7XN7
 
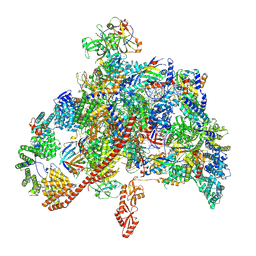 | | RNA polymerase II elongation complex containing Spt4/5, Elf1, Spt6, Spn1 and Paf1C | | 分子名称: | Chromatin elongation factor SPT5, Component of the Paf1p complex, Constituent of Paf1 complex with RNA polymerase II, ... | | 著者 | Ehara, H, Kujirai, T, Shirouzu, M, Kurumizaka, H, Sekine, S. | | 登録日 | 2022-04-28 | | 公開日 | 2022-09-07 | | 最終更新日 | 2024-07-03 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Structural basis of nucleosome disassembly and reassembly by RNAPII elongation complex with FACT.
Science, 377, 2022
|
|
3D70
 
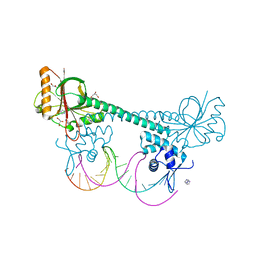 | |
4RAY
 
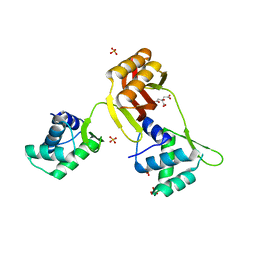 | |
4RAZ
 
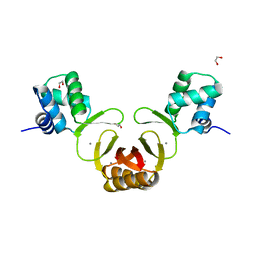 | |
4RB0
 
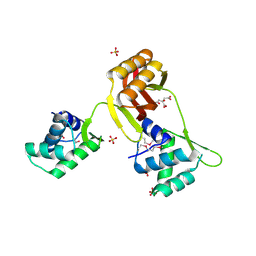 | |
1F5T
 
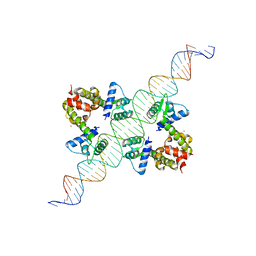 | | DIPHTHERIA TOX REPRESSOR (C102D MUTANT) COMPLEXED WITH NICKEL AND DTXR CONSENSUS BINDING SEQUENCE | | 分子名称: | 43MER DNA CONTAINING DXTR CONSENSUS BINDING SEQUENCE, DIPHTHERIA TOXIN REPRESSOR, NICKEL (II) ION | | 著者 | Chen, S, White, A, Love, J, Murphy, J.R, Ringe, D. | | 登録日 | 2000-06-15 | | 公開日 | 2000-09-25 | | 最終更新日 | 2024-02-07 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Methyl groups of thymine bases are important for nucleic acid recognition by DtxR.
Biochemistry, 39, 2000
|
|
6XAT
 
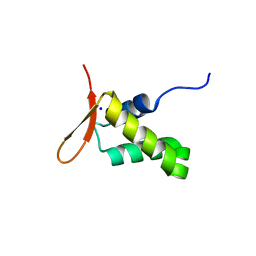 | | Crystal structure of the human FoxP4 DNA binding Domain | | 分子名称: | FOXP4 protein, SODIUM ION | | 著者 | VIllalobos, P, Castro-Fernandez, V, Medina, E, Gonzalez-Ordenes, F, Maturana, P, Herrera-Morande, A, Ramirez-Sarmiento, C.A, Babul, J. | | 登録日 | 2020-06-04 | | 公開日 | 2021-06-09 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | Unraveling the folding and dimerization properties of the human FoxP subfamily of transcription factors.
Febs Lett., 597, 2023
|
|
7ASV
 
