7WAY
 
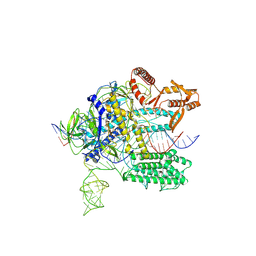 | |
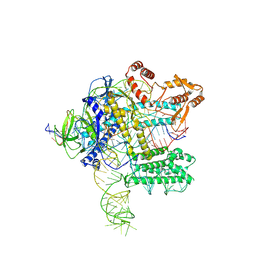
7WB1
 
 | |
4MIA
 
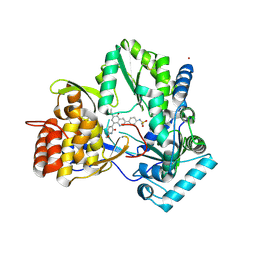 | | Hepatitis C Virus polymerase NS5B genotype 1b (BK) in complex with RG7109 (N-{4-[6-tert-butyl-5-methoxy-8-(6-methoxy-2-oxo-2,5-dihydropyridin-3-yl)quinolin-3-yl]phenyl}methanesulfonamide) | | Descriptor: | N-{4-[6-tert-butyl-5-methoxy-8-(6-methoxy-2-oxo-2,5-dihydropyridin-3-yl)quinolin-3-yl]phenyl}methanesulfonamide, RNA-directed RNA polymerase, ZINC ION | | Authors: | Harris, S.F, Villasenor, A.G. | | Deposit date: | 2013-08-30 | | Release date: | 2014-05-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Discovery of N-[4-[6-tert-butyl-5-methoxy-8-(6-methoxy-2-oxo-1H-pyridin-3-yl)-3-quinolyl]phenyl]methanesulfonamide (RG7109), a potent inhibitor of the hepatitis C virus NS5B polymerase.
J.Med.Chem., 57, 2014
|
|
4MON
 
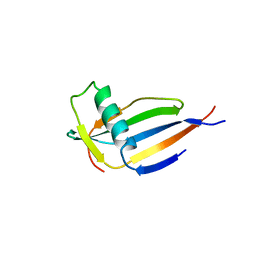 | | ORTHORHOMBIC MONELLIN | | Descriptor: | MONELLIN | | Authors: | Bujacz, G, Wlodawer, A. | | Deposit date: | 1997-03-04 | | Release date: | 1997-07-07 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of monellin refined to 2.3 a resolution in the orthorhombic crystal form.
Acta Crystallogr.,Sect.D, 53, 1997
|
|
7WLW
 
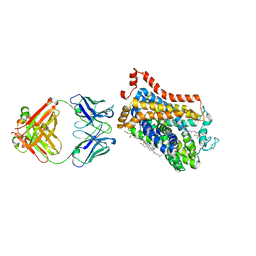 | | X-ray structure of thermostabilized Drosophila dopamine transporter with GABA transporter1-like substitutions in the binding site, in complex with SKF89976a | | Descriptor: | (3S)-1-(4,4-diphenylbut-3-enyl)piperidine-3-carboxylic acid, Antibody Fragment heavy chain, Antibody fragment light chain, ... | | Authors: | Penmatsa, A, Joseph, D. | | Deposit date: | 2022-01-13 | | Release date: | 2022-06-29 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural insights into GABA transport inhibition using an engineered neurotransmitter transporter.
Embo J., 41, 2022
|
|
7WGD
 
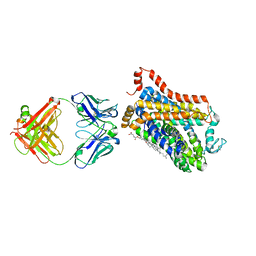 | |
4MKA
 
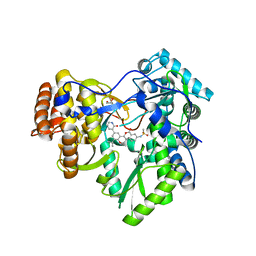 | | Hepatitis C Virus polymerase NS5B genotype 1b (BK) in complex with inhibitor 13 (N-{2-[3-tert-butyl-2-methoxy-5-(2-oxo-1,2-dihydropyridin-3-yl)phenyl]-1,3-benzoxazol-5-yl}methanesulfonamide) | | Descriptor: | DIMETHYL SULFOXIDE, GLYCEROL, N-{3-[3-tert-butyl-2-methoxy-5-(2-oxo-1,2-dihydropyridin-3-yl)phenyl]-1-oxo-1H-isochromen-7-yl}methanesulfonamide, ... | | Authors: | Harris, S.F, Wong, A. | | Deposit date: | 2013-09-04 | | Release date: | 2013-10-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Discovery of a Novel Series of Potent Non-Nucleoside Inhibitors of Hepatitis C Virus NS5B.
J.Med.Chem., 56, 2013
|
|
7WGT
 
