2LSC
 
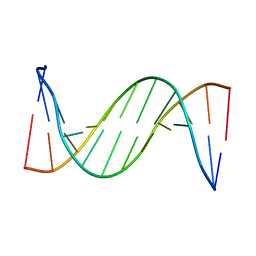 | | Solution structure of 2'F-ANA and ANA self-complementary duplex | | Descriptor: | DNA (5'-D(*(CFL)P*(GFL)P*(CFL)P*(GFL)P*(A5O)P*(A5O)P*(UAR)P*(UAR)P*(CFL)P*(GFL)P*(CFL)P*(GFL))-3') | | Authors: | Martin-Pintado, N, Yahyaee, M, Campos, R, Noronha, A, Wilds, C, Damha, M, Gonzalez, C. | | Deposit date: | 2012-04-26 | | Release date: | 2012-07-18 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The solution structure of double helical arabino nucleic acids (ANA and 2'F-ANA): effect of arabinoses in duplex-hairpin interconversion.
Nucleic Acids Res., 40, 2012
|
|
1B5K
 
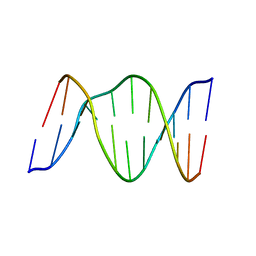 | | 3,N4-ETHENO-2'-DEOXYCYTIDINE OPPOSITE THYMIDINE IN AN 11-MER DUPLEX, SOLUTION STRUCTURE FROM NMR AND MOLECULAR DYNAMICS | | Descriptor: | DNA (5'-D(*CP*GP*TP*AP*CP*EDCP*CP*AP*TP*GP*C)-3'), DNA (5'-D(*GP*CP*AP*TP*GP*TP*GP*TP*AP*CP*G)-3') | | Authors: | Cullinan, D, Korobka, A, Grollman, A.P, Patel, D.J, Eisenberg, M, De Santos, C.L. | | Deposit date: | 1999-01-07 | | Release date: | 1999-01-13 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of an oligodeoxynucleotide duplex containing the exocyclic lesion 3,N4-etheno-2'-deoxycytidine opposite thymidine: comparison with the duplex containing deoxyadenosine opposite the adduct.
Biochemistry, 35, 1996
|
|
5YEG
 
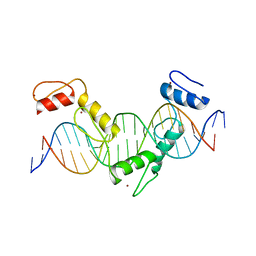 | | Crystal structure of CTCF ZFs4-8-Hs5-1a complex | | Descriptor: | DNA (5'-D(*AP*CP*TP*TP*TP*AP*AP*CP*CP*AP*GP*CP*AP*GP*AP*GP*GP*GP*CP*G)-3'), DNA (5'-D(*TP*CP*GP*CP*CP*CP*TP*CP*TP*GP*CP*TP*GP*GP*TP*TP*AP*AP*AP*G)-3'), Transcriptional repressor CTCF, ... | | Authors: | Yin, M, Wang, J, Wang, M, Li, X. | | Deposit date: | 2017-09-17 | | Release date: | 2017-11-29 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular mechanism of directional CTCF recognition of a diverse range of genomic sites
Cell Res., 27, 2017
|
|
2N9Q
 
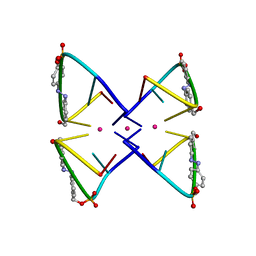 | | Photoswitchable G-quadruplex | | Descriptor: | DNA (5'-D(*GP*GP*(AZW)P*GP*G)-3'), POTASSIUM ION | | Authors: | Thevarpadam, J, Bessi, I, Binas, O, Goncalves, D.P.N, Slavov, C, Jonker, H.R.A, Richter, C, Wachtveitl, J, Schwalbe, H, Heckel, A. | | Deposit date: | 2015-12-02 | | Release date: | 2016-02-17 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Photoresponsive Formation of an Intermolecular Minimal G-Quadruplex Motif.
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
4B3G
 
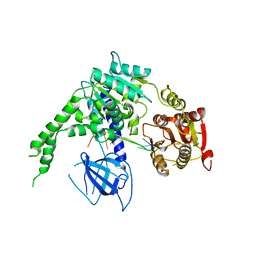 | | crystal structure of Ighmbp2 helicase in complex with RNA | | Descriptor: | DNA-BINDING PROTEIN SMUBP-2, RNA (5'-(AP*AP*AP*AP*AP*AP*AP*AP*AP)-3') | | Authors: | Lim, S.C, Song, H. | | Deposit date: | 2012-07-24 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | The Ighmbp2 Helicase Structure Reveals the Molecular Basis for Disease-Causing Mutations in Dmsa1.
Nucleic Acids Res., 40, 2012
|
|
7BWD
 
