3F4B
 | |
3TJB
 
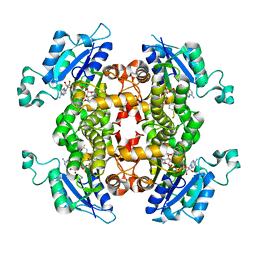
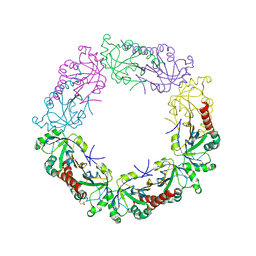 | | Crystal structure of wild-type human peroxiredoxin IV | | Descriptor: | Peroxiredoxin-4 | | Authors: | Cao, Z, Tavender, T.J, Roszak, A.W, Cogdell, R.J, Bulleid, N.J. | | Deposit date: | 2011-08-24 | | Release date: | 2011-10-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Crystal Structure of Reduced and of Oxidized Peroxiredoxin IV Enzyme Reveals a Stable Oxidized Decamer and a Non-disulfide-bonded Intermediate in the Catalytic Cycle.
J.Biol.Chem., 286, 2011
|
|
2KLV
 
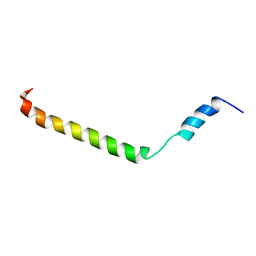 | | Membrane-bound structure of the Pf1 major coat protein in DHPC micelle | | Descriptor: | Capsid protein G8P | | Authors: | Park, S, Son, W, Mukhopadhyay, R, Valafar, H, Opella, S.J. | | Deposit date: | 2009-07-08 | | Release date: | 2009-10-06 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Phage-induced alignment of membrane proteins enables the measurement and structural analysis of residual dipolar couplings with dipolar waves and lambda-maps.
J.Am.Chem.Soc., 131, 2009
|
|
3TJJ
 
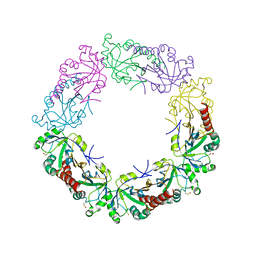 | | Crystal structure of human peroxiredoxin IV C245A mutant in sulfenylated form | | Descriptor: | Peroxiredoxin-4 | | Authors: | Cao, Z, Tavender, T.J, Roszak, A.W, Cogdell, R.J, Bulleid, N.J. | | Deposit date: | 2011-08-24 | | Release date: | 2011-10-12 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Crystal Structure of Reduced and of Oxidized Peroxiredoxin IV Enzyme Reveals a Stable Oxidized Decamer and a Non-disulfide-bonded Intermediate in the Catalytic Cycle.
J.Biol.Chem., 286, 2011
|
|
3TJF
 
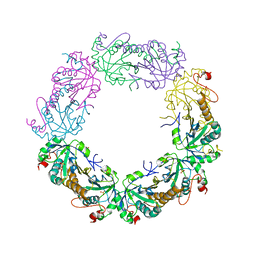 | | Crystal Structure of human peroxiredoxin IV C51A mutant in reduced form | | Descriptor: | Peroxiredoxin-4, SULFATE ION | | Authors: | Cao, Z, Tavender, T.J, Roszak, A.W, Cogdell, R.J, Bulleid, N.J. | | Deposit date: | 2011-08-24 | | Release date: | 2011-10-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Crystal Structure of Reduced and of Oxidized Peroxiredoxin IV Enzyme Reveals a Stable Oxidized Decamer and a Non-disulfide-bonded Intermediate in the Catalytic Cycle.
J.Biol.Chem., 286, 2011
|
|
3STU
 
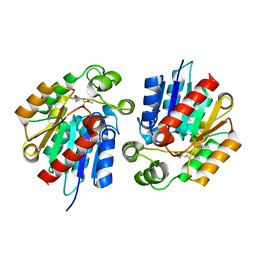 | |
2IDJ
 
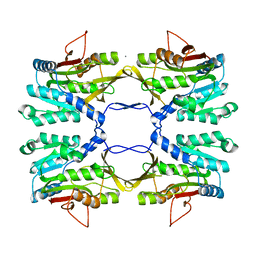 | | Crystal Structure of Rat Glycine N-Methyltransferase Apoprotein, Monoclinic Form | | Descriptor: | CALCIUM ION, Glycine N-methyltransferase | | Authors: | Luka, Z, Pakhomova, S, Loukachevitch, L.V, Egli, M, Newcomer, M.E, Wagner, C. | | Deposit date: | 2006-09-15 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | 5-methyltetrahydrofolate is bound in intersubunit areas of rat liver folate-binding protein glycine N-methyltransferase.
J.Biol.Chem., 282, 2007
|
|
3DSZ
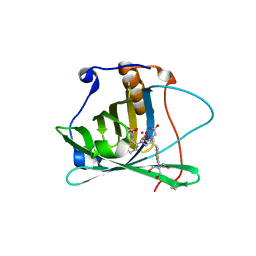 
 | | Engineered human lipocalin 2 in complex with Y-DTPA | | Descriptor: | N-{(1S,2S)-2-[bis(carboxymethyl)amino]cyclohexyl}-N-{(2R)-2-[bis(carboxymethyl)amino]-3-[4-({[2-hydroxy-1,1-bis(hydroxymethyl)ethyl]carbamothioyl}amino)phenyl]propyl}glycine, YTTRIUM (III) ION, engineered human lipocalin 2 | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2008-07-14 | | Release date: | 2009-05-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High-affinity recognition of lanthanide(III) chelate complexes by a reprogrammed human lipocalin 2
J.Am.Chem.Soc., 131, 2009
|
|
3RVG
 
