4ZYH
 
 | |
2BOZ
 
 | | Photosynthetic Reaction Center Mutant With Gly M203 Replaced With Leu | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, DECANE, ... | | Authors: | Potter, J.P, Fyfe, P.K, Jones, M.R. | | Deposit date: | 2005-04-15 | | Release date: | 2005-05-20 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Strong Effects of an Individual Water Molecule on the Rate of Light-Driven Charge Separation in the Rhodobacter Sphaeroides Reaction Center.
J.Biol.Chem., 280, 2005
|
|
4WAM
 
 | |
2BHH
 
 | | HUMAN CYCLIN DEPENDENT PROTEIN KINASE 2 IN COMPLEX WITH THE INHIBITOR 4-HYDROXYPIPERINDINESULFONYL-INDIRUBINE | | Descriptor: | (2E,3S)-3-HYDROXY-5'-[(4-HYDROXYPIPERIDIN-1-YL)SULFONYL]-3-METHYL-1,3-DIHYDRO-2,3'-BIINDOL-2'(1'H)-ONE, CELL DIVISION PROTEIN KINASE 2 | | Authors: | Schaefer, M, Jautelat, R, Brumby, T, Briem, H, Eisenbrand, G, Schwahn, S, Krueger, M, Luecking, U, Prien, O, Siemeister, G. | | Deposit date: | 2005-01-11 | | Release date: | 2005-03-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | From the Insoluble Dye Indirubin Towards Highly Active, Soluble Cdk2-Inhibitors
Chembiochem, 6, 2005
|
|
4UQ8
 
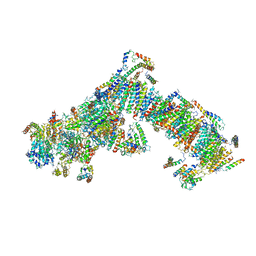 | | Electron cryo-microscopy of bovine Complex I | | Descriptor: | ACYL CARRIER PROTEIN, MITOCHONDRIAL, FE2/S2 (INORGANIC) CLUSTER, ... | | Authors: | Vinothkumar, K.R, Zhu, J, Hirst, J. | | Deposit date: | 2014-06-21 | | Release date: | 2014-10-01 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (4.95 Å) | | Cite: | Architecture of Mammalian Respiratory Complex I.
Nature, 515, 2014
|
|
4ZYG
 
 | |
8CWW
 
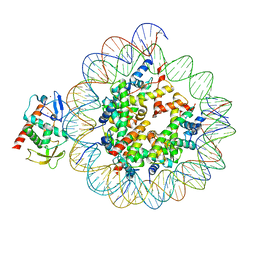 | | Structure of S. cerevisiae Hop1 CBR bound to a nucleosome | | Descriptor: | Histone H2A, Histone H2B, Histone H3, ... | | Authors: | Gu, Y, Ur, S.N, Milano, C.R, Tromer, E.C, Vale-Silva, L.A, Hochwagen, A, Corbett, K.D. | | Deposit date: | 2022-05-19 | | Release date: | 2023-06-07 | | Last modified: | 2024-04-10 | | Method: | ELECTRON MICROSCOPY (2.74 Å) | | Cite: | Chromatin binding by HORMAD proteins regulates meiotic recombination initiation.
Embo J., 43, 2024
|
|
8CZE
 
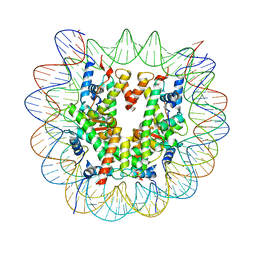 | | Structure of a Xenopus Nucleosome with Widom 601 DNA | | Descriptor: | Histone H2A, Histone H2B, Histone H3, ... | | Authors: | Gu, Y, Ur, S.N, Milano, C.R, Tromer, E.C, Vale-Silva, L.A, Hochwagen, A, Corbett, K.D. | | Deposit date: | 2022-05-24 | | Release date: | 2023-06-07 | | Last modified: | 2024-04-10 | | Method: | ELECTRON MICROSCOPY (2.58 Å) | | Cite: | Chromatin binding by HORMAD proteins regulates meiotic recombination initiation.
Embo J., 43, 2024
|
|
4UST
 
