1SHR
 
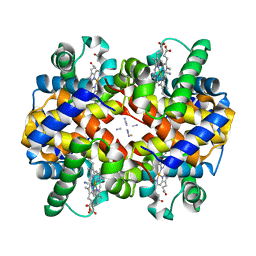 | | Crystal structure of ferrocyanide bound human hemoglobin A2 at 1.88A resolution | | Descriptor: | CYANIDE ION, FE (III) ION, Hemoglobin alpha chain, ... | | Authors: | Sen, U, Dasgupta, J, Choudhury, D, Datta, P, Chakrabarti, A, Chakrabarty, S.B, Chakrabarty, A, Dattagupta, J.K. | | Deposit date: | 2004-02-26 | | Release date: | 2004-10-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Crystal structures of HbA2 and HbE and modeling of hemoglobin delta4: interpretation of the thermal stability and the antisickling effect of HbA2 and identification of the ferrocyanide binding site in Hb
Biochemistry, 43, 2004
|
|
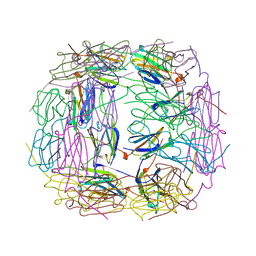
1SHS
 
 | |
1SHT
 
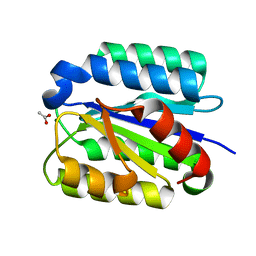 | | Crystal Structure of the von Willebrand factor A domain of human capillary morphogenesis protein 2: an anthrax toxin receptor | | Descriptor: | ACETATE ION, Anthrax toxin receptor 2, MAGNESIUM ION | | Authors: | Lacy, D.B, Wigelsworth, D.J, Scobie, H.M, Young, J.A.T, Collier, R.J. | | Deposit date: | 2004-02-26 | | Release date: | 2004-04-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Crystal Structure of the von Willebrand factor A domain of human capillary morphogenesis protein 2: an anthrax toxin receptor
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1SHU
 
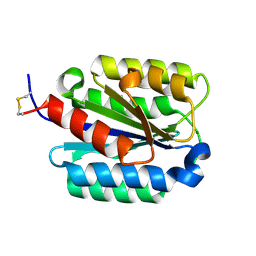 | | Crystal Structure of the von Willebrand factor A domain of human capillary morphogenesis protein 2: an anthrax toxin receptor | | Descriptor: | Anthrax toxin receptor 2, MAGNESIUM ION | | Authors: | Lacy, D.B, Wigelsworth, D.J, Scobie, H.M, Young, J.A.T, Collier, R.J. | | Deposit date: | 2004-02-26 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystal Structure of the von Willebrand factor A domain of human capillary morphogenesis protein 2: an anthrax toxin receptor
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1SHV
 
 | | STRUCTURE OF SHV-1 BETA-LACTAMASE | | Descriptor: | CYCLOHEXYL-HEXYL-BETA-D-MALTOSIDE, PROTEIN (BETA-LACTAMASE SHV-1) | | Authors: | Kuzin, A.P, Nukaga, M, Nukaga, Y, Hujer, A, Bonomo, R.A, Knox, J.R. | | Deposit date: | 1999-02-23 | | Release date: | 1999-05-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structure of the SHV-1 beta-lactamase.
Biochemistry, 38, 1999
|
|
1SHW
 
 | | EphB2 / EphrinA5 Complex Structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Ephrin type-B receptor 2, Ephrin-A5, ... | | Authors: | Himanen, J.P, Chumley, M.J, Lackmann, M, Li, C, Barton, W.A, Jeffrey, P.D, Vearing, C, Geleick, D, Feldheim, D.A, Boyd, A.W. | | Deposit date: | 2004-02-26 | | Release date: | 2004-05-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Repelling class discrimination: ephrin-A5 binds to and activates EphB2 receptor signaling
Nat.Neurosci., 7, 2004
|
|
1SHX
 
 | | Ephrin A5 ligand structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Ephrin-A5 | | Authors: | Himanen, J.P, Barton, W.A, Nikolov, D.B, Jeffrey, P.D. | | Deposit date: | 2004-02-26 | | Release date: | 2005-04-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Three distinct molecular surfaces in ephrin-A5 are essential for a functional interaction with EphA3.
J.Biol.Chem., 280, 2005
|
|
1SHY
 
