
8R8P
 
 | |
6C2U
 
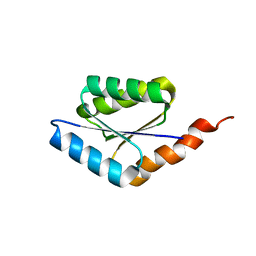 | | Solution structure of a phosphate-loop protein | | Descriptor: | phosphate-loop protein | | Authors: | Yang, F, Yang, W, Lin, Y.R, Romero Romero, M.L, Tawfik, D, Baker, D, Varani, G. | | Deposit date: | 2018-01-09 | | Release date: | 2018-11-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Simple yet functional phosphate-loop proteins.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
7T2F
 
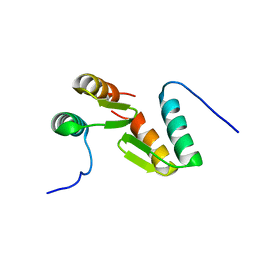 | | Solution structure of the model HEEH mini protein homodimer HEEH_TK_rd5_0341 | | Descriptor: | HEEH mini protein HEEH_TK_rd5_0341 | | Authors: | Lemak, A, Houliston, S, Kim, T.-E, Martel, C, Rocklin, G.J, Arrowsmith, C.H. | | Deposit date: | 2021-12-04 | | Release date: | 2022-10-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Dissecting the stability determinants of a challenging de novo protein fold using massively parallel design and experimentation.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
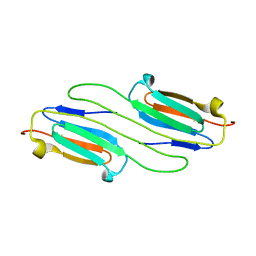
6BP9
 
 | |
8P7G
 
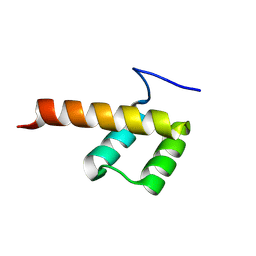 | |
6C2V
 
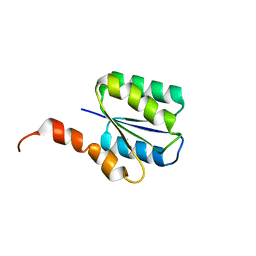 | | Solution structure of a phosphate-loop protein | | Descriptor: | phosphate-loop protein | | Authors: | Yang, F, Yang, W, Lin, Y.R, Romero Romero, M.L, Tawfik, D, Baker, D, Varani, G. | | Deposit date: | 2018-01-09 | | Release date: | 2018-11-21 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Simple yet functional phosphate-loop proteins.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3KVD
 
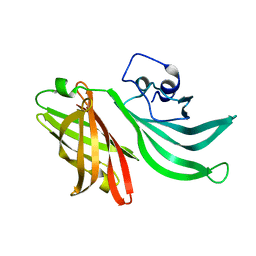 | | Crystal structure of the Neisseria meningitidis Factor H binding protein, fHbp (GNA1870) at 2.0 A resolution | | Descriptor: | Lipoprotein | | Authors: | Cendron, L, Veggi, D, Girardi, E, Zanotti, G. | | Deposit date: | 2009-11-30 | | Release date: | 2010-12-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the uncomplexed Neisseria meningitidis factor H-binding protein fHbp (rLP2086).
Acta Crystallogr.,Sect.F, 67, 2011
|
|
8R63
 
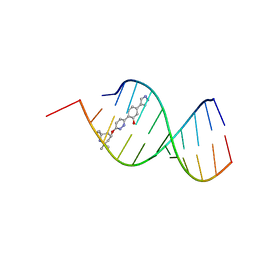 | | Solution structure of branaplam bound to the RNA duplex formed upon 5'-splice site recognition | | Descriptor: | 5-(1~{H}-pyrazol-4-yl)-2-[6-(2,2,6,6-tetramethylpiperidin-4-yl)oxypyridazin-3-yl]phenol, RNA (5'-R(*AP*UP*AP*CP*(PSU)P*(PSU)P*AP*CP*CP*UP*G)-3'), RNA (5'-R(P*GP*GP*AP*GP*UP*AP*AP*GP*UP*CP*U)-3') | | Authors: | Malard, F, Campagne, S. | | Deposit date: | 2023-11-20 | | Release date: | 2024-03-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The diversity of splicing modifiers acting on A-1 bulged 5'-splice sites reveals rules for rational drug design.
Nucleic Acids Res., 52, 2024
|
|
2V1V
 | |
8QTQ
 | | Thermostable WW domain | | Descriptor: | WW domain | | Authors: | Kovermann, M, Thomas, F. | | Deposit date: | 2023-10-13 | | Release date: | 2024-07-03 | | Method: | SOLUTION NMR | | Cite: | Thermostable WW-Domain Scaffold to Design Functional beta-Sheet Miniproteins.
J.Am.Chem.Soc., 2024
|
|
2WZD
 
