8U1D
 
 | | Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, DH1285 Heavy Chain, DH1285 Light Chain, ... | | 著者 | Thakur, B, Stalls, V.D, Acharya, P. | | 登録日 | 2023-08-31 | | 公開日 | 2024-01-03 | | 最終更新日 | 2025-06-04 | | 実験手法 | ELECTRON MICROSCOPY (4.25 Å) | | 主引用文献 | Vaccine induction of CD4-mimicking HIV-1 broadly neutralizing antibody precursors in macaques.
Cell, 187, 2024
|
|
6QV6
 
 | |
6QVB
 
 | |
8YVR
 
 | |
7RI1
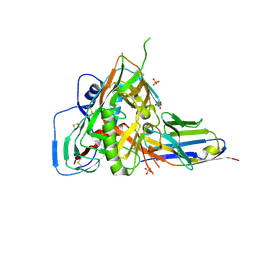 
 | |
6QVU
 
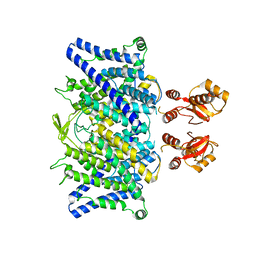 | |
8D3Z
 
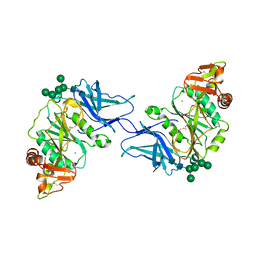 | | Crystal structure of GalS1 in complex with Manganese from Populus trichocarpas | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Galactan synthase, ... | | 著者 | Pereira, J.H, Prabhakar, P.K, Urbanowicz, B.R, Adams, P.D. | | 登録日 | 2022-06-01 | | 公開日 | 2023-03-15 | | 最終更新日 | 2024-10-30 | | 実験手法 | X-RAY DIFFRACTION (2.56 Å) | | 主引用文献 | Structural and biochemical insight into a modular beta-1,4-galactan synthase in plants.
Nat.Plants, 9, 2023
|
|
8HRV
 
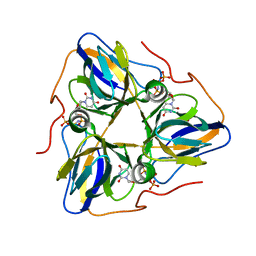 | | dutpase of helicobacter pylori 26695 | | 分子名称: | 1,2-ETHANEDIOL, 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, Deoxyuridine 5'-triphosphate nucleotidohydrolase, ... | | 著者 | Kumari, K, Gourinath, S. | | 登録日 | 2022-12-16 | | 公開日 | 2024-01-17 | | 最終更新日 | 2025-03-26 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Structural analysis of dUTPase from Helicobacter pylori reveals unusual activity for dATP.
Int.J.Biol.Macromol., 282, 2024
|
|
8SAL
 
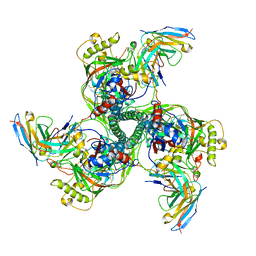 | | CryoEM structure of VRC01-CH848.0358.80 | | 分子名称: | CH0848.3.D0358.80.06CHIM.DS.6R.SOSIP gp120, CH0848.3.D0358.80.06CHIM.DS.6R.SOSIP gp41, VCR01 variable heavy chain, ... | | 著者 | Henderson, R, Zhou, Y, Stalls, V, Bartesaghi, B, Acharya, P. | | 登録日 | 2023-04-01 | | 公開日 | 2023-04-19 | | 最終更新日 | 2025-05-21 | | 実験手法 | ELECTRON MICROSCOPY (4.9 Å) | | 主引用文献 | Structural basis for breadth development in the HIV-1 V3-glycan targeting DH270 antibody clonal lineage.
Nat Commun, 14, 2023
|
|
8SAT
 
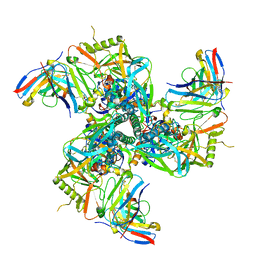 | | CryoEM structure of VRC01-CH848.10.17 | | 分子名称: | CH848.10.17 gp120, CH848.10.17 gp41, VRC01-variable heavy chain, ... | | 著者 | Henderson, R, Zhou, Y, Stalls, V, Bartesaghi, B, Acharya, P. | | 登録日 | 2023-04-02 | | 公開日 | 2023-04-19 | | 最終更新日 | 2025-05-28 | | 実験手法 | ELECTRON MICROSCOPY (4.5 Å) | | 主引用文献 | Structural basis for breadth development in the HIV-1 V3-glycan targeting DH270 antibody clonal lineage.
Nat Commun, 14, 2023
|
|
8UA2
 
