1M78
 
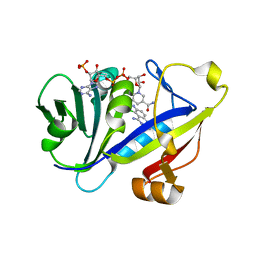 | | CANDIDA ALBICANS DIHYDROFOLATE REDUCTASE COMPLEXED WITH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE (NADPH) AND 5-CHLORYL-2,4,6-QUINAZOLINETRIAMINE (GW1225) | | Descriptor: | 5-CHLORYL-2,4,6-QUINAZOLINETRIAMINE, DIHYDROFOLATE REDUCTASE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Whitlow, M, Howard, A.J, Kuyper, L.F. | | Deposit date: | 2002-07-19 | | Release date: | 2003-03-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | X-Ray Crystallographic Studies of Candida Albicans Dihydrofolate Reductase. High Resolution Structures of the Holoenzyme and an Inhibited Ternary Complex.
J.Biol.Chem., 272, 1997
|
|
2QZA
 
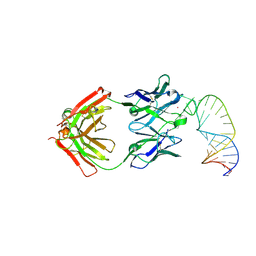
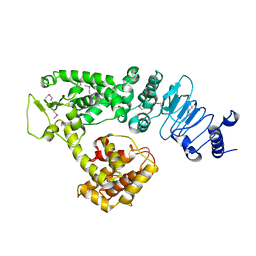 | | Crystal structure of Salmonella effector protein SopA | | Descriptor: | Secreted effector protein | | Authors: | Diao, J, Chen, J. | | Deposit date: | 2007-08-16 | | Release date: | 2007-12-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of SopA, a Salmonella effector protein mimicking a eukaryotic ubiquitin ligase.
Nat.Struct.Mol.Biol., 15, 2008
|
|
3BCA
 
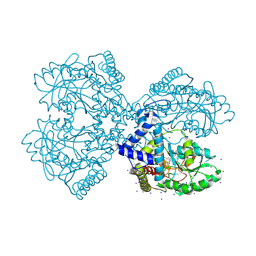 | |
1M7A
 
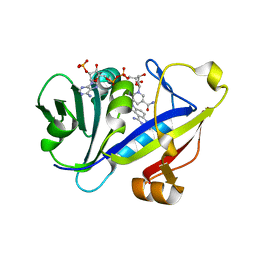 | | CANDIDA ALBICANS DIHYDROFOLATE REDUCTASE COMPLEXED WITH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE (NADPH) AND 7-[2-methoxy-1-(methoxymethyl)ethyl]-7H-pyrrolo[3,2-f] quinazoline-1,3-diamine (GW557) | | Descriptor: | 7-[2-METHOXY-1-(METHOXYMETHYL)ETHYL]-7H-PYRROLO[3,2-F] QUINAZOLINE-1,3-DIAMINE, DIHYDROFOLATE REDUCTASE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Whitlow, M, Howard, A.J, Kuyper, L.F. | | Deposit date: | 2002-07-19 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | X-Ray Crystallographic Studies of Candida Albicans Dihydrofolate Reductase. High Resolution Structures of the Holoenzyme and an Inhibited Ternary Complex.
J.Biol.Chem., 272, 1997
|
|
3ZX7
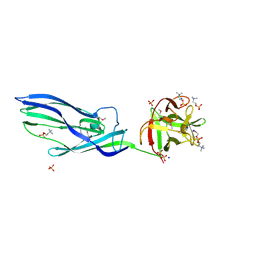 
 | | Complex of lysenin with phosphocholine | | Descriptor: | LYSENIN, PHOSPHATE ION, PHOSPHOCHOLINE, ... | | Authors: | De Colibus, L, Sonnen, A.F.P, Morris, K.J, Siebert, C.A, Abrusci, P, Plitzko, J, Hodnik, V, Leippe, M, Volpi, E, Anderluh, G, Gilbert, R.J.C. | | Deposit date: | 2011-08-08 | | Release date: | 2012-09-19 | | Last modified: | 2012-10-03 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Structures of Lysenin Reveal a Shared Evolutionary Origin for Pore-Forming Proteins and its Mode of Sphingomyelin Recognition.
Structure, 20, 2012
|
|
8D28
 
 | |
8D2A
 
 | |
8D2B
 
 | |
8D5L
 
 | | Crystal structure of theophylline aptamer in complex with TAL1 | | Descriptor: | 4-amino-8-methylpteridine-2,7(1H,8H)-dione, CALCIUM ION, MAGNESIUM ION, ... | | Authors: | Menichelli, E, Spraggon, G. | | Deposit date: | 2022-06-05 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Discovery of small molecules that target a tertiary-structured RNA.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
8D5O
 