 | |
1SHZ
 
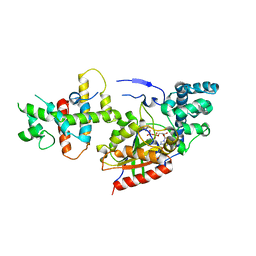 | | Crystal Structure of the p115RhoGEF rgRGS Domain in A Complex with Galpha(13):Galpha(i1) Chimera | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Guanine Nucleotide-Binding Protein Galpha(13):Galpha(i1) Chimera, MAGNESIUM ION, ... | | Authors: | Chen, Z, Singer, W.D, Sternweis, P.C, Sprang, S.R. | | Deposit date: | 2004-02-26 | | Release date: | 2005-01-18 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structure of the p115RhoGEF rgRGS domain-Galpha13/i1 chimera complex suggests convergent evolution of a GTPase activator.
Nat.Struct.Mol.Biol., 12, 2005
|
|
1SI0
 
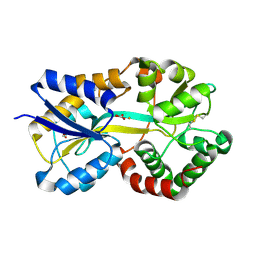 | | Crystal Structure of Mannheimia haemolytica Ferric iron-Binding Protein A in a closed conformation | | Descriptor: | 1,2-ETHANEDIOL, CARBONATE ION, FE (III) ION, ... | | Authors: | Shouldice, S.R, Skene, R.J, Dougan, D.R, Snell, G, McRee, D.E, Schryvers, A.B, Tari, L.W. | | Deposit date: | 2004-02-26 | | Release date: | 2004-06-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structural basis for iron binding and release by a novel class of periplasmic iron-binding proteins found in gram-negative pathogens.
J.Bacteriol., 186, 2004
|
|
1SI1
 | | Crystal Structure of Mannheimia haemolytica Ferric iron-Binding Protein A in an open conformation | | Descriptor: | FE (III) ION, iron binding protein FbpA | | Authors: | Shouldice, S.R, Skene, R.J, Dougan, D.R, Snell, G, McRee, D.E, Schryvers, A.B, Tari, L.W. | | Deposit date: | 2004-02-26 | | Release date: | 2004-06-08 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Structural basis for iron binding and release by a novel class of periplasmic iron-binding proteins found in gram-negative pathogens.
J.Bacteriol., 186, 2004
|
|
1SI2
 
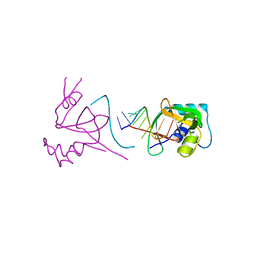 | |
1SI3
 
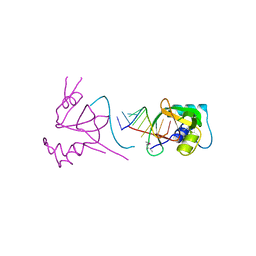 | |
1SI4
 
 | | Crystal structure of Human hemoglobin A2 (in R2 state) at 2.2 A resolution | | Descriptor: | CYANIDE ION, Hemoglobin alpha chain, Hemoglobin delta chain, ... | | Authors: | Sen, U, Dasgupta, J, Choudhury, D, Datta, P, Chakrabarti, A, Chakrabarty, S.B, Chakrabarty, A, Dattagupta, J.K. | | Deposit date: | 2004-02-27 | | Release date: | 2004-10-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of HbA2 and HbE and modeling of hemoglobin delta4: interpretation of the thermal stability and the antisickling effect of HbA2 and identification of the ferrocyanide binding site in Hb.
Biochemistry, 43, 2004
|
|
1SI5
 
 | | Protease-like domain from 2-chain hepatocyte growth factor | | Descriptor: | hepatocyte growth factor | | Authors: | Kirchhofer, D, Yao, X, Peek, M, Eigenbrot, C, Lipari, M.T, Billeci, K.L, Maun, H.R, Moran, P, Santell, L, Lazarus, R.A. | | Deposit date: | 2004-02-27 | | Release date: | 2004-12-28 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Structural and functional basis of the serine protease-like hepatocyte growth factor beta-chain in Met binding and signaling
J.Biol.Chem., 279, 2004
|
|
1SI6
 
 | |
1SI7
 
 | | Structure of E. coli tRNA psi 13 pseudouridine synthase TruD | | Descriptor: | tRNA pseudouridine synthase D | | Authors: | Kaya, Y, Del Campo, M, Ofengand, J, Malhotra, A. | | Deposit date: | 2004-02-27 | | Release date: | 2004-03-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of TruD, a novel pseudouridine synthase with a new protein fold
J.Biol.Chem., 279, 2004
|
|
1SI8
 