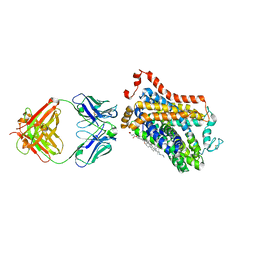 | | X-ray structure of thermostabilized Drosophila dopamine transporter with GABA transporter1-like substitutions in the binding site, in complex with NO711. | | Descriptor: | 1-(2-{[(diphenylmethylidene)amino]oxy}ethyl)-1,2,5,6-tetrahydropyridine-3-carboxylic acid, Antibody fragment (9D5) heavy chain, Antibody fragment (9D5) light chain, ... | | Authors: | Joseph, D, Penmatsa, A. | | Deposit date: | 2021-12-28 | | Release date: | 2022-06-29 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural insights into GABA transport inhibition using an engineered neurotransmitter transporter.
Embo J., 41, 2022
|
|
4MR1
 
 | |
4KXT
 
 | |
4LD1
 | |
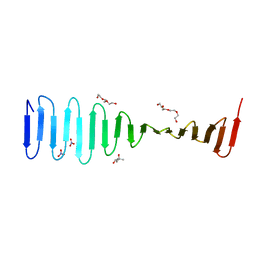
4LM5
 
 | | Crystal structure of Pim1 in complex with 2-{4-[(3-aminopropyl)amino]quinazolin-2-yl}phenol (resulting from displacement of SKF86002) | | Descriptor: | 2-{4-[(3-aminopropyl)amino]quinazolin-2-yl}phenol, GLYCEROL, Serine/threonine-protein kinase pim-1 | | Authors: | Parker, L.J, Tanaka, A, Handa, N, Honda, K, Tomabechi, Y, Shirouzu, M, Yokoyama, S. | | Deposit date: | 2013-07-10 | | Release date: | 2014-02-12 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Kinase crystal identification and ATP-competitive inhibitor screening using the fluorescent ligand SKF86002.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
7WAK
 
 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, Glutamyl-tRNA synthetase, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.782 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAL
 
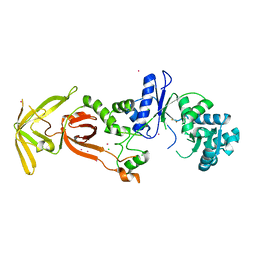 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with Co | | Descriptor: | BETA-MERCAPTOETHANOL, CHLORIDE ION, COBALT (II) ION, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAJ
 
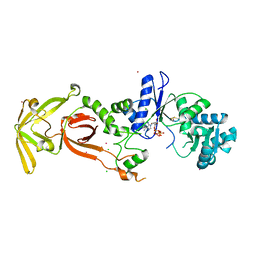 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) complexed with ATP and Co | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.254 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
7WAI
 
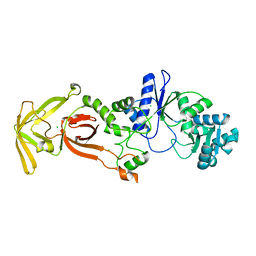 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) | | Descriptor: | CHLORIDE ION, Glutamyl-tRNA synthetase | | Authors: | Sharma, V, Manickam, Y, Harlos, K, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.105 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
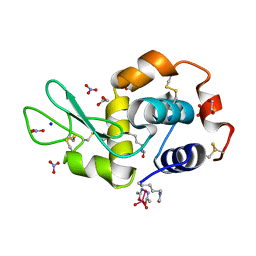
5NJ1
 
 | |
7WAO
 
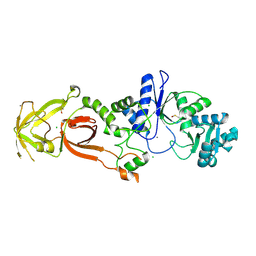 | | Glutamyl-tRNA synthetase from Plasmodium falciparum (PfERS) in complex with Mn | | Descriptor: | CHLORIDE ION, Glutamyl-tRNA synthetase, MANGANESE (II) ION | | Authors: | Sharma, V, Manickam, Y, Babbar, P, Sharma, A. | | Deposit date: | 2021-12-14 | | Release date: | 2022-12-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.582 Å) | | Cite: | Structural characterization of glutamyl-tRNA synthetase (GluRS) from Plasmodium falciparum.
Mol.Biochem.Parasitol., 253, 2022
|
|
5MJL
 
