5FKY
 
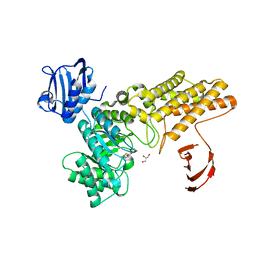 | | Structure of a hydrolase bound with an inhibitor | | Descriptor: | (3aR,5R,6S,7R,7aR)-2-amino-5-(hydroxymethyl)-5,6,7,7a-tetrahydro-3aH-pyrano[3,2-d][1,3]thiazole-6,7-diol, GLYCEROL, O-GLCNACASE BT_4395 | | Authors: | Cekic, N, Heinonen, J.E, Stubbs, K.A, Roth, C, McEachern, E.J, Davies, G.J, Vocadlo, D.J. | | Deposit date: | 2015-10-20 | | Release date: | 2016-01-27 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Analysis of transition state mimicry by tight binding aminothiazoline inhibitors provides insight into catalysis by humanO-GlcNAcase.
Chem Sci, 7, 2016
|
|
5FL1
 
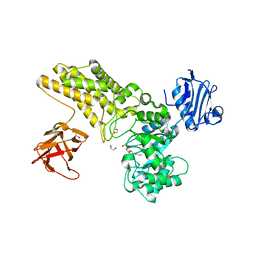 | | Structure of a hydrolase with an inhibitor | | Descriptor: | (3~{a}~{R},5~{R},6~{S},7~{R},7~{a}~{R})-5-(hydroxymethyl)-2-(prop-2-enylamino)-5,6,7,7~{a}-tetrahydro-3~{a}~{H}-pyrano[3,2-d][1,3]thiazole-6,7-diol, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Cekic, N, Heinonen, J.E, Stubbs, K.A, Roth, C, McEachern, E.J, Davies, G.J, Vocadlo, D.J. | | Deposit date: | 2015-10-20 | | Release date: | 2016-08-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Analysis of transition state mimicry by tight binding aminothiazoline inhibitors provides insight into catalysis by humanO-GlcNAcase.
Chem Sci, 7, 2016
|
|
5FL0
 
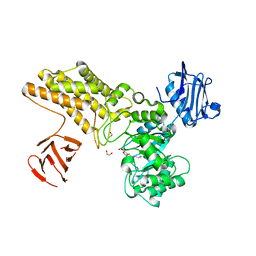 | | Structure of a hydrolase with an inhibitor | | Descriptor: | (3~{a}~{R},5~{R},6~{S},7~{R},7~{a}~{R})-2-(butylamino)-5-(hydroxymethyl)-5,6,7,7~{a}-tetrahydro-3~{a}~{H}-pyrano[3,2-d] [1,3]thiazole-6,7-diol, 1,2-ETHANEDIOL, CALCIUM ION, ... | | Authors: | Cekic, N, Heinonen, J.E, Stubbs, K.A, Roth, C, McEachern, E.J, Davies, G.J, Vocadlo, D.J. | | Deposit date: | 2015-10-20 | | Release date: | 2016-08-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Analysis of transition state mimicry by tight binding aminothiazoline inhibitors provides insight into catalysis by humanO-GlcNAcase.
Chem Sci, 7, 2016
|
|
5OXD
 
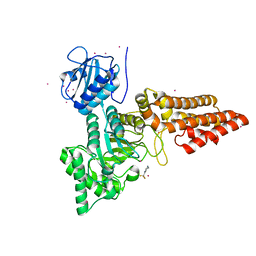 | |
2WB5
 
 | | GlcNAcstatins are nanomolar inhibitors of human O-GlcNAcase inducing cellular hyper-O-GlcNAcylation | | Descriptor: | (5R,6R,7R,8S)-6,7-dihydroxy-5-(hydroxymethyl)-2-(2-phenylethyl)-8-(propanoylamino)-5,6,7,8-tetrahydro-1H-imidazo[1,2-a]pyridin-4-ium, CHLORIDE ION, O-GLCNACASE NAGJ, ... | | Authors: | Dorfmueller, H.C, Borodkin, V.S, Schimpl, M, van Aalten, D.M.F. | | Deposit date: | 2009-02-20 | | Release date: | 2009-03-31 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Glcnacstatins are Nanomolar Inhibitors of Human O-Glcnacase Inducing Cellular Hyper-O-Glcnacylation
Biochem.J., 420, 2009
|
|
2XJ7
 
 | | BtGH84 in complex with 6-acetamido-6-deoxy-castanospermine | | Descriptor: | 6-ACETAMIDO-6-DEOXY-CASTANOSPERMINE, CALCIUM ION, O-GLCNACASE BT_4395 | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2010-07-02 | | Release date: | 2010-09-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Inhibition of O-Glcnacase Using a Potent and Cell-Permeable Inhibitor Does not Induce Insulin Resistance in 3T3-L1 Adipocytes.
Chem.Biol., 17, 2010
|
|
2X0Y
 
