2IKB
 
 | |
2IKC
 
 | | Crystal structure of sheep lactoperoxidase at 3.25 A resolution reveals the binding sites for formate | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Sheikh, I.A, Singh, N, Singh, A.K, Sharma, S, Singh, T.P. | | Deposit date: | 2006-10-02 | | Release date: | 2006-10-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Crystal structure of sheep lactoperoxidase at 3.25 A resolution reveals the binding sites for formate
To be Published
|
|
2IKD
 
 | | Solution Structure of the first Clip domain in PAP2 | | Descriptor: | Prophenoloxidase activating proteinase-2 | | Authors: | Huang, R.D, Lv, Z.Q, Dai, H.E, Velde, D.V, Prakash, O, Jiang, H.B. | | Deposit date: | 2006-10-02 | | Release date: | 2007-10-16 | | Last modified: | 2020-09-09 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Clip domain in PAP2
To be Published
|
|
2IKE
 
 | | Solution Structure of the second Clip domain in PAP2 | | Descriptor: | Prophenoloxidase activating proteinase-2 | | Authors: | Huang, R.D, Lv, Z.Q, Dai, H.E, Velde, D.V, Prakash, O, Jiang, H.B. | | Deposit date: | 2006-10-02 | | Release date: | 2007-10-16 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Clip domain in PAP2
To be Published
|
|
2IKF
 
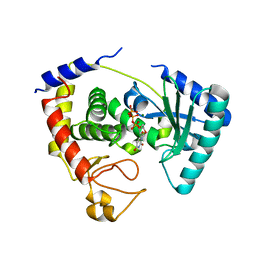 | |
2IKG
 
 | | Aldose reductase complexed with nitrophenyl-oxadiazol type inhibitor at 1.43 A | | Descriptor: | 4-[3-(3-NITROPHENYL)-1,2,4-OXADIAZOL-5-YL]BUTANOIC ACID, Aldose reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Steuber, H, Koch, C, Heine, A, Klebe, G. | | Deposit date: | 2006-10-02 | | Release date: | 2007-05-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Structural and Thermodynamic Study on Aldose Reductase: Nitro-substituted Inhibitors with Strong Enthalpic Binding Contribution
J.Mol.Biol., 368, 2007
|
|
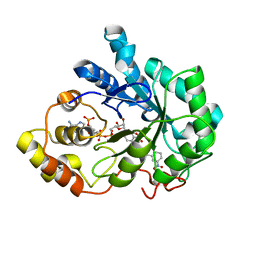
2IKH
 
 | | Human aldose reductase complexed with nitrofuryl-oxadiazol inhibitor at 1.55 A | | Descriptor: | Aldose reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, {[5-(5-NITRO-2-FURYL)-1,3,4-OXADIAZOL-2-YL]THIO}ACETIC ACID | | Authors: | Steuber, H, Koch, C, Heine, A, Klebe, G. | | Deposit date: | 2006-10-02 | | Release date: | 2007-05-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural and Thermodynamic Study on Aldose Reductase: Nitro-substituted Inhibitors with Strong Enthalpic Binding Contribution
J.Mol.Biol., 368, 2007
|
|
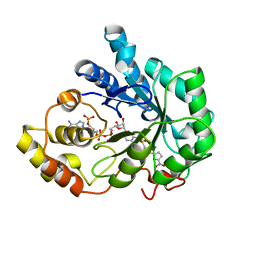
2IKI
 
 | | Human aldose reductase complexed with halogenated IDD-type inhibitor | | Descriptor: | (2-{[(4-BROMO-2-FLUOROBENZYL)AMINO]CARBONYL}-5-CHLOROPHENOXY)ACETIC ACID, Aldose reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Steuber, H, Koch, C, Heine, A, Klebe, G. | | Deposit date: | 2006-10-02 | | Release date: | 2007-05-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Structural and Thermodynamic Study on Aldose Reductase: Nitro-substituted Inhibitors with Strong Enthalpic Binding Contribution
J.Mol.Biol., 368, 2007
|
|
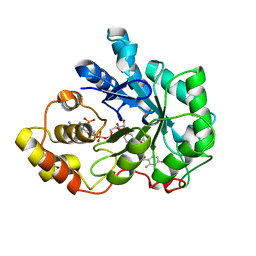
2IKJ
 
