1JRN
 
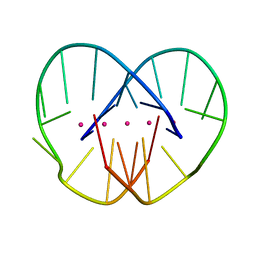 | |
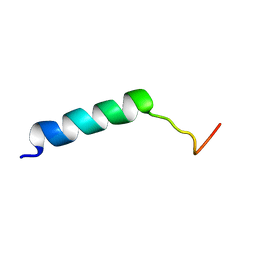
7M1E
 
 | |
4JAY
 
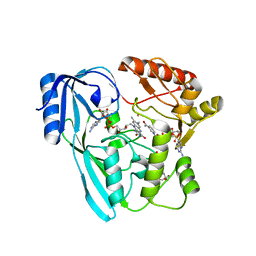 | | Crystal structure of P. aeruginosa MurB in complex with NADP+ | | Descriptor: | 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Chen, M.W, Lohkamp, B, Schnell, R, Lescar, J, Schneider, G. | | Deposit date: | 2013-02-19 | | Release date: | 2013-07-17 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Substrate Channel Flexibility in Pseudomonas aeruginosa MurB Accommodates Two Distinct Substrates.
Plos One, 8, 2013
|
|
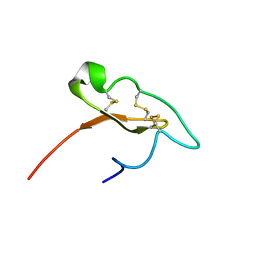
7M1D
 
 | |
1LUP
 
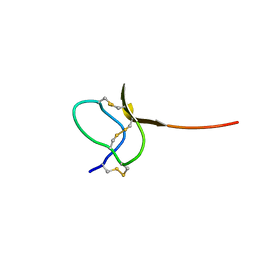 | | Solution structure of a toxin (GsMTx2) from the tarantula, Grammostola spatulata, which inhibits mechanosensitive ion channels | | Descriptor: | GsMTx2 | | Authors: | Oswald, R.E, Suchyna, T.M, McFeeters, R, Gottlieb, P, Sachs, F. | | Deposit date: | 2002-05-23 | | Release date: | 2002-08-07 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of peptide toxins that block mechanosensitive ion channels
J.Biol.Chem., 277, 2002
|
|
2BKO
 
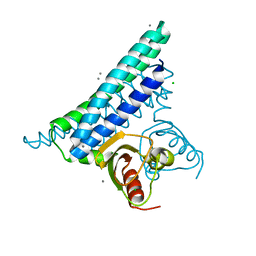 | | STRUCTURE ANALYSIS OF UNKNOWN FUNCTION PROTEIN | | Descriptor: | CALCIUM ION, CHLORIDE ION, HYPOTHETICAL PROTEIN PH0236 | | Authors: | Inagaki, E, Tahirov, T.H. | | Deposit date: | 2005-02-17 | | Release date: | 2006-06-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Analysis of Putative Potassium Channel Related Protein from Pyrococcus Horikoshii
To be Published
|
|
2BKN
 
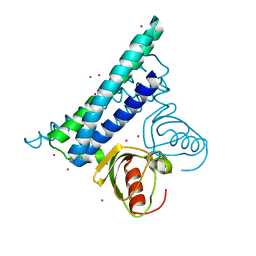 | |
2MVT
 
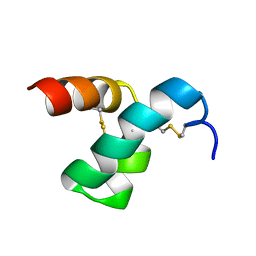 | | Solution structure of scoloptoxin SSD609 from Scolopendra mutilans | | Descriptor: | Scoloptoxin SSD609 | | Authors: | Wu, F, Sun, P, Wang, C, He, Y, Zhang, L, Tian, C. | | Deposit date: | 2014-10-14 | | Release date: | 2015-09-23 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | A distinct three-helix centipede toxin SSD609 inhibits Iks channels by interacting with the KCNE1 auxiliary subunit.
Sci Rep, 5, 2015
|
|
3OXQ
 