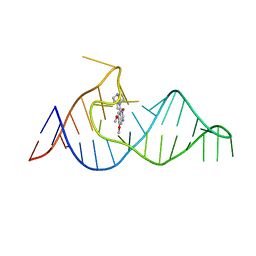 | |
8D29
 
 | |
8CSR
 
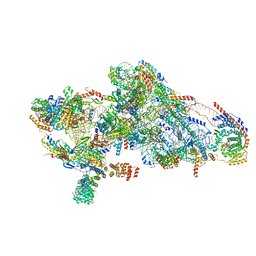 | | Human mitochondrial small subunit assembly intermediate (State C) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2023-02-15 | | Method: | ELECTRON MICROSCOPY (2.54 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
8CSS
 
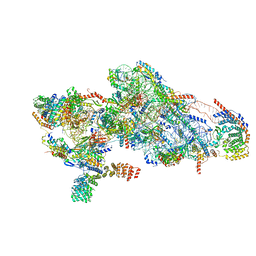 | | Human mitochondrial small subunit assembly intermediate (State D) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2023-02-08 | | Method: | ELECTRON MICROSCOPY (2.36 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
8D8L
 
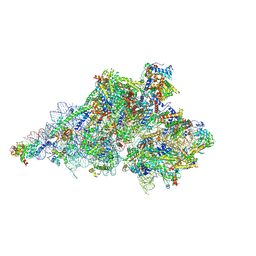 | | Yeast mitochondrial small subunit assembly intermediate (State 3) | | Descriptor: | 15S ribosomal RNA, 3-hydroxyisobutyryl-CoA hydrolase, mitochondrial, ... | | Authors: | Burnside, C, Harper, N, Klinge, S. | | Deposit date: | 2022-06-08 | | Release date: | 2022-12-14 | | Last modified: | 2023-02-15 | | Method: | ELECTRON MICROSCOPY (2.6 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
8CSQ
 
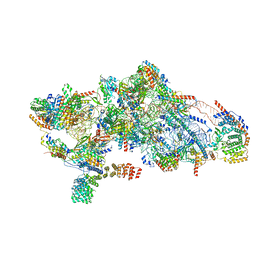 | | Human mitochondrial small subunit assembly intermediate (State B) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.54 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
8CSP
 
 | | Human mitochondrial small subunit assembly intermediate (State A) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.66 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
8CSU
 
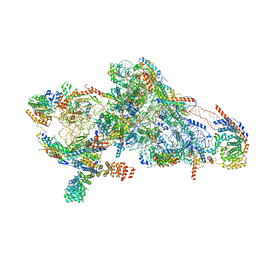 | | Human mitochondrial small subunit assembly intermediate (State C*) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2023-02-08 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
4UFU
 
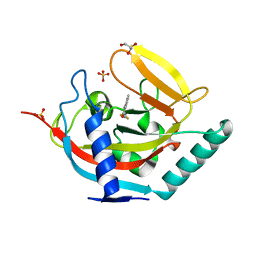 | | Crystal structure of human tankyrase 2 in complex with TA-12 | | Descriptor: | 8-methyl-2-[4-(trifluoromethyl)phenyl]-3H-quinazolin-4-one, GLYCEROL, SULFATE ION, ... | | Authors: | Haikarainen, T, Lehtio, L. | | Deposit date: | 2015-03-19 | | Release date: | 2016-04-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure-Activity Relationships of 2-Arylquinazolin-4-Ones as Highly Selective and Potent Inhibitors of the Tankyrases.
Eur.J.Med.Chem., 118, 2016
|
|
8CST
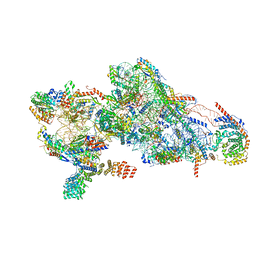 
 | | Human mitochondrial small subunit assembly intermediate (State E) | | Descriptor: | 12S mitochondrial rRNA, 28S ribosomal protein S10, mitochondrial, ... | | Authors: | Harper, N.J, Burnside, C, Klinge, S. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-14 | | Last modified: | 2023-02-08 | | Method: | ELECTRON MICROSCOPY (2.85 Å) | | Cite: | Principles of mitoribosomal small subunit assembly in eukaryotes.
Nature, 614, 2023
|
|
4UFY
 