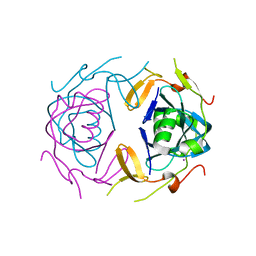 | | Crystal Structure of infected cell protein 0 (ICP0) from herpes simplex virus 1 (proteolyzed fragment) | | 分子名称: | IODIDE ION, RL2 | | 著者 | Lovell, S, Kashipathy, M, Battaile, K.P, Cooper, A, Davido, D. | | 登録日 | 2023-09-20 | | 公開日 | 2024-02-28 | | 最終更新日 | 2024-06-19 | | 実験手法 | X-RAY DIFFRACTION (2.65 Å) | | 主引用文献 | HSV-1 ICP0 dimer domain adopts a novel beta-barrel fold.
Proteins, 92, 2024
|
|
8DJ1
 
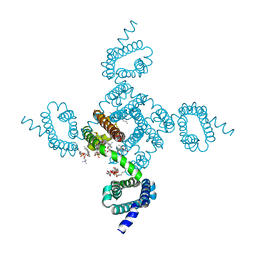 | | Crystal structure of NavAb V126T as a basis for the human Nav1.7 Inherited Erythromelalgia S241T mutation | | 分子名称: | 1,2-DIMYRISTOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, Ion transport protein | | 著者 | Wisedchaisri, G, Gamal El-Din, T.M, Zheng, N, Catterall, W.A. | | 登録日 | 2022-06-29 | | 公開日 | 2023-04-12 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (3.1 Å) | | 主引用文献 | Structural basis for severe pain caused by mutations in the S4-S5 linkers of voltage-gated sodium channel Na V 1.7.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8TY6
 
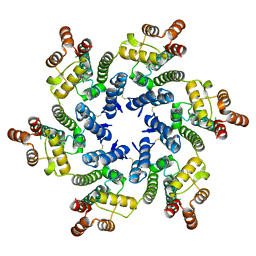 | |
6GPK
 
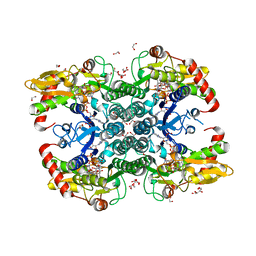 | | Crystal structure of human GDP-D-mannose 4,6-dehydratase (E157Q) in complex with GDP-Man | | 分子名称: | 1,2-ETHANEDIOL, GDP-mannose 4,6 dehydratase, GLYCEROL, ... | | 著者 | Pfeiffer, M, Krojer, T, Johansson, C, von Delft, F, Bountra, C, Arrowsmith, C.H, Edwards, A, Nidetzky, B, Oppermann, U, Structural Genomics Consortium (SGC) | | 登録日 | 2018-06-06 | | 公開日 | 2018-07-18 | | 最終更新日 | 2024-05-15 | | 実験手法 | X-RAY DIFFRACTION (1.47 Å) | | 主引用文献 | A Parsimonious Mechanism of Sugar Dehydration by Human GDP-Mannose-4,6-dehydratase.
Acs Catalysis, 9, 2019
|
|
6QVC
 
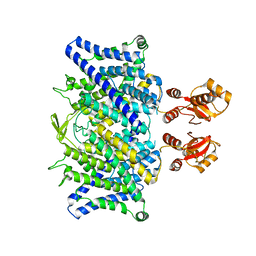 | |
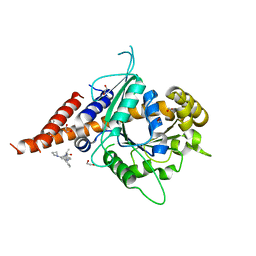
8SKN
 
 | |
6YB4
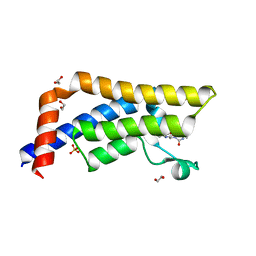 
 | |
8ZD8
 
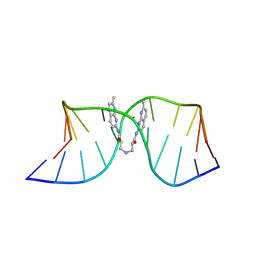 | | NMR structure of the (CGG-dsDNA:ND=) 1:1 complex | | 分子名称: | DNA (5'-D(*CP*AP*TP*TP*CP*GP*GP*TP*TP*AP*G)-3'), DNA (5'-D(*CP*TP*AP*AP*CP*GP*GP*AP*AP*TP*G)-3'), ~{N}-(7-methyl-1,8-naphthyridin-2-yl)-3-[[3-[(7-methyl-1,8-naphthyridin-2-yl)amino]-3-oxidanylidene-propyl]amino]propanamide | | 著者 | Sakurabayashi, S, Furuita, K, Yamada, T, Nomura, M, Nakatani, K, Kojima, C. | | 登録日 | 2024-05-01 | | 公開日 | 2025-04-30 | | 実験手法 | SOLUTION NMR | | 主引用文献 | NMR structure of the (CGG-dsDNA:ND=) 1:1 complex
To Be Published
|
|
7R97
 