 | | Screening-based discovery of drug-like O-GlcNAcase inhibitor scaffolds | | Descriptor: | 7-[(2S)-2,3-DIHYDROXYPROPYL]-1,3-DIMETHYL-3,7-DIHYDRO-1H-PURINE-2,6-DIONE, O-GLCNACASE NAGJ | | Authors: | Dorfmueller, H.C, van Aalten, D.M.F. | | Deposit date: | 2009-12-18 | | Release date: | 2010-01-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Screening-Based Discovery of Drug-Like O-Glcnacase Inhibitor Scaffolds
FEBS Lett., 584, 2010
|
|
2XM2
 
 | | BtGH84 in complex with LOGNAc | | Descriptor: | GLYCEROL, N-acetylglucosaminono-1,5-lactone (Z)-oxime, O-GLCNACASE BT_4395 | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2010-07-22 | | Release date: | 2011-08-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Inhibition of a Bacterial O-Glcnacase Homologue by Lactone and Lactam Derivatives: Structural, Kinetic and Thermodynamic Analyses.
Amino Acids, 40, 2011
|
|
2XM1
 
 | | BtGH84 in complex with N-acetyl gluconolactam | | Descriptor: | GLYCEROL, N-ACETYL GLUCONOLACTAM, O-GLCNACASE BT_4395 | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2010-07-22 | | Release date: | 2011-08-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Inhibition of a Bacterial O-Glcnacase Homologue by Lactone and Lactam Derivatives: Structural, Kinetic and Thermodynamic Analyses.
Amino Acids, 40, 2011
|
|
2XPK
 
 | | Cell-penetrant, nanomolar O-GlcNAcase inhibitors selective against lysosomal hexosaminidases | | Descriptor: | N-[(5R,6R,7R,8S)-6,7-DIHYDROXY-5-(HYDROXYMETHYL)-2-(2-PHENYLETHYL)-5,6,7,8-TETRAHYDROIMIDAZO[1,2-A]PYRIDIN-8-YL]-3-SULFANYLPROPANAMIDE, O-GLCNACASE NAGJ | | Authors: | Dorfmueller, H.C, Borodkin, V.S, Schimpl, M, Zheng, X, Kime, R, Read, K.D, van Aalten, D.M.F. | | Deposit date: | 2010-08-26 | | Release date: | 2011-03-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Cell-Penetrant, Nanomolar O-Glcnacase Inhibitors Selective Against Lysosomal Hexosaminidases.
Chem.Biol, 17, 2010
|
|
2WZH
 
 | | BtGH84 D242N in complex with MeUMB-derived oxazoline | | Descriptor: | 2-METHYL-4,5-DIHYDRO-(1,2-DIDEOXY-ALPHA-D-GLUCOPYRANOSO)[2,1-D]-1,3-OXAZOLE, CALCIUM ION, GLYCEROL, ... | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2009-11-30 | | Release date: | 2010-01-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Visualizing the Reaction Coordinate of an O-Glcnac Hydrolase
J.Am.Chem.Soc., 132, 2010
|
|
2X0H
 
 | | BtGH84 Michaelis complex | | Descriptor: | 3,4-difluorophenyl 2-deoxy-2-[(difluoroacetyl)amino]-beta-D-glucopyranoside, CALCIUM ION, GLYCEROL, ... | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2009-12-08 | | Release date: | 2010-01-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Visualizing the Reaction Coordinate of an O-Glcnac Hydrolase
J.Am.Chem.Soc., 132, 2010
|
|
2WZI
 
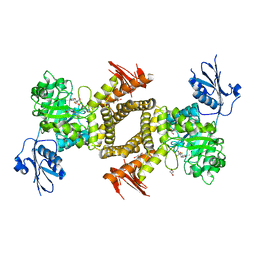 | | BtGH84 D243N in complex with 5F-oxazoline | | Descriptor: | (3AS,5S,6S,7R,7AR)-5-FLUORO-5-(HYDROXYMETHYL)-2-METHYL-5,6,7,7A-TETRAHYDRO-3AH-PYRANO[3,2-D][1,3]OXAZOLE-6,7-DIOL, CALCIUM ION, GLYCEROL, ... | | Authors: | He, Y, Davies, G.J. | | Deposit date: | 2009-11-30 | | Release date: | 2010-01-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Visualizing the Reaction Coordinate of an O-Glcnac Hydrolase
J.Am.Chem.Soc., 132, 2010
|
|
1R3E
 