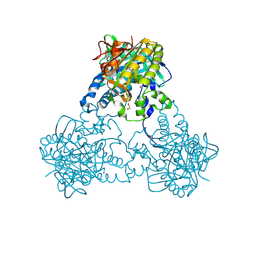 | |
3V7Q
 
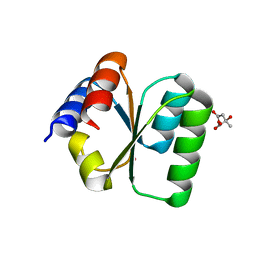 | | Crystal structure of B. subtilis YlxQ at 1.55 A resolution | | Descriptor: | CITRIC ACID, POTASSIUM ION, Probable ribosomal protein ylxQ | | Authors: | Baird, N.J, Zhang, J, Hamma, T, Ferre-D'Amare, A.R. | | Deposit date: | 2011-12-21 | | Release date: | 2012-03-07 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | YbxF and YlxQ are bacterial homologs of L7Ae and bind K-turns but not K-loops.
Rna, 18, 2012
|
|
4WFM
 
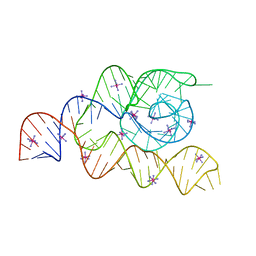 | | Structure of the complete bacterial SRP Alu domain | | Descriptor: | Bacillus subtilis small cytoplasmic RNA (scRNA),RNA, COBALT HEXAMMINE(III), MAGNESIUM ION | | Authors: | Kempf, G, Wild, K, Sinning, I. | | Deposit date: | 2014-09-15 | | Release date: | 2014-10-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure of the complete bacterial SRP Alu domain.
Nucleic Acids Res., 42, 2014
|
|
4USV
 
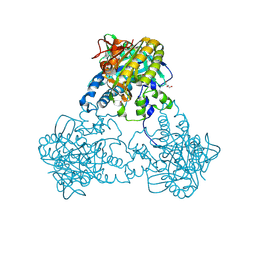 | |
4USW
 
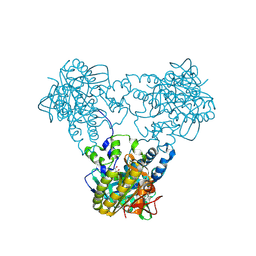 | | Crystal structure of human soluble Adenylyl Cyclase with ATP | | Descriptor: | ACETATE ION, ADENOSINE-5'-TRIPHOSPHATE, ADENYLATE CYCLASE TYPE 10, ... | | Authors: | Kleinboelting, S, Steegborn, C. | | Deposit date: | 2014-07-13 | | Release date: | 2014-07-30 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural Analysis of Human Soluble Adenylyl Cyclase and Crystal Structures of its Nucleotide Complexes -Implications for Cyclase Catalysis and Evolution.
FEBS J., 281, 2014
|
|
7M3T
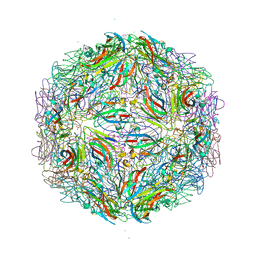 
 | |
7M50
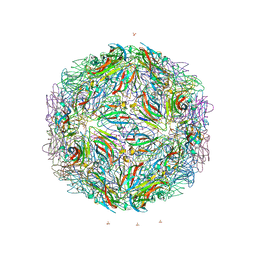 
 | |
5AJ0
 
 | | Cryo electron microscopy of actively translating human polysomes (POST state). | | Descriptor: | 18S ribosomal RNA, 28S ribosomal RNA, 40S ribosomal protein S10, ... | | Authors: | Behrmann, E, Loerke, J, Budkevich, T.V, Yamamoto, K, Schmidt, A, Penczek, P.A, Vos, M.R, Burger, J, Mielke, T, Scheerer, P, Spahn, C.M.T. | | Deposit date: | 2015-02-19 | | Release date: | 2015-05-20 | | Last modified: | 2019-12-18 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural Snapshots of Actively Translating Human Ribosomes
Cell(Cambridge,Mass.), 161, 2015
|
|
6CIT
 
