2KQD
 
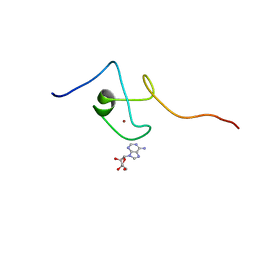 | | First PBZ domain of human APLF protein in complex with ribofuranosyladenosine | | Descriptor: | ADENOSINE, Aprataxin and PNK-like factor, ZINC ION, ... | | Authors: | Neuhaus, D, Eustermann, S, Brockmann, C, Yang, J. | | Deposit date: | 2009-11-04 | | Release date: | 2010-01-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the two PBZ domains from human APLF and their interaction with poly(ADP-ribose).
Nat.Struct.Mol.Biol., 17, 2010
|
|
2KQE
 
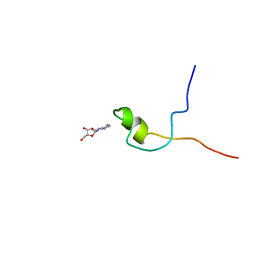 | | Second PBZ domain of human APLF protein in complex with ribofuranosyladenosine | | Descriptor: | ADENOSINE, Aprataxin and PNK-like factor, ZINC ION, ... | | Authors: | Neuhaus, D, Eustermann, S, Brockmann, C, Yang, J. | | Deposit date: | 2009-11-04 | | Release date: | 2010-01-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the two PBZ domains from human APLF and their interaction with poly(ADP-ribose).
Nat.Struct.Mol.Biol., 17, 2010
|
|
2KQF
 
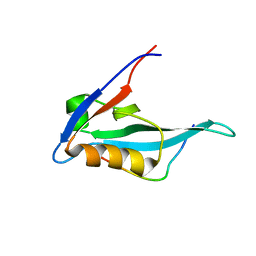 | | Solution structure of MAST205-PDZ complexed with the C-terminus of a rabies virus G protein | | Descriptor: | C-terminal motif from Glycoprotein, Microtubule-associated serine/threonine-protein kinase 2 | | Authors: | Terrien, E, Wolff, N, Cordier, F, Simenel, C, Bernard, A, Lafon, M, Delepierre, M. | | Deposit date: | 2009-11-04 | | Release date: | 2010-11-10 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | 1H, 13C and 15N resonance assignments of the PDZ of Microtubule-associated serine/threonine kinase 205 (MAST205) in complex with the C-Terminal motif from the rabies virus glycoprotein
To be Published
|
|
2KQG
 
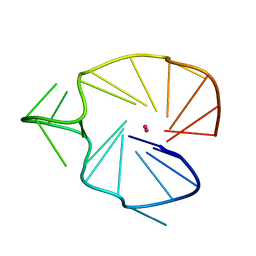 | | A G-rich sequence within the c-kit oncogene promoter forms a parallel G-quadruplex having asymmetric G-tetrad dynamics | | Descriptor: | 5'-D(*CP*GP*GP*GP*CP*GP*GP*GP*CP*AP*CP*GP*AP*GP*GP*GP*AP*GP*GP*GP*T)-3', POTASSIUM ION | | Authors: | Hsu, S.-T.D, Varnai, P, Bugaut, A, Reszka, A.P, Neidle, S, Balasubramanian, S. | | Deposit date: | 2009-11-05 | | Release date: | 2009-11-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A G-rich sequence within the c-kit oncogene promoter forms a parallel G-quadruplex having asymmetric G-tetrad dynamics
J.Am.Chem.Soc., 131, 2009
|
|
2KQH
 
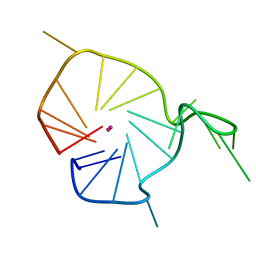 | | A G-rich sequence within the c-kit oncogene promoter forms a parallel G-quadruplex having asymmetric G-tetrad dynamics | | Descriptor: | 5'-D(*CP*GP*GP*GP*CP*GP*GP*GP*CP*GP*CP*GP*AP*GP*GP*GP*AP*GP*GP*GP*T)-3', POTASSIUM ION | | Authors: | Hsu, S.-T.D, Varnai, P, Bugaut, A, Reszka, A.P, Neidle, S, Balasubramanian, S. | | Deposit date: | 2009-11-05 | | Release date: | 2009-11-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A G-rich sequence within the c-kit oncogene promoter forms a parallel G-quadruplex having asymmetric G-tetrad dynamics
J.Am.Chem.Soc., 131, 2009
|
|
2KQK
 