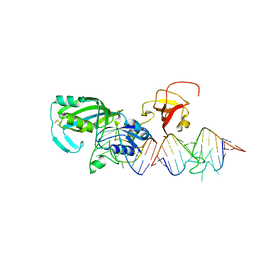 | | Crystal Structure of tRNA Pseudouridine Synthase TruB and Its RNA Complex: RNA-protein Recognition Through a Combination of Rigid Docking and Induced Fit | | Descriptor: | 5'-R(*CP*UP*GP*UP*GP*UP*(FHU)P*CP*GP*AP*UP*CP*CP*AP*CP*AP*G)-3', 5'-R(*CP*UP*GP*UP*GP*UP*UP*CP*GP*AP*UP*CP*CP*AP*CP*AP*G)-3', tRNA pseudouridine synthase B | | Authors: | Pan, H, Agarwalla, S, Moustakas, D.T, Finer-Moore, J, Stroud, R.M. | | Deposit date: | 2003-10-01 | | Release date: | 2003-11-04 | | Last modified: | 2018-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of tRNA Pseudouridine Synthase TruB and Its RNA Complex: RNA Recognition Through a Combination of Rigid Docking and Induced Fit
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
5LAD
 
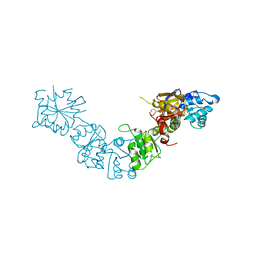 | |
6Q44
 
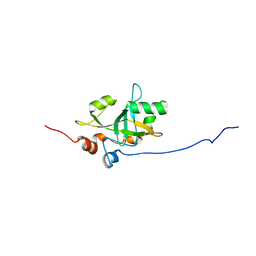 | | Est3 telomerase subunit in the yeast Hansenula polymorpha | | Descriptor: | Uncharacterized protein | | Authors: | Mantsyzov, A.B, Mariasina, S.S, Petrova, O.A, Efimov, S.V, Dontsova, O.A, Polshakov, V.I. | | Deposit date: | 2018-12-05 | | Release date: | 2019-12-25 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Insights into the structure and function of Est3 from the Hansenula polymorpha telomerase.
Sci Rep, 10, 2020
|
|
2AZL
 
 | | Crystal structure for the mutant F117E of Thermotoga maritima octaprenyl pyrophosphate synthase | | Descriptor: | octoprenyl-diphosphate synthase | | Authors: | Sun, H.Y, Ko, T.P, Kuo, C.J, Guo, R.T, Chou, C.C, Liang, P.H, Wang, A.H. | | Deposit date: | 2005-09-12 | | Release date: | 2006-03-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Homodimeric hexaprenyl pyrophosphate synthase from the thermoacidophilic crenarchaeon Sulfolobus solfataricus displays asymmetric subunit structures
J.Bacteriol., 187, 2005
|
|
2H2I
 
 | | The Structural basis of Sirtuin Substrate Affinity | | Descriptor: | (2S,5R,8R,11S,14S,17S,21R)-5,8,11,14,17-PENTAMETHYL-4,7,10,13,16,19-HEXAOXADOCOSANE-2,21-DIOL, NAD-dependent deacetylase, ZINC ION | | Authors: | Cosgrove, M.S, Wolberger, C. | | Deposit date: | 2006-05-18 | | Release date: | 2006-12-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structural basis of sirtuin substrate affinity
Biochemistry, 45, 2006
|
|
2KA0
 
 | | NMR structure of the protein TM1367 | | Descriptor: | uncharacterized protein TM1367 | | Authors: | Mohanty, B, Pedrini, B, Serrano, P, Geralt, M, Horst, R, Herrmann, T, Wilson, I.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-27 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures for the proteins TM1112 and TM1367.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
2H2H
 
 | |
2H2D
 
 | |
2H2F
 
 | |
2KA5
 
 | | NMR Structure of the protein TM1081 | | Descriptor: | Putative anti-sigma factor antagonist TM_1081 | | Authors: | Serrano, P, Geralt, M, Mohanty, B, Pedrini, B, Horst, R, Wuthrich, K, Wilson, I, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-30 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures highlights conformational isomerism in protein active sites.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
1WA3
 
 | | Mechanism of the Class I KDPG aldolase | | Descriptor: | 2-KETO-3-DEOXY-6-PHOSPHOGLUCONATE ALDOLASE, PYRUVIC ACID, SULFATE ION | | Authors: | Fullerton, S.W.B, Griffiths, J.S, Merkel, A.B, Wymer, N.J, Hutchins, M.J, Fierke, C.A, Toone, E.J, Naismith, J.H. | | Deposit date: | 2004-10-22 | | Release date: | 2005-01-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanism of the Class I Kdpg Aldolase.
Bioorg.Med.Chem., 14, 2006
|
|
1LW4
 | |