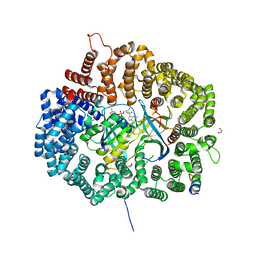 | |
6CF1
 
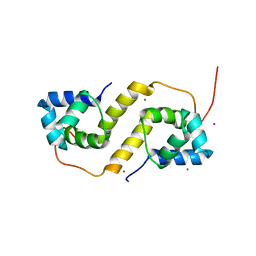 | | Proteus vulgaris HigA antitoxin structure | | Descriptor: | Antitoxin HigA, POTASSIUM ION | | Authors: | Schureck, M.A, Hoffer, E.D, Ei Cho, S, Dunham, C.M. | | Deposit date: | 2018-02-13 | | Release date: | 2019-02-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis of transcriptional regulation by the HigA antitoxin.
Mol.Microbiol., 111, 2019
|
|
7AT8
 
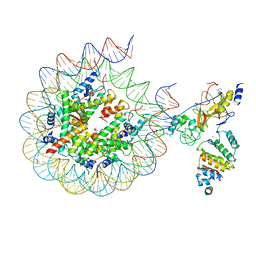 | | Histone H3 recognition by nucleosome-bound PRC2 subunit EZH2. | | Descriptor: | Histone H2A, Histone H2B 1.1, Histone H3.2, ... | | Authors: | Finogenova, K, Benda, C, Schaefer, I.B, Poepsel, S, Strauss, M, Mueller, J. | | Deposit date: | 2020-10-29 | | Release date: | 2020-12-09 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structural basis for PRC2 decoding of active histone methylation marks H3K36me2/3.
Elife, 9, 2020
|
|
6PHR
 
 | |
7B06
 
 | | TgoT_RT521 apo | | Descriptor: | DNA polymerase | | Authors: | Samson, C, Legrand, P, Tekpinar, M, Rozenski, J, Abramov, M, Holliger, P, Pinheiro, V, Herdewijn, P, Delarue, M. | | Deposit date: | 2020-11-18 | | Release date: | 2020-12-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.349 Å) | | Cite: | Structural Studies of HNA Substrate Specificity in Mutants of an Archaeal DNA Polymerase Obtained by Directed Evolution.
Biomolecules, 10, 2020
|
|
6PI1
 
 | |
6W3Z
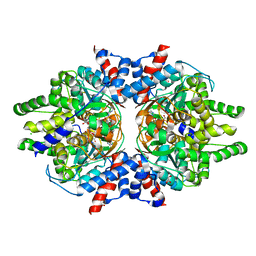 
 | | Crystal Structure of Brugia malayi Deoxyhypusine synthase (DHPS) | | Descriptor: | BMA-DHPS-1, isoform a, CHLORIDE ION, ... | | Authors: | Santiago, A.S, dos Reis, C.V, Ramos, P.Z, Klippel, H.A, Silva, S.F, Zanelli, C.F, Massirer, K.B, Arruda, P, Edwards, A.M, Counago, R.M, Structural Genomics Consortium (SGC) | | Deposit date: | 2020-03-09 | | Release date: | 2020-11-04 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural features and development of an assay platform of the parasite target deoxyhypusine synthase of Brugia malayi and Leishmania major.
Plos Negl Trop Dis, 14, 2020
|
|
6CZL
 
 | |
6CB5
 
 | |