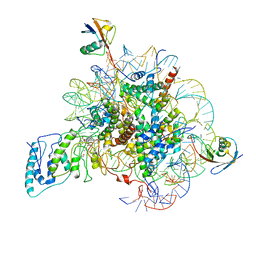 | | Structure of Dot1L-H2BK34ub Nucleosome Complex | | Descriptor: | DNA (146-MER), Histone H2A type 1-B/E, Histone H2B type 1-K, ... | | Authors: | Lou, Z.Y, Liu, L, Cao, L, Ai, H.S, Sun, Z.X. | | Deposit date: | 2020-04-14 | | Release date: | 2021-04-14 | | Method: | ELECTRON MICROSCOPY (4.32 Å) | | Cite: | Structure of the Dot1L-H2BK34ub Nucleosome Complex Reveals an Unprecedented Crosstalk Activation Mechanism through Nucleosome Shape Distortion
To Be Published
|
|
4B3F
 
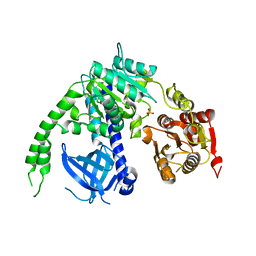 | | crystal structure of Ighmbp2 helicase | | Descriptor: | DNA-BINDING PROTEIN SMUBP-2, PHOSPHATE ION | | Authors: | Lim, S.C, Song, H. | | Deposit date: | 2012-07-24 | | Release date: | 2012-09-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Ighmbp2 Helicase Structure Reveals the Molecular Basis for Disease-Causing Mutations in Dmsa1.
Nucleic Acids Res., 40, 2012
|
|
1WI3
 
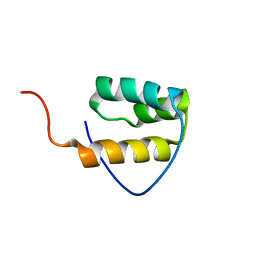 | | Solution structure of the homeodomain of KIAA1034 protein | | Descriptor: | DNA-binding protein SATB2 | | Authors: | Izumi, K, Yoshida, M, Hayashi, F, Hatta, R, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-05-28 | | Release date: | 2004-11-28 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the homeodomain of KIAA1034 protein
TO BE PUBLISHED
|
|
1NR3
 
 | | SOLUTION STRUCTURE OF THE PROTEIN MTH0916: THE NORTHEAST STRUCTURAL GENOMICS CONSORTIUM TARGET TT212 | | Descriptor: | DNA-binding protein tfx | | Authors: | Shen, Y, Liu, G, Bhaskaran, R, Yee, A, Arrowsmith, C, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2003-01-23 | | Release date: | 2003-06-10 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | SOLUTION STRUCTURE OF THE PROTEIN MTH0916: THE NORTHEAST STRUCTURAL GENOMICS CONSORTIUM TARGET TT212
To be Published
|
|
3PGW
 
 | | Crystal structure of human U1 snRNP | | Descriptor: | DNA 5'-D(*AP*GP*GP*TP*AP*AP*GP*TP*A)-3', Sm B, Sm G, ... | | Authors: | Weber, G, Trowitzsch, S, Kastner, B, Luehrmann, R, Wahl, M.C. | | Deposit date: | 2010-11-02 | | Release date: | 2010-12-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (4.4 Å) | | Cite: | Functional organization of the Sm core in the crystal structure of human U1 snRNP.
Embo J., 29, 2010
|
|
3TCI
 | |
3LPX
 
 | | Crystal structure of GyrA | | Descriptor: | DNA gyrase, A subunit | | Authors: | Jung, H.Y, Heo, Y.S. | | Deposit date: | 2010-02-07 | | Release date: | 2011-02-16 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of GyrA
To be Published
|
|
3TU4
 
 | | Crystal structure of the Sir3 BAH domain in complex with a nucleosome core particle. | | Descriptor: | DNA (146-MER), Histone H2A, Histone H2B 1.1, ... | | Authors: | Armache, K.-J, Garlick, J.D, Canzio, D, Narlikar, G.J, Kingston, R.E. | | Deposit date: | 2011-09-15 | | Release date: | 2011-11-23 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis of silencing: Sir3 BAH domain in complex with a nucleosome at 3.0 A resolution.
Science, 334, 2011
|
|
6XLX
 
 | |
5LM5
 
 | | Structure of C-terminal domain from S. cerevisiae Pat1 decapping activator bound to Dcp2 HLM2 peptide (region 435-451) | | Descriptor: | DNA topoisomerase 2-associated protein PAT1, mRNA decapping protein 2 | | Authors: | Charenton, C, Gaudon-Plesse, C, Fourati, Z, Taverniti, V, Back, R, Kolesnikova, O, Seraphin, B, Graille, M. | | Deposit date: | 2016-07-29 | | Release date: | 2017-08-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A unique surface on Pat1 C-terminal domain directly interacts with Dcp2 decapping enzyme and Xrn1 5'-3' mRNA exonuclease in yeast.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
6OQM
 