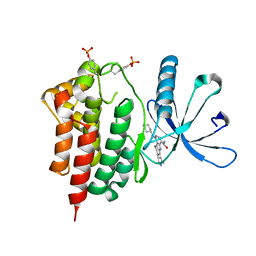 | | Crystals structure of Jak2 with a 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitor | | Descriptor: | 1-(cyclohexylamino)-7-(1-methyl-1H-pyrazol-4-yl)-5H-pyrido[4,3-b]indole-4-carboxamide, Tyrosine-protein kinase JAK2 | | Authors: | Lim, J, Taoka, B, Otte, R.D, Spencer, K, Dinsmore, C.J, Altman, M.D, Chan, G, Rosenstein, C, Sharma, S, Su, H.P, Szewczak, A.A, Xu, L, Yin, H, Zugay-Murphy, J, Marshall, C.G, Young, J.R. | | Deposit date: | 2011-05-06 | | Release date: | 2012-03-21 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | Discovery of 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitors of Janus kinase 2 (JAK2) for the treatment of myeloproliferative disorders.
J.Med.Chem., 54, 2011
|
|
3QVS
 
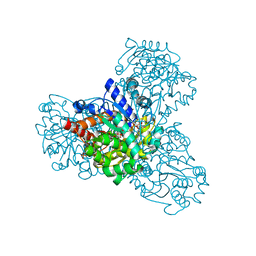 | | L-myo-inositol 1-phosphate synthase from Archaeoglobus fulgidus wild type | | Descriptor: | GLYCEROL, Myo-inositol-1-phosphate synthase (Ino1), NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Neelon, K, Roberts, M.F, Stec, B. | | Deposit date: | 2011-02-25 | | Release date: | 2012-01-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of a trapped catalytic intermediate suggests that forced atomic proximity drives the catalysis of mIPS.
Biophys.J., 101, 2011
|
|
3DTQ
 
 | |
3QW2
 
 | | L-myo-inositol 1-phosphate synthase from Archaeoglobus mutant N255A | | Descriptor: | GLYCEROL, Myo-inositol-1-phosphate synthase (Ino1), NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Neelon, K, Roberts, M.F, Stec, B. | | Deposit date: | 2011-02-26 | | Release date: | 2012-01-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Crystal structure of a trapped catalytic intermediate suggests that forced atomic proximity drives the catalysis of mIPS.
Biophys.J., 101, 2011
|
|
3QGW
 
 | | Crystal Structure of ITK kinase bound to an inhibitor | | Descriptor: | 3-[(8-phenylthieno[2,3-h]quinazolin-2-yl)amino]benzenesulfonamide, N-(6-oxo-1,6-dihydro-3,4'-bipyridin-5-yl)benzamide, Tyrosine-protein kinase ITK/TSK | | Authors: | Brown, K, Cheetham, G.M.T. | | Deposit date: | 2011-01-25 | | Release date: | 2011-06-22 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Discovery and structure-activity relationship of 3-aminopyrid-2-ones as potent and selective interleukin-2 inducible T-cell kinase (Itk) inhibitors
J.Med.Chem., 54, 2011
|
|
3QCP
 
 | | QSOX from Trypanosoma brucei | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, QSOX from Trypanosoma brucei (TbQSOX) | | Authors: | Alon, A, Fass, D. | | Deposit date: | 2011-01-17 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The dynamic disulphide relay of quiescin sulphydryl oxidase.
Nature, 488, 2012
|
|
3DKI
 
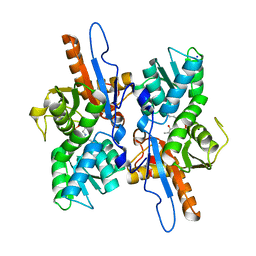 | |
3TJM
 
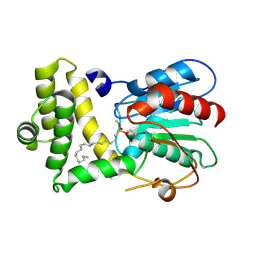 | | Crystal Structure of the Human Fatty Acid Synthase Thioesterase Domain with an Activate Site-Specific Polyunsaturated Fatty Acyl Adduct | | Descriptor: | Fatty acid synthase, methyl (R)-(6Z,9Z,12Z)-octadeca-6,9,12-trien-1-ylphosphonofluoridate | | Authors: | Zhang, W, Zheng, F, Florante, A.Q. | | Deposit date: | 2011-08-24 | | Release date: | 2011-09-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Crystal structure of FAS thioesterase domain with polyunsaturated fatty acyl adduct and inhibition by dihomo-gamma-linolenic acid.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2KIR
 