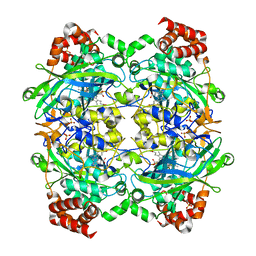 | | Crystal structure of E. faecalis catalase | | Descriptor: | CHLORIDE ION, Catalase, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Hakansson, K.O, Brugna, M, Tasse, L. | | Deposit date: | 2004-02-28 | | Release date: | 2004-05-04 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The three-dimensional structure of catalase from Enterococcus faecalis.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1SI9
 
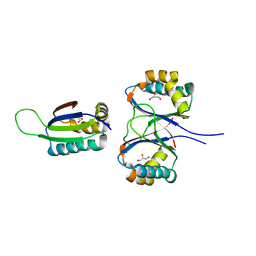 | | Boiling stable protein isolated from Populus tremula | | Descriptor: | GLYCEROL, stable protein 1 | | Authors: | Almog, O, Gonzales, A, Shoseyov, O, Dgany, O, Sofer, O, Wolf, S.G. | | Deposit date: | 2004-02-29 | | Release date: | 2004-09-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | The Structural Basis of the Thermostability of SP1, a Novel Plant (Populus tremula) Boiling Stable Protein.
J.Biol.Chem., 279, 2004
|
|
1SIB
 
 | |
1SID
 
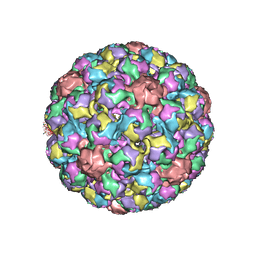 | | MURINE POLYOMAVIRUS COMPLEXED WITH 3'SIALYL LACTOSE | | Descriptor: | N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, POLYOMAVIRUS COAT PROTEIN VP1 | | Authors: | Stehle, T, Harrison, S.C. | | Deposit date: | 1995-12-12 | | Release date: | 1996-06-20 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.65 Å) | | Cite: | Crystal structures of murine polyomavirus in complex with straight-chain and branched-chain sialyloligosaccharide receptor fragments.
Structure, 4, 1996
|
|
1SIE
 
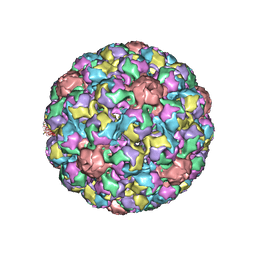 | | MURINE POLYOMAVIRUS COMPLEXED WITH A DISIALYLATED OLIGOSACCHARIDE | | Descriptor: | N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-3)-[N-acetyl-alpha-neuraminic acid-(2-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, POLYOMAVIRUS COAT PROTEIN VP1 | | Authors: | Stehle, T, Harrison, S.C. | | Deposit date: | 1995-12-12 | | Release date: | 1996-06-20 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.65 Å) | | Cite: | Crystal structures of murine polyomavirus in complex with straight-chain and branched-chain sialyloligosaccharide receptor fragments.
Structure, 4, 1996
|
|
1SIF
 
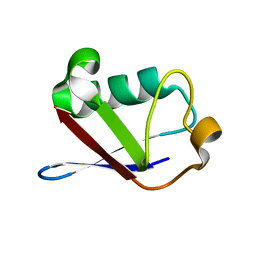 | | Crystal structure of a multiple hydrophobic core mutant of ubiquitin | | Descriptor: | ubiquitin | | Authors: | Benitez-Cardoza, C.G, Stott, K, Hirshberg, M, Went, H.M, Woolfson, D.N, Jackson, S.E. | | Deposit date: | 2004-02-29 | | Release date: | 2004-07-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Exploring sequence/folding space: folding studies on multiple hydrophobic core mutants of ubiquitin
Biochemistry, 43, 2004
|
|
1SIG
 
 | |
1SIH
 
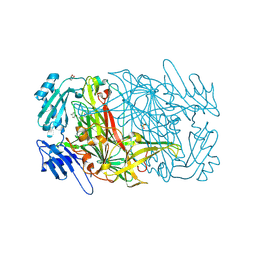 | | AGAO in covalent complex with the inhibitor MOBA ("4-(4-methylphenoxy)-2-butyn-1-amine") | | Descriptor: | COPPER (II) ION, GLYCEROL, Phenylethylamine oxidase, ... | | Authors: | Guss, J.M, Langley, D.B, Duff, A.P. | | Deposit date: | 2004-02-29 | | Release date: | 2004-09-07 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Differential Inhibition of Six Copper Amine Oxidases by a Family of 4-(Aryloxy)-2-butynamines: Evidence for a New Mode of Inactivation.
Biochemistry, 43, 2004
|
|