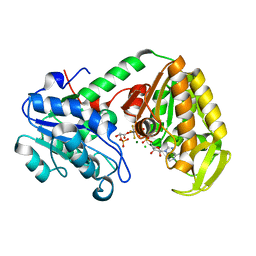 | | The catalytically active fully closed conformation of human phosphoglycerate kinase K219A mutant in complex with ADP, 3PG and aluminium trifluoride | | Descriptor: | 3-PHOSPHOGLYCERIC ACID, ADENOSINE-5'-DIPHOSPHATE, ALUMINUM FLUORIDE, ... | | Authors: | Bowler, M.W, Cliff, M.J, Marston, J.P.M, Baxter, N.J, Hownslow, A.M.H, Varga, A.V, Szabo, J, Vas, M, Blackburn, G.M, Waltho, J.P. | | Deposit date: | 2009-11-27 | | Release date: | 2010-04-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Transition State Analogue Structures of Human Phosphoglycerate Kinase Establish the Importance of Charge Balance in Catalysis.
J.Am.Chem.Soc., 132, 2010
|
|
6CO4
 
 | |
6CPI
 
 | | Solution structure of SH3 domain from Shank1 | | Descriptor: | SH3 and multiple ankyrin repeat domains protein 1 | | Authors: | Ishida, H, Vogel, H.J. | | Deposit date: | 2018-03-13 | | Release date: | 2018-08-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the SH3 domains from Shank scaffold proteins and their interactions with Cav1.3 calcium channels.
FEBS Lett., 592, 2018
|
|
6CPK
 
 | | Solution structure of SH3 domain from Shank3 | | Descriptor: | SH3 and multiple ankyrin repeat domains protein 3 | | Authors: | Ishida, H, Vogel, H.J. | | Deposit date: | 2018-03-13 | | Release date: | 2018-08-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the SH3 domains from Shank scaffold proteins and their interactions with Cav1.3 calcium channels.
FEBS Lett., 592, 2018
|
|
6CL3
 | | LyeTxI-b, a synthetic peptide derived from Lycosa erythrognatha spider venom, shows potent antibiotic activity, in vitro and in vivo | | Descriptor: | Toxin LyeTx 1 | | Authors: | de Lima, M.E, dos Reis, P.V, Resende, J.M, Verly, R.M. | | Deposit date: | 2018-03-01 | | Release date: | 2018-05-30 | | Method: | SOLUTION NMR | | Cite: | LyeTxI-b, a Synthetic Peptide Derived FromLycosa erythrognathaSpider Venom, Shows Potent Antibiotic Activityin Vitroandin Vivo.
Front Microbiol, 9, 2018
|
|
6CPJ
 
 | | Solution structure of SH3 domain from Shank2 | | Descriptor: | SH3 and multiple ankyrin repeat domains protein 2 | | Authors: | Ishida, H, Vogel, H.J. | | Deposit date: | 2018-03-13 | | Release date: | 2018-08-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the SH3 domains from Shank scaffold proteins and their interactions with Cav1.3 calcium channels.
FEBS Lett., 592, 2018
|
|
6D2H
 
 | |
3IAP
 
 | | E. coli (lacZ) beta-galactosidase (E416Q) | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Beta-galactosidase, DIMETHYL SULFOXIDE, ... | | Authors: | Lo, S, Dugdale, M.L, Jeerh, N, Ku, T, Roth, N.J, Huber, R.E. | | Deposit date: | 2009-07-14 | | Release date: | 2009-12-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Studies of Glu-416 variants of beta-galactosidase (E. coli) show that the active site Mg(2+) is not important for structure and indicate that the main role of Mg (2+) is to mediate optimization of active site chemistry
Protein J., 29, 2010
|
|
6CUC
 
 | |
6CFA
 
 | | peptide PaAMP1R3 | | Descriptor: | peptide PaAMP1R3 | | Authors: | Alves, E.S.F, Liao, L.M. | | Deposit date: | 2018-02-14 | | Release date: | 2019-03-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Synthetic peptide PaAMP1R3
To be Published
|
|
7R6P
 
 | |
7MJ3
 
 | | Structure of Hact-4 | | Descriptor: | Hact-4 | | Authors: | Schmidt, C.A, Daly, N.L. | | Deposit date: | 2021-04-19 | | Release date: | 2022-08-03 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Diversity of coral derived peptides
To Be Published
|
|
7NIP
 
 | |
7RUL
 
 | |
7PMP
 
 | |