 | | crystal structure of the MSH6 PWWP domain | | Descriptor: | DNA mismatch repair protein Msh6, MAGNESIUM ION, SULFATE ION, ... | | Authors: | Qin, S, Tempel, W, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium, Structural Genomics Consortium (SGC) | | Deposit date: | 2019-04-26 | | Release date: | 2019-05-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | crystal structure of the MSH6 PWWP domain
To Be Published
|
|
8PSE
 
 | |
8Q7D
 
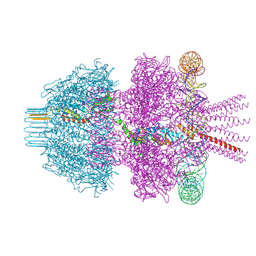 | | Neck of phage 812 after tail contraction (C12) | | Descriptor: | DNA forward strand (120-MER), DNA reverse strand (120-MER), Portal protein, ... | | Authors: | Cienikova, Z, Siborova, M, Fuzik, T, Plevka, P. | | Deposit date: | 2023-08-16 | | Release date: | 2024-08-28 | | Method: | ELECTRON MICROSCOPY (3.07 Å) | | Cite: | Genome anchoring, retention, and release by neck proteins of Herelleviridae phage 812
To Be Published
|
|
6XLV
 
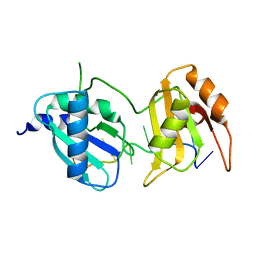 | |
2AUJ
 
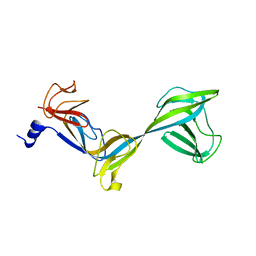 | | Structure of Thermus aquaticus RNA polymerase beta'-subunit insert | | Descriptor: | DNA-directed RNA polymerase beta' chain | | Authors: | Chlenov, M, Masuda, S, Murakami, K.S, Nikiforov, V, Darst, S.A, Mustaev, A. | | Deposit date: | 2005-08-27 | | Release date: | 2005-10-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure and Function of Lineage-specific Sequence Insertions in the Bacterial RNA Polymerase beta' Subunit
J.Mol.Biol., 353, 2005
|
|
2KOW
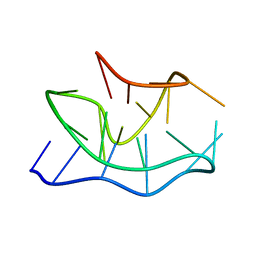 
 | |
2AUK
 
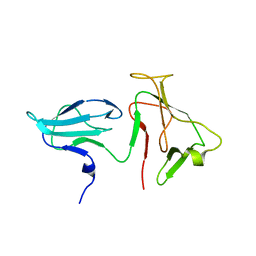 | | Structure of E. coli RNA polymerase beta' G/G' insert | | Descriptor: | DNA-directed RNA polymerase beta' chain | | Authors: | Chlenov, M, Masuda, S, Murakami, K.S, Nikiforov, V, Darst, S.A, Mustaev, A. | | Deposit date: | 2005-08-28 | | Release date: | 2005-10-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure and Function of Lineage-specific Sequence Insertions in the Bacterial RNA Polymerase beta' Subunit
J.Mol.Biol., 353, 2005
|
|
2DK8
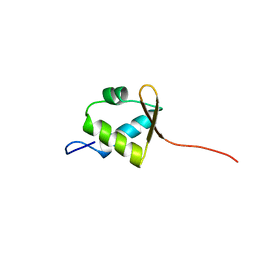 
 | | Solution structure of rpc34 subunit in RNA polymerase III from mouse | | Descriptor: | DNA-directed RNA polymerase III 39 kDa polypeptide | | Authors: | He, F, Muto, Y, Inoue, M, Kigawa, T, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-04-06 | | Release date: | 2006-10-06 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of rpc34 subunit in RNA polymerase III from mouse
To be Published
|
|
5UMV
 
 | |
6RNL
 
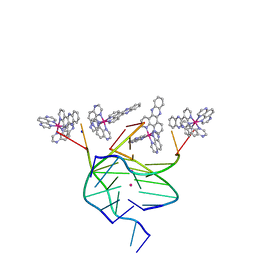 | | L-[Ru(TAP)2(dppz)]2+ bound to the G-quadruplex forming sequence d(TAGGGTT) | | Descriptor: | DNA (5'-D(*TP*AP*GP*GP*GP*TP*T)-3'), POTASSIUM ION, Ru(tap)2(dppz) complex, ... | | Authors: | McQuaid, K.T, Hall, J.P, Cardin, C.J. | | Deposit date: | 2019-05-09 | | Release date: | 2019-08-28 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Three thymine/adenine binding modes of the ruthenium complex Lambda-[Ru(TAP)2(dppz)]2+to the G-quadruplex forming sequence d(TAGGGTT) shown by X-ray crystallography.
Chem.Commun.(Camb.), 55, 2019
|
|