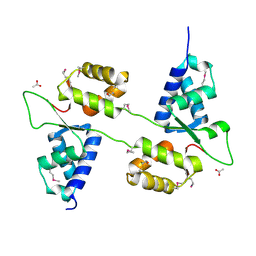 | |
2C7A
 
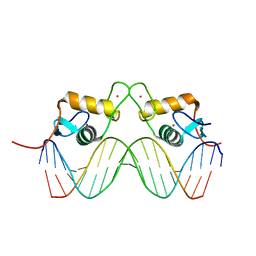 | | STRUCTURE OF THE PROGESTERONE RECEPTOR-DNA COMPLEX | | 分子名称: | 5'-D(*CP*CP*AP*GP*AP*AP*CP*AP*AP*AP *CP*TP*GP*TP*TP*CP*TP*G)-3', 5'-D(*CP*CP*AP*GP*AP*AP*CP*AP*GP*TP *TP*TP*GP*TP*TP*CP*TP*G)-3', PROGESTERONE RECEPTOR, ... | | 著者 | Roemer, S.C, Donham, D.C, Sherman, L, Pon, V.H, Edwards, D.P, Churchill, M.E.A. | | 登録日 | 2005-11-19 | | 公開日 | 2006-08-30 | | 最終更新日 | 2024-05-08 | | 実験手法 | X-RAY DIFFRACTION (2.5 Å) | | 主引用文献 | Structure of the Progesterone Receptor-Deoxyribonucleic Acid Complex: Novel Interactions Required for Binding to Half-Site Response Elements.
Mol.Endocrinol., 20, 2006
|
|
3B02
 
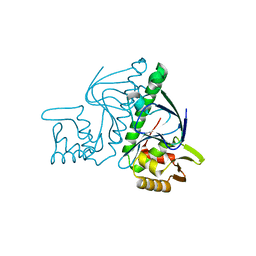 | | Crystal structure of TTHB099, a transcriptional regulator CRP family from Thermus thermophilus HB8 | | 分子名称: | Transcriptional regulator, Crp family | | 著者 | Agari, Y, Kuramitsu, S, Shinkai, A, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | 登録日 | 2011-06-03 | | 公開日 | 2011-06-15 | | 最終更新日 | 2023-11-01 | | 実験手法 | X-RAY DIFFRACTION (1.92 Å) | | 主引用文献 | X-ray crystal structure of TTHB099, a CRP/FNR superfamily transcriptional regulator from Thermus thermophilus HB8, reveals a DNA-binding protein with no required allosteric effector molecule
Proteins, 80, 2012
|
|
5ZKO
 
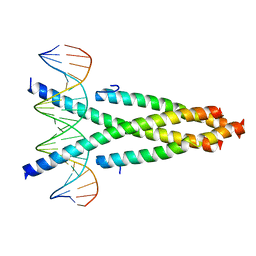 | | Crystal structure of the CRTC2-CREB-CRE complex | | 分子名称: | CREB-regulated transcription coactivator 2, Cyclic AMP-responsive element-binding protein 1, DNA (5'-D(*CP*TP*TP*GP*GP*CP*TP*GP*AP*CP*GP*TP*CP*AP*GP*CP*CP*AP*AP*G)-3') | | 著者 | Xiang, S, Zhai, L, Valecia-Swain, J. | | 登録日 | 2018-03-24 | | 公開日 | 2018-06-20 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (3.05 Å) | | 主引用文献 | Structural Insights into the CRTC2-CREB Complex Assembly on CRE.
J. Mol. Biol., 430, 2018
|
|
4S20
 
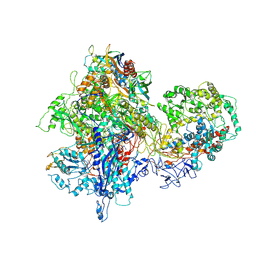 | | Structural basis for transcription reactivation by RapA | | 分子名称: | 5'-D(P*AP*CP*GP*AP*CP*TP*GP*AP*GP*CP*CP*GP*AP*TP*G)-3', 5'-R(P*AP*UP*CP*GP*GP*CP*UP*CP*A)-3', DNA-directed RNA polymerase subunit alpha, ... | | 著者 | Liu, B, Zuo, Y, Steitz, T.A. | | 登録日 | 2015-01-16 | | 公開日 | 2015-02-04 | | 最終更新日 | 2023-09-20 | | 実験手法 | X-RAY DIFFRACTION (4.7 Å) | | 主引用文献 | Structural basis for transcription reactivation by RapA.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
5ZK1
 