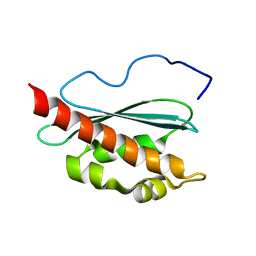 | | Solution structure of apo-IscU(D39A) | | Descriptor: | NifU-like protein | | Authors: | Kim, J.H, Fuzery, A.K, Tonelli, M, Vickery, L.E, Markley, J.L. | | Deposit date: | 2009-11-10 | | Release date: | 2010-11-17 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Three-Dimensional Structure and Determinants of Stability of the Iron-Sulfur Cluster Scaffold Protein IscU from Escherichia coli.
Biochemistry, 51, 2012
|
|
2KQL
 
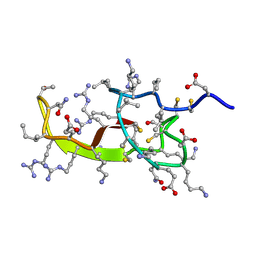 | |
2KQM
 
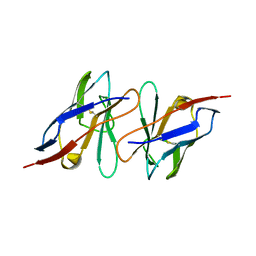 | |
2KQN
 
 | |
2KQO
 
 | |
2KQP
 
 | |
2KQQ
 
 | | NMR structure of human insulin mutant gly-b8-d-ala, his-b10-asp, pro-b28-lys, lys-b29-pro, 20 structures | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Hua, Q.X, Wan, Z.L, Zhao, M, Jia, W, Huang, K, Weiss, M.A. | | Deposit date: | 2009-11-13 | | Release date: | 2010-11-24 | | Last modified: | 2021-10-13 | | Method: | SOLUTION NMR | | Cite: | Structure and Dynamics of an Turn-Stabilize But Ina Insulin Analog.
To be Published
|
|
2KQR
 
 | |
2KQS
 
 | |
2KQT
 
 | | Solid-state NMR structure of the M2 transmembrane peptide of the influenza A virus in DMPC lipid bilayers bound to deuterated amantadine | | Descriptor: | (3S,5S,7S)-tricyclo[3.3.1.1~3,7~]decan-1-amine, M2 protein | | Authors: | Cady, S.D, Schmidt-Rohr, K, Wang, J, Soto, C.S, DeGrado, W.F, Hong, M. | | Deposit date: | 2009-11-18 | | Release date: | 2010-02-09 | | Last modified: | 2024-05-08 | | Method: | SOLID-STATE NMR | | Cite: | Structure of the amantadine binding site of influenza M2 proton channels in lipid bilayers
Nature, 463, 2010
|
|
2KQU
 
 | |
2KQV
 
 | |
2KQW
 | |
2KQX
 | |
2KQY
 
 | | Solution structure of Avian Thymic Hormone | | Descriptor: | Parvalbumin, thymic | | Authors: | Henzl, M.T. | | Deposit date: | 2009-11-24 | | Release date: | 2010-04-07 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structure of Avian Thymic Hormone, a High-Affinity Avian beta-Parvalbumin, in the Ca(2+)-Free and Ca(2+)-Bound States.
J.Mol.Biol., 397, 2010
|
|
2KQZ
 
 | |
2KR0
 
 | |
2KR1
 
 | | Solution NMR structure of zinc binding N-terminal domain of ubiquitin-protein ligase E3A from Homo Sapiens. Northeast Structural Genomics Consortium (NESG) target HR3662 | | Descriptor: | Ubiquitin protein ligase E3A, ZINC ION | | Authors: | Lemak, A, Yee, A, Fares, C, Semesi, A, Xiao, R, Montelione, G, Dhe-Paganon, S, Arrowsmith, C, Northeast Structural Genomics Consortium (NESG), Structural Genomics Consortium (SGC) | | Deposit date: | 2009-11-27 | | Release date: | 2009-12-22 | | Last modified: | 2020-02-26 | | Method: | SOLUTION NMR | | Cite: | Zn-binding AZUL domain of human ubiquitin protein ligase Ube3A.
J.Biomol.Nmr, 51, 2011
|
|
2KR2
 
 | |
2KR3
 
 | | Solution structure of SHA-D | | Descriptor: | Spectrin alpha chain, brain | | Authors: | Khristoforov, V.S, Prokhorov, D.A, Timchenko, M.A, Kudrevatykh, Y.A, Gushchina, L.V, Filimonov, V.V, Kutyshenko, V.P. | | Deposit date: | 2009-12-03 | | Release date: | 2010-09-15 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Chimeric SHA-D domain "SH3-Bergerac": 3D structure and dynamics studies
Russ.J.Bioorganic Chem., 36, 2010
|
|