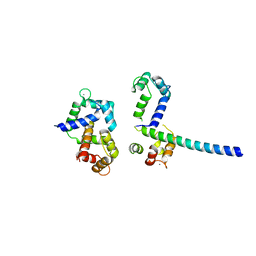 | | Crystal Structure of Ca2+/CaM-CaV1.2 pre-IQ/IQ domain complex | | Descriptor: | CALCIUM ION, Calmodulin, Voltage-dependent L-type calcium channel subunit alpha-1C | | Authors: | Kim, E.Y, Rumpf, C.H, Van Petegem, F, Arant, R, Findeisen, F, Cooley, E.S, Isacoff, E.Y, Minor, D.L. | | Deposit date: | 2010-09-21 | | Release date: | 2010-11-03 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Multiple C-terminal tail Ca(2+)/CaMs regulate Ca(V)1.2 function but do not mediate channel dimerization.
Embo J., 29, 2010
|
|
3RBX
 
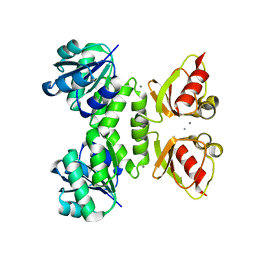 | | MthK RCK domain D184N mutant, Ca2+-bound | | Descriptor: | CALCIUM ION, Calcium-gated potassium channel mthK | | Authors: | Samakai, E, Rothberg, B.S. | | Deposit date: | 2011-03-30 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structures of multiple Ca2+-binding sites in a K+ channel RCK domain
Proc.Natl.Acad.Sci.USA, 2011
|
|
3RDE
 
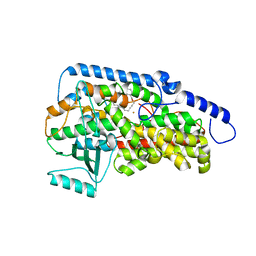 | | Crystal structure of the catalytic domain of porcine leukocyte 12-lipoxygenase | | Descriptor: | 3-{4-[(tridec-2-yn-1-yloxy)methyl]phenyl}propanoic acid, Arachidonate 12-lipoxygenase, 12S-type, ... | | Authors: | Funk, M.O, Xu, S, Marnett, L.J, Mueser, T.C. | | Deposit date: | 2011-04-01 | | Release date: | 2012-04-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.892 Å) | | Cite: | Crystal structure of 12-lipoxygenase catalytic-domain-inhibitor complex identifies a substrate-binding channel for catalysis.
Structure, 20, 2012
|
|
7L8V
 
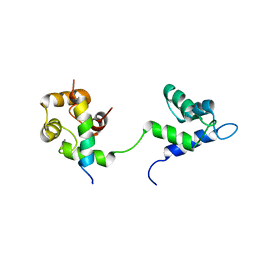 | |
3SFH
 
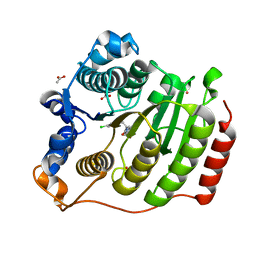 | | Crystal Structure of Human HDAC8 Inhibitor Complex, an Amino Acid Derived Inhibitor | | Descriptor: | (2R)-2-amino-3-(2,4-dichlorophenyl)-1-(1,3-dihydro-2H-isoindol-2-yl)propan-1-one, ACETATE ION, Histone deacetylase 8, ... | | Authors: | Stams, T, Vash, B. | | Deposit date: | 2011-06-13 | | Release date: | 2011-07-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors.
Bioorg.Med.Chem., 19, 2011
|
|
3SFF
 
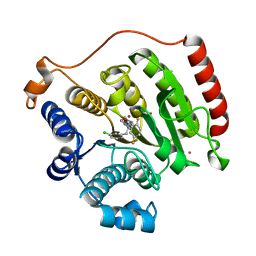 | | Crystal Structure of Human HDAC8 Inhibitor Complex, an Amino Acid Derived Inhibitor | | Descriptor: | (2R)-2-amino-3-(3-chlorophenyl)-1-[4-(2,5-difluorobenzoyl)piperazin-1-yl]propan-1-one, Histone deacetylase 8, POTASSIUM ION, ... | | Authors: | Stams, T, Vash, B. | | Deposit date: | 2011-06-13 | | Release date: | 2011-07-20 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Human HDAC isoform selectivity achieved via exploitation of the acetate release channel with structurally unique small molecule inhibitors.
Bioorg.Med.Chem., 19, 2011
|
|
4NMC
 