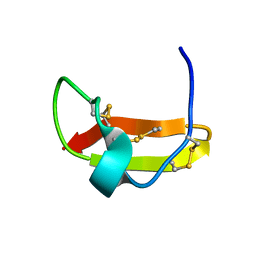 | | Solution structure of a designer toxin, mokatoxin-1 | | Descriptor: | Designer toxin | | Authors: | Biancalana, M, Koide, A, Takacs, Z, Goldstein, S, Koide, S. | | Deposit date: | 2009-05-07 | | Release date: | 2009-12-29 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | A designer ligand specific for Kv1.3 channels from a scorpion neurotoxin-based library.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3GAR
 
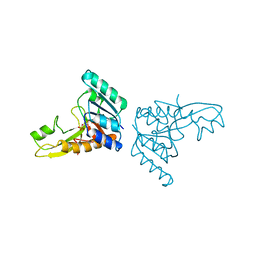 | | A PH-DEPENDENT STABLIZATION OF AN ACTIVE SITE LOOP OBSERVED FROM LOW AND HIGH PH CRYSTAL STRUCTURES OF MUTANT MONOMERIC GLYCINAMIDE RIBONUCLEOTIDE TRANSFORMYLASE | | Descriptor: | GLYCINAMIDE RIBONUCLEOTIDE TRANSFORMYLASE, PHOSPHATE ION | | Authors: | Su, Y, Yamashita, M.M, Greasley, S.E, Mullen, C.A, Shim, J.H, Jennings, P.A, Benkovic, S.J, Wilson, I.A. | | Deposit date: | 1998-05-13 | | Release date: | 1998-08-12 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A pH-dependent stabilization of an active site loop observed from low and high pH crystal structures of mutant monomeric glycinamide ribonucleotide transformylase at 1.8 to 1.9 A.
J.Mol.Biol., 281, 1998
|
|
2GP6
 
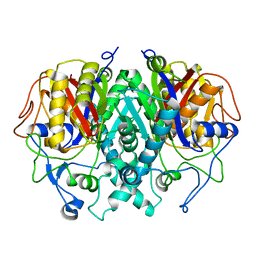 | |
2KT8
 
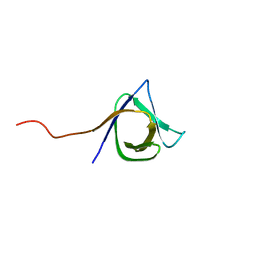 | | Solution NMR structure of the CPE1231(468-535) protein from Clostridium perfringens, Northeast Structural Genomics Consortium Target CpR82B | | Descriptor: | Probable surface protein | | Authors: | Yang, Y, Ramelot, T.A, Lee, D, Ciccosanti, C, Hamilton, K, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-01-21 | | Release date: | 2010-02-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the CPE1231(468-535) protein from Clostridium perfringens, Northeast Structural Genomics Consortium Target CpR82B
To be Published
|
|
2HS4
 
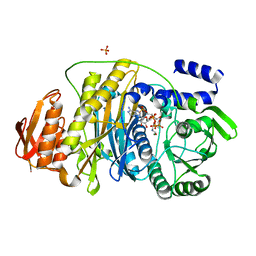 | | T. maritima PurL complexed with FGAR and AMPPCP | | Descriptor: | MAGNESIUM ION, N-(N-FORMYLGLYCYL)-5-O-PHOSPHONO-BETA-D-RIBOFURANOSYLAMINE, PHOSPHATE ION, ... | | Authors: | Ealick, S.E, Morar, M. | | Deposit date: | 2006-07-21 | | Release date: | 2007-01-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Complexed Structures of Formylglycinamide Ribonucleotide Amidotransferase from Thermotoga maritima Describe a Novel ATP Binding Protein Superfamily
Biochemistry, 45, 2006
|
|
3STT
 
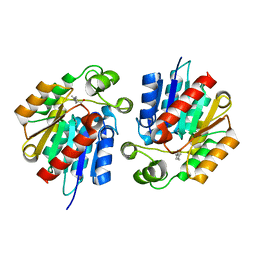 | |
3T58
 
 | |
3TBA
 
 | | Structure of Yeast Ribonucleotide Reductase 1 Q288A with dGTP and ADP | | Descriptor: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Ahmad, M.F, Kaushal, P.S, Wan, Q, Wijeratna, S.R, Huang, M, Dealwis, C. | | Deposit date: | 2011-08-05 | | Release date: | 2012-04-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Role of Arginine 293 and Glutamine 288 in Communication between Catalytic and Allosteric Sites in Yeast Ribonucleotide Reductase.
J.Mol.Biol., 419, 2012
|
|
3FB1
 
 | |