 | | Human aldose reductase complexed with nitro-substituted IDD-type inhibitor | | Descriptor: | (5-CHLORO-2-{[(3-NITROBENZYL)AMINO]CARBONYL}PHENOXY)ACETIC ACID, Aldose reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Steuber, H, Koch, C, Heine, A, Klebe, G. | | Deposit date: | 2006-10-02 | | Release date: | 2007-05-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural and Thermodynamic Study on Aldose Reductase: Nitro-substituted Inhibitors with Strong Enthalpic Binding Contribution
J.Mol.Biol., 368, 2007
|
|
2IKK
 
 | | Structural Genomics, the crystal structure of the C-terminal domain of Yurk from Bacillus subtilis subsp. subtilis str. 168 | | Descriptor: | Hypothetical transcriptional regulator yurK, SULFATE ION | | Authors: | Tan, K, Hatzos, C, Abdullah, J, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-10-02 | | Release date: | 2006-10-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The crystal structure of the C-terminal domain of Yurk from Bacillus subtilis subsp. subtilis str. 168
To be Published
|
|
2IKO
 
 | | Crystal Structure of Human Renin Complexed with Inhibitor | | Descriptor: | 5-{4-[(3,5-DIFLUOROBENZYL)AMINO]PHENYL}-6-ETHYLPYRIMIDINE-2,4-DIAMINE, Renin | | Authors: | Mochalkin, I. | | Deposit date: | 2006-10-02 | | Release date: | 2006-11-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Binding thermodynamics of substituted diaminopyrimidine renin inhibitors.
Anal.Biochem., 360, 2007
|
|
2IKQ
 
 | |
2IKS
 
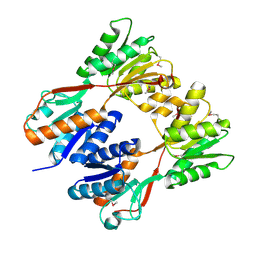 | | Crystal structure of N-terminal truncated DNA-binding transcriptional dual regulator from Escherichia coli K12 | | Descriptor: | DNA-binding transcriptional dual regulator | | Authors: | Chang, C, Evdokimova, E, Kagan, O, Savchenko, A, Edwards, A.M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-10-02 | | Release date: | 2006-10-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of N-terminal truncated DNA-binding transcriptional
dual regulator from Escherichia coli K12
To be Published
|
|
2IKU
 
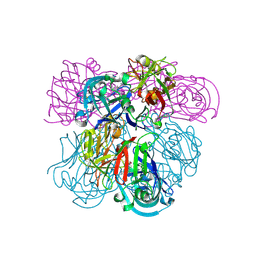 | | Crystal Structure of Human Renin Complexed with Inhibitors | | Descriptor: | 6-ETHYL-5-[(2S)-1-(3-METHOXYPROPYL)-2-PHENYL-1,2,3,4-TETRAHYDROQUINOLIN-7-YL]PYRIMIDINE-2,4-DIAMINE, Renin | | Authors: | Mochalkin, I. | | Deposit date: | 2006-10-02 | | Release date: | 2006-12-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Binding thermodynamics of substituted diaminopyrimidine renin inhibitors.
Anal.Biochem., 360, 2007
|
|
2IL1
 
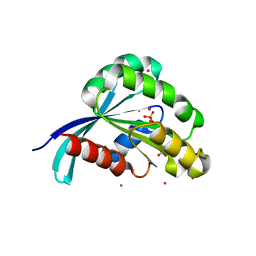 | | Crystal structure of a predicted human GTPase in complex with GDP | | Descriptor: | CALCIUM ION, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Wang, J, Shen, Y, Tempel, W, Landry, R, Arrowsmith, C.H, Edwards, A.M, Sundstrom, M, Weigelt, J, Bochkarev, A, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-10-02 | | Release date: | 2006-10-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of a predicted human GTPase in complex with GDP
To be Published
|
|
2IL2
 