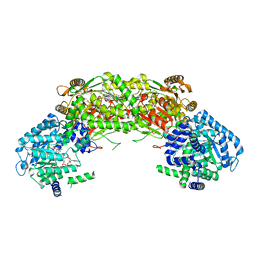 | |
6ESQ
 
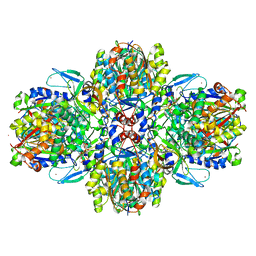 | | Structure of the acetoacetyl-CoA thiolase/HMG-CoA synthase complex from Methanothermococcus thermolithotrophicus soaked with acetyl-CoA | | Descriptor: | CHLORIDE ION, COENZYME A, HydroxyMethylGlutaryl-CoA synthase, ... | | Authors: | Voegeli, B, Engilberge, S, Girard, E, Riobe, F, Maury, O, Erb, J.T, Shima, S, Wagner, T. | | Deposit date: | 2017-10-24 | | Release date: | 2018-03-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Archaeal acetoacetyl-CoA thiolase/HMG-CoA synthase complex channels the intermediate via a fused CoA-binding site.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6ET9
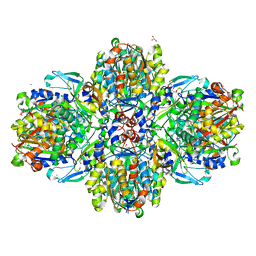 
 | | Structure of the acetoacetyl-CoA-thiolase/HMG-CoA-synthase complex from Methanothermococcus thermolithotrophicus at 2.75 A | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Acetyl-CoA acetyltransferase thiolase, CHLORIDE ION, ... | | Authors: | Engilberge, S, Voegeli, B, Girard, E, Riobe, F, Maury, O, Erb, T.J, Shima, S, Wagner, T. | | Deposit date: | 2017-10-25 | | Release date: | 2018-03-14 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Archaeal acetoacetyl-CoA thiolase/HMG-CoA synthase complex channels the intermediate via a fused CoA-binding site.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3BXK
 
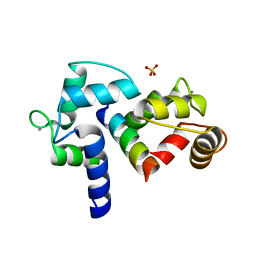 | | Crystal structure of the P/Q-type calcium channel (CaV2.1) IQ domain and CA2+calmodulin complex | | Descriptor: | CALCIUM ION, Calmodulin, SULFATE ION, ... | | Authors: | Mori, M.X, Vander Kooi, C.W, Leahy, D.J, Yue, D.T. | | Deposit date: | 2008-01-14 | | Release date: | 2008-03-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of the P/Q-type calcium channel (CaV2.1) IQ domain and CA2+calmodulin complex
To be Published
|
|
1UYU
 
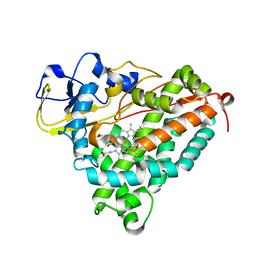 | | Xenon COMPLEX OF wildtype P450CAM FROM PSEUDOMONAS PUTIDA | | Descriptor: | CAMPHOR, CYTOCHROME P450-CAM, POTASSIUM ION, ... | | Authors: | Wade, R.C, Winn, P.J, Schlichting, I, Sudarko, X. | | Deposit date: | 2004-03-02 | | Release date: | 2005-03-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A Survey of Active Site Access Channels in Cytochromes P450
J.Inorg.Biochem., 98, 2004
|
|
4J90
 
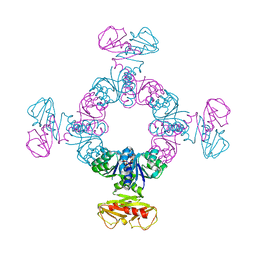 | |
4J91
 
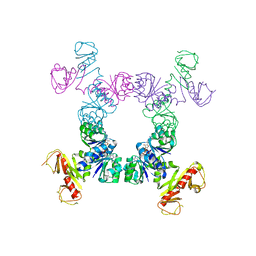 | |
6BBG
 
 | |
6BBI
 
 | |
3WFN
 
 | |
7NMN
 
 | | Rabbit HCN4 stabilised in amphipol A8-35 | | Descriptor: | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4,Rabbit HCN4 | | Authors: | Chaves-Sanjuan, A. | | Deposit date: | 2021-02-23 | | Release date: | 2021-06-30 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Gating movements and ion permeation in HCN4 pacemaker channels.
Mol.Cell, 81, 2021
|
|