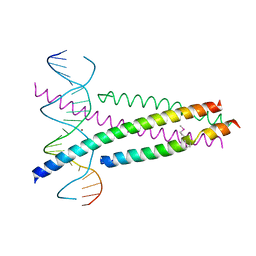 | | Crystal Structure of the CRTC2(SeMet)-CREB-CRE complex | | 分子名称: | CREB-regulated transcription coactivator 2, Cyclic AMP-responsive element-binding protein 1, DNA (5'-D(*CP*TP*TP*GP*GP*CP*TP*GP*AP*CP*GP*TP*CP*AP*GP*CP*CP*AP*AP*G)-3'), ... | | 著者 | Xiang, S, Zhai, L, Valencia-Swain, J. | | 登録日 | 2018-03-22 | | 公開日 | 2018-06-20 | | 最終更新日 | 2023-11-22 | | 実験手法 | X-RAY DIFFRACTION (3.05 Å) | | 主引用文献 | Structural Insights into the CRTC2-CREB Complex Assembly on CRE.
J. Mol. Biol., 430, 2018
|
|
5O6D
 
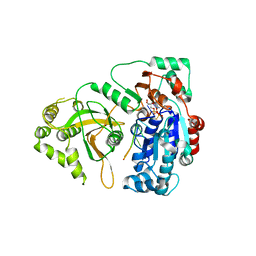 | | Structure of ScPif1 in complex with polydT and ATPgS | | 分子名称: | ATP-dependent DNA helicase PIF1, DNA (5'-D(P*TP*TP*TP*TP*TP*T)-3'), MAGNESIUM ION, ... | | 著者 | Lu, K.Y, Chen, W.F, Rety, S, Liu, N.N, Xu, X.G. | | 登録日 | 2017-06-06 | | 公開日 | 2017-12-13 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (3.283 Å) | | 主引用文献 | Insights into the structural and mechanistic basis of multifunctional S. cerevisiae Pif1p helicase.
Nucleic Acids Res., 46, 2018
|
|
7Y8R
 
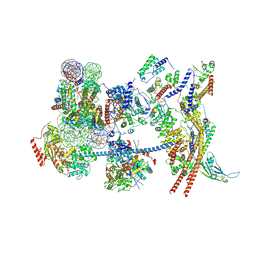 | | The nucleosome-bound human PBAF complex | | 分子名称: | ACTB protein (Fragment), ADENOSINE-5'-DIPHOSPHATE, AT-rich interactive domain-containing protein 2, ... | | 著者 | Wang, L, Yu, J, Yu, Z, Wang, Q, He, S, Xu, Y. | | 登録日 | 2022-06-24 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-07-03 | | 実験手法 | ELECTRON MICROSCOPY (4.4 Å) | | 主引用文献 | Structure of nucleosome-bound human PBAF complex.
Nat Commun, 13, 2022
|
|
1BHI
 
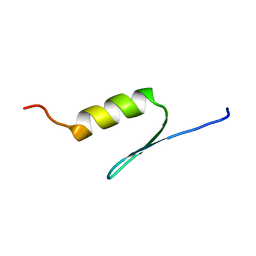 | | STRUCTURE OF TRANSACTIVATION DOMAIN OF CRE-BP1/ATF-2, NMR, 20 STRUCTURES | | 分子名称: | CRE-BP1 | | 著者 | Nagadoi, A, Nakazawa, K, Uda, H, Maekawa, T, Ishii, S, Nishimura, Y. | | 登録日 | 1998-06-09 | | 公開日 | 1999-06-15 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Solution structure of the transactivation domain of ATF-2 comprising a zinc finger-like subdomain and a flexible subdomain.
J.Mol.Biol., 287, 1999
|
|
4WOI
 
 | | 4,5-linked aminoglycoside antibiotics regulate the bacterial ribosome by targeting dynamic conformational processes within intersubunit bridge B2 | | 分子名称: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | 著者 | Pulk, A, Cate, J.H.D, Blanchard, S, Wasserman, M, Altman, R, Zhou, Z, Zinder, J, Green, K, Garneau-Tsodikova, S. | | 登録日 | 2014-10-15 | | 公開日 | 2015-08-05 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Chemically related 4,5-linked aminoglycoside antibiotics drive subunit rotation in opposite directions.
Nat Commun, 6, 2015
|
|
2H27
 