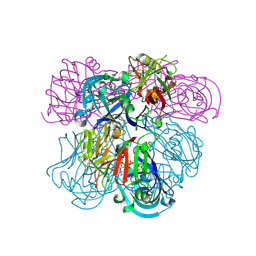 | | Crystal Structure of Human Renin Complexed with Inhibitor | | Descriptor: | CITRIC ACID, N-[2-({2-AMINO-6-ETHYL-5-[4-(3-METHOXYPROPYL)-2,2-DIMETHYL-3-OXO-3,4-DIHYDRO-2H-1,4-BENZOXAZIN-6-YL]PYRIMIDIN-4-YL}AMINO)ETHYL]NAPHTHALENE-2-SULFONAMIDE, Renin | | Authors: | Mochalkin, I. | | Deposit date: | 2006-10-02 | | Release date: | 2006-12-05 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Binding thermodynamics of substituted diaminopyrimidine renin inhibitors.
Anal.Biochem., 360, 2007
|
|
2IL3
 
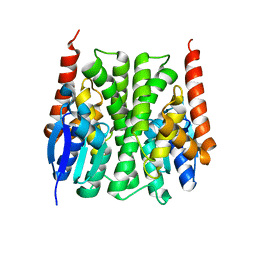 | | Structures of an Insect Epsilon-class Glutathione S-transferase from the Malaria Vector Anopheles Gambiae: Evidence for High DDT-detoxifying Activity | | Descriptor: | Epsilon-class Glutathione S-transferase | | Authors: | Wang, Y, Hemingway, J, Ranson, H, Meehan, E.J, Chen, L. | | Deposit date: | 2006-10-02 | | Release date: | 2007-10-09 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of an insect epsilon class glutathione S-transferase from the malaria vector Anopheles gambiae provides an explanation for the high DDT-detoxifying activity
J.Struct.Biol., 164, 2008
|
|
2IL4
 
 | | Crystal structure of At1g77540-Coenzyme A Complex | | Descriptor: | COENZYME A, Protein At1g77540 | | Authors: | Bitto, E, Wesenberg, G.E, Phillips Jr, G.N, Bingman, C.A, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-10-02 | | Release date: | 2006-10-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.054 Å) | | Cite: | Structure of Arabidopsis thaliana At1g77540 Protein, a Minimal Acetyltransferase from the COG2388 Family.
Biochemistry, 45, 2006
|
|
2IL5
 | |
2IL6
 
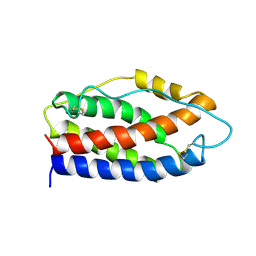 | | HUMAN INTERLEUKIN-6, NMR, 32 STRUCTURES | | Descriptor: | INTERLEUKIN-6 | | Authors: | Xu, G.Y, Yu, H.A, Hong, J, Stahl, M, Mcdonagh, T, Kay, L.E, Cumming, D.A. | | Deposit date: | 1997-01-31 | | Release date: | 1998-02-04 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of recombinant human interleukin-6.
J.Mol.Biol., 268, 1997
|
|
2IL8
 
 | |
2IL9
 
 | |
2ILA
 
 | |
2ILI
 
 | | Refine atomic structure of human carbonic anhydrase II | | Descriptor: | Carbonic anhydrase 2, ZINC ION | | Authors: | Fisher, S.Z. | | Deposit date: | 2006-10-02 | | Release date: | 2007-06-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | Atomic crystal and molecular dynamics simulation structures of human carbonic anhydrase II: insights into the proton transfer mechanism
Biochemistry, 46, 2007
|
|
2ILK
 
 | |