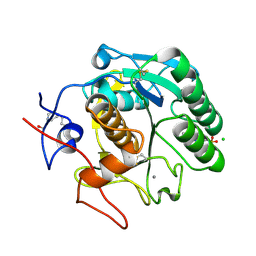 | | Single-shot pink beam serial crystallography: Proteinase K | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CALCIUM ION, ... | | Authors: | Meents, A, Oberthuer, D, Lieske, J, Srajer, V. | | Deposit date: | 2016-12-01 | | Release date: | 2017-11-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.21013784 Å) | | Cite: | Pink-beam serial crystallography.
Nat Commun, 8, 2017
|
|
7WZ8
 
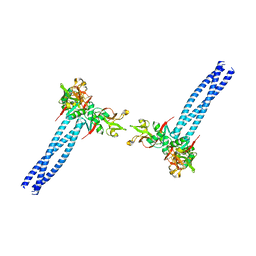 | | Structure of human langerin complex in Birbeck granules | | Descriptor: | SNAP-tag,C-type lectin domain family 4 member K | | Authors: | Oda, T, Yanagisawa, H. | | Deposit date: | 2022-02-17 | | Release date: | 2022-06-22 | | Last modified: | 2023-08-02 | | Method: | ELECTRON MICROSCOPY (6.4 Å) | | Cite: | Cryo-electron tomography of Birbeck granules reveals the molecular mechanism of langerin lattice formation.
Elife, 11, 2022
|
|
7XGO
 
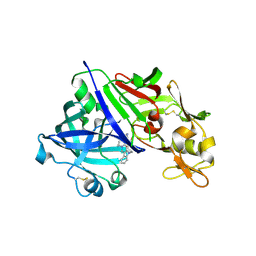 | | Human renin in complex with compound2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Renin, UNKNOWN LIGAND | | Authors: | Kashima, A. | | Deposit date: | 2022-04-05 | | Release date: | 2022-08-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Discovery of Novel 2-Carbamoyl Morpholine Derivatives as Highly Potent and Orally Active Direct Renin Inhibitors.
Acs Med.Chem.Lett., 13, 2022
|
|
7XGK
 
 | | Human renin in complex with compound1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Renin, UNKNOWN LIGAND | | Authors: | Kashima, A. | | Deposit date: | 2022-04-05 | | Release date: | 2022-08-31 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Discovery of Novel 2-Carbamoyl Morpholine Derivatives as Highly Potent and Orally Active Direct Renin Inhibitors.
Acs Med.Chem.Lett., 13, 2022
|
|
7XGP
 
 | | Human renin in complex with compound3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Renin, UNKNOWN LIGAND | | Authors: | Kashima, A. | | Deposit date: | 2022-04-05 | | Release date: | 2022-09-07 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Discovery of SPH3127: A Novel, Highly Potent, and Orally Active Direct Renin Inhibitor.
J.Med.Chem., 65, 2022
|
|
4MXV
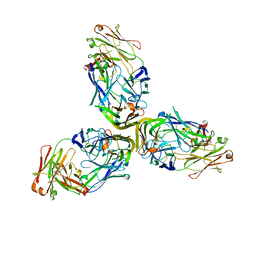 
 | | Structure of Lymphotoxin alpha bound to anti-LTa Fab | | Descriptor: | Lymphotoxin-alpha, anti-Lymphotoxin alpha antibody heavy chain, anti-Lymphotoxin alpha antibody light chain | | Authors: | Yin, J.P, Hymowitz, S.G. | | Deposit date: | 2013-09-26 | | Release date: | 2013-11-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Dimerization of LT beta R by LT alpha 1 beta 2 is necessary and sufficient for signal transduction.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
5NEN
 | |