 | |
9FOF
 | |
9FOR
 
 | |
7SHL
 
 | | Structure of Xenopus laevis CRL2Lrr1 (State 2) | | 分子名称: | CULLIN_2 domain-containing protein, Elongin-C, Lrr1, ... | | 著者 | Zhou, H, Brown, A. | | 登録日 | 2021-10-09 | | 公開日 | 2021-12-08 | | 最終更新日 | 2024-06-05 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Structure of CRL2Lrr1, the E3 ubiquitin ligase that promotes DNA replication termination in vertebrates.
Nucleic Acids Res., 49, 2021
|
|
7SHK
 
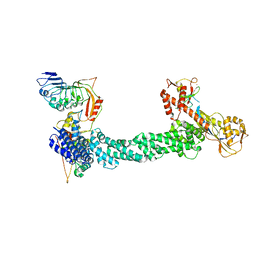 | | Structure of Xenopus laevis CRL2Lrr1 (State 1) | | 分子名称: | CULLIN_2 domain-containing protein, Elongin-C, Lrr1, ... | | 著者 | Zhou, H, Brown, A. | | 登録日 | 2021-10-09 | | 公開日 | 2021-12-08 | | 最終更新日 | 2024-06-05 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Structure of CRL2Lrr1, the E3 ubiquitin ligase that promotes DNA replication termination in vertebrates.
Nucleic Acids Res., 49, 2021
|
|
6SKL
 
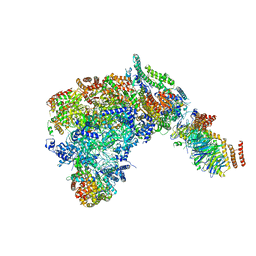 | | Cryo-EM structure of the CMG Fork Protection Complex at a replication fork - Conformation 1 | | 分子名称: | Cell division control protein 45, Chromosome segregation in meiosis protein 3, DNA fork, ... | | 著者 | Yeeles, J, Baretic, D, Jenkyn-Bedford, M. | | 登録日 | 2019-08-16 | | 公開日 | 2020-05-06 | | 最終更新日 | 2024-05-22 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Cryo-EM Structure of the Fork Protection Complex Bound to CMG at a Replication Fork.
Mol.Cell, 78, 2020
|
|
5U8S
 
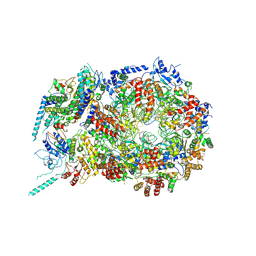 | | Structure of eukaryotic CMG helicase at a replication fork | | 分子名称: | ADENOSINE-5'-TRIPHOSPHATE, Cell division control protein 45, DNA (26-MER), ... | | 著者 | Li, H, Li, B, Georgescu, R, Yuan, Z, Santos, R, Sun, J, Zhang, D, Yurieva, O, O'Donnell, M.E. | | 登録日 | 2016-12-14 | | 公開日 | 2017-01-25 | | 最終更新日 | 2020-01-01 | | 実験手法 | ELECTRON MICROSCOPY (6.101 Å) | | 主引用文献 | Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
8DQ1
 
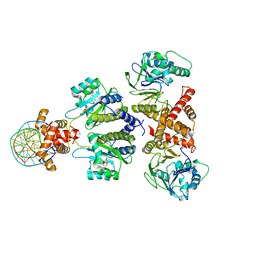 | | Quorum-sensing receptor RhlR bound to PqsE | | 分子名称: | 2-aminobenzoylacetyl-CoA thioesterase, 4-(3-bromophenoxy)-N-[(3S)-2-oxothiolan-3-yl]butanamide, DNA (5'-D(*AP*CP*CP*TP*GP*CP*CP*AP*GP*AP*CP*TP*GP*CP*AP*CP*AP*G)-3'), ... | | 著者 | Paczkowski, J.E, Fromme, J.C, Feathers, J.R. | | 登録日 | 2022-07-18 | | 公開日 | 2022-12-07 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (4.1 Å) | | 主引用文献 | Structure of the RhlR-PqsE complex from Pseudomonas aeruginosa reveals mechanistic insights into quorum-sensing gene regulation.
Structure, 30, 2022
|
|