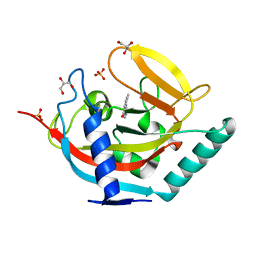 | | Crystal structure of human tankyrase 2 in complex with TA-13 | | Descriptor: | 8-METHYL-2-(4-METHYLPHENYL)-3,4-DIHYDROQUINAZOLIN-4-ONE, GLYCEROL, SULFATE ION, ... | | Authors: | Haikarainen, T, Lehtio, L. | | Deposit date: | 2015-03-20 | | Release date: | 2016-04-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-Activity Relationships of 2-Arylquinazolin-4-Ones as Highly Selective and Potent Inhibitors of the Tankyrases.
Eur.J.Med.Chem., 118, 2016
|
|
4UHG
 
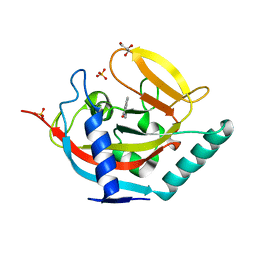 | | Crystal structure of human tankyrase 2 in complex with TA-21 | | Descriptor: | 2-(4-METHYLPHENYL)-3,4-DIHYDROQUINAZOLIN-4-ONE, GLYCEROL, SULFATE ION, ... | | Authors: | Haikarainen, T, Lehtio, L. | | Deposit date: | 2015-03-24 | | Release date: | 2016-04-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-Activity Relationships of 2-Arylquinazolin-4-Ones as Highly Selective and Potent Inhibitors of the Tankyrases.
Eur.J.Med.Chem., 118, 2016
|
|
7Z4E
 
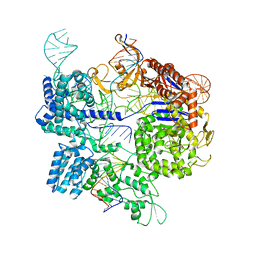 | | SpCas9 bound to 8-nucleotide complementary DNA substrate | | Descriptor: | CRISPR-associated endonuclease Cas9/Csn1, Non-target strand of 8 nucleotide complementary DNA substrate, Target strand of 8 nucleotide complementary DNA substrate, ... | | Authors: | Pacesa, M, Jinek, M. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-31 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (4.14 Å) | | Cite: | R-loop formation and conformational activation mechanisms of Cas9.
Nature, 609, 2022
|
|
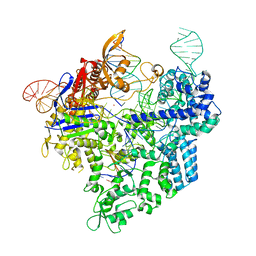
7Z4I
 
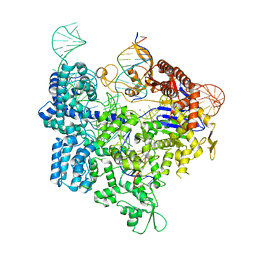 | | SpCas9 bound to 16-nucleotide complementary DNA substrate | | Descriptor: | CRISPR-associated endonuclease Cas9/Csn1, Non-target strand of 16-nucleotide complementary DNA substrate, POTASSIUM ION, ... | | Authors: | Pacesa, M, Jinek, M. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-31 | | Last modified: | 2024-07-17 | | Method: | ELECTRON MICROSCOPY (3.12 Å) | | Cite: | R-loop formation and conformational activation mechanisms of Cas9.
Nature, 609, 2022
|
|
7Z4L
 
 | |
4UI6
 
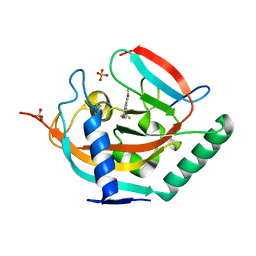 | | Crystal structure of human tankyrase 2 in complex with TA-47 | | Descriptor: | 8-METHOXY-2-[4-(TRIFLUOROMETHYL)PHENYL]-3,4-DIHYDROQUINAZOLIN-4-ONE, GLYCEROL, SULFATE ION, ... | | Authors: | Haikarainen, T, Lehtio, L. | | Deposit date: | 2015-03-27 | | Release date: | 2016-04-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure-Activity Relationships of 2-Arylquinazolin-4-Ones as Highly Selective and Potent Inhibitors of the Tankyrases.
Eur.J.Med.Chem., 118, 2016
|
|