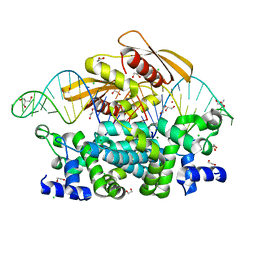 | | Crystal structure of postcleavge complex of Escherichia coli RNase III | | 分子名称: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, ... | | 著者 | Dharavath, S, Shaw, G.X, Ji, X. | | 登録日 | 2021-06-28 | | 公開日 | 2022-07-20 | | 最終更新日 | 2023-10-18 | | 実験手法 | X-RAY DIFFRACTION (1.804 Å) | | 主引用文献 | Structural basis for Dicer-like function of an engineered RNase III variant and insights into the reaction trajectory of two-Mg 2+ -ion catalysis.
Rna Biol., 19, 2022
|
|
6HY3
 
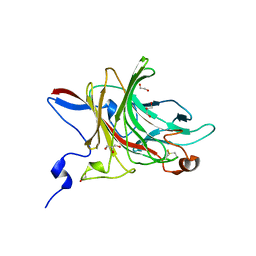 | | Three-dimensional structure of AgaC from Zobellia galactanivorans | | 分子名称: | 1,2-ETHANEDIOL, Beta-agarase C, GLYCEROL, ... | | 著者 | Naretto, A, Fanuel, M, Ropartz, D, Rogniaux, H, Larocque, R, Czjzek, M, Tellier, C, Michel, G. | | 登録日 | 2018-10-19 | | 公開日 | 2019-03-13 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (1.3 Å) | | 主引用文献 | The agar-specific hydrolaseZgAgaC from the marine bacteriumZobellia galactanivoransdefines a new GH16 protein subfamily.
J.Biol.Chem., 294, 2019
|
|
5XXD
 
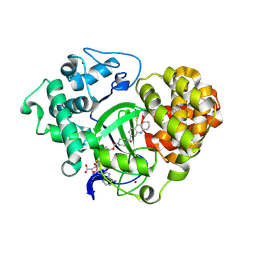 | |
7RJ2
 
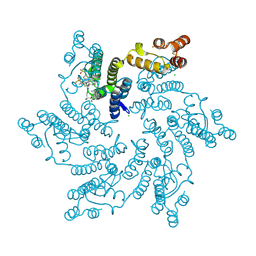 | |
7RHM
 
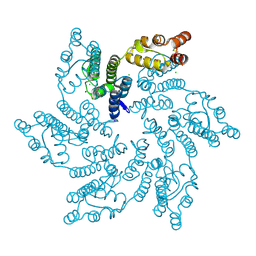 | |
6ZK6
 
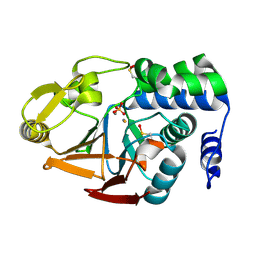 | | Protein Phosphatase 1 (PP1) T320E mutant | | 分子名称: | FE (III) ION, MANGANESE (II) ION, PHOSPHATE ION, ... | | 著者 | Salvi, F, Barabas, O, Koehn, M. | | 登録日 | 2020-06-29 | | 公開日 | 2020-11-18 | | 最終更新日 | 2024-11-06 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Towards Dissecting the Mechanism of Protein Phosphatase-1 Inhibition by Its C-Terminal Phosphorylation.
Chembiochem, 22, 2021
|
|
6Z4V
 
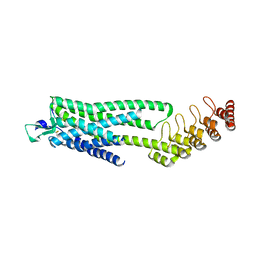 | | Crystal structure of the neurotensin receptor 1 (NTSR1-H4bmx) in complex with NTS8-13 | | 分子名称: | ARG-PRO-TYR-ILE-LEU, Neurotensin receptor type 1,Neurotensin receptor type 1,DARPin | | 著者 | Deluigi, M, Klipp, A, Hilge, M, Merklinger, L, Klenk, C, Plueckthun, A. | | 登録日 | 2020-05-25 | | 公開日 | 2021-02-10 | | 最終更新日 | 2024-11-13 | | 実験手法 | X-RAY DIFFRACTION (2.595 Å) | | 主引用文献 | Complexes of the neurotensin receptor 1 with small-molecule ligands reveal structural determinants of full, partial, and inverse agonism.
Sci Adv, 7, 2021
|
|
