3I5X
 
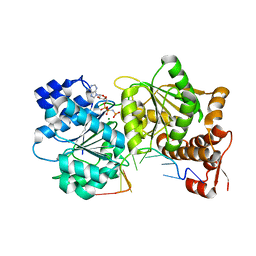 | | Structure of Mss116p bound to ssRNA and AMP-PNP | | Descriptor: | 5'-R(*UP*UP*UP*UP*UP*UP*UP*UP*UP*U)-3', ATP-dependent RNA helicase MSS116, MAGNESIUM ION, ... | | Authors: | Del Campo, M, Lambowitz, A.M. | | Deposit date: | 2009-07-06 | | Release date: | 2009-09-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA
Mol.Cell, 35, 2009
|
|
3SQX
 
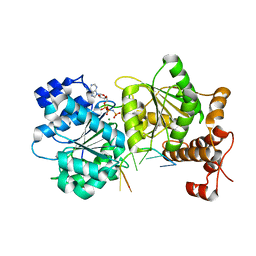 | |
3SQW
 
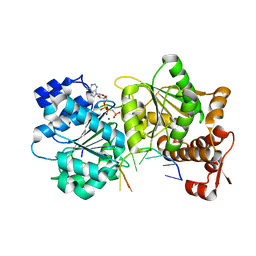 | |
3I62
 
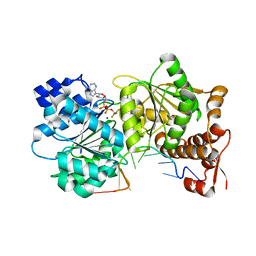 | |
3I61
 
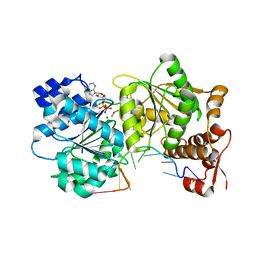 | |
3I5Y
 
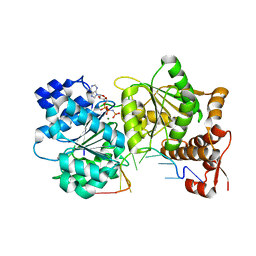 | |
1XPI
 
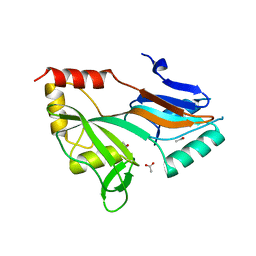 | |
1QYU
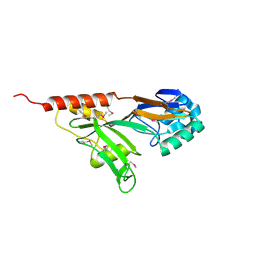 
 | |
1SI7
 
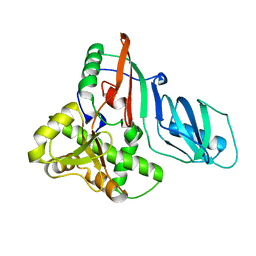 | | Structure of E. coli tRNA psi 13 pseudouridine synthase TruD | | Descriptor: | tRNA pseudouridine synthase D | | Authors: | Kaya, Y, Del Campo, M, Ofengand, J, Malhotra, A. | | Deposit date: | 2004-02-27 | | Release date: | 2004-03-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of TruD, a novel pseudouridine synthase with a new protein fold
J.Biol.Chem., 279, 2004
|
|
4DB2
 
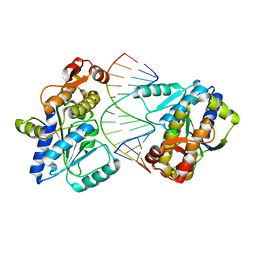 | | Mss116p DEAD-box helicase domain 2 bound to an RNA duplex | | Descriptor: | 5'-R(*GP*GP*GP*CP*GP*GP*GP*CP*CP*CP*GP*CP*CP*C)-3', ATP-dependent RNA helicase MSS116, mitochondrial | | Authors: | Mallam, A.L, Del Campo, M, Gilman, B.D, Sidote, D.J, Lambowitz, A. | | Deposit date: | 2012-01-13 | | Release date: | 2012-08-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.157 Å) | | Cite: | Structural basis for RNA-duplex recognition and unwinding by the DEAD-box helicase Mss116p.
Nature, 490, 2012
|
|
4DB4
 
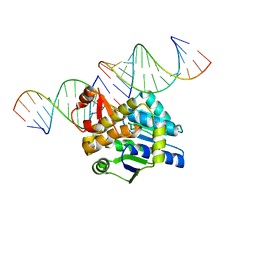 | | Mss116p DEAD-box helicase domain 2 bound to a chimaeric RNA-DNA duplex | | Descriptor: | 5'-R(*GP*GP*GP*CP*GP*GP*G)-D(P*CP*CP*CP*GP*CP*CP*C)-3', ATP-dependent RNA helicase MSS116, mitochondrial | | Authors: | Mallam, A.L, Del Campo, M, Gilman, B.D, Sidote, D.J, Lambowitz, A. | | Deposit date: | 2012-01-13 | | Release date: | 2012-08-29 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.599 Å) | | Cite: | Structural basis for RNA-duplex recognition and unwinding by the DEAD-box helicase Mss116p.
Nature, 490, 2012
|
|
7KWM
 
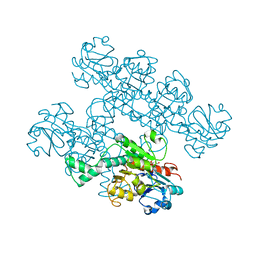 | | CtBP1 (28-375) L182F/V185T - AMP | | Descriptor: | ADENOSINE MONOPHOSPHATE, C-terminal-binding protein 1, CALCIUM ION | | Authors: | Royer, W.E, Del Campo, M. | | Deposit date: | 2020-12-01 | | Release date: | 2021-02-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | NAD(H) phosphates mediate tetramer assembly of human C-terminal binding protein (CtBP).
J.Biol.Chem., 296, 2021
|
|
