6F04
 
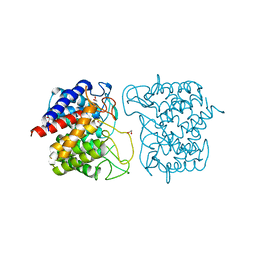 | | N-acetylglucosamine-2-epimerase | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, N-acetylglucosamine-2-epimerase | | Authors: | Halsoer, M.J, Rothweiler, U. | | Deposit date: | 2017-11-17 | | Release date: | 2018-09-26 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.699 Å) | | Cite: | The crystal structure of the N-acetylglucosamine 2-epimerase from Nostoc sp. KVJ10 reveals the true dimer.
Acta Crystallogr D Struct Biol, 75, 2019
|
|
5NNK
 
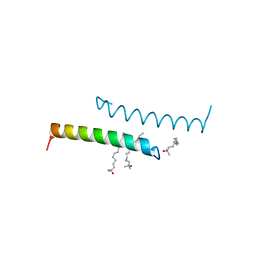 | | The structure of LL-37 crystallized in the presence LDAO | | Descriptor: | Cathelicidin antimicrobial peptide, LAURYL DIMETHYLAMINE-N-OXIDE | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.798 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
8OUI
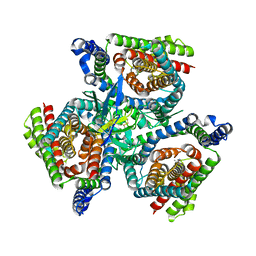 
 | | Complex of ASCT2 with Suppressyn | | Descriptor: | ALANINE, Neutral amino acid transporter B(0), Suppressyn | | Authors: | Khare, S, Kumar, A, Reyes, N. | | Deposit date: | 2023-04-23 | | Release date: | 2024-05-01 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.39 Å) | | Cite: | Receptor-recognition and antiviral mechanisms of retrovirus-derived human proteins.
Nat.Struct.Mol.Biol., 2024
|
|
8OUD
 
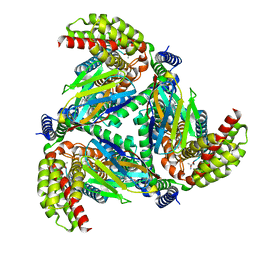 | |
8OUH
 
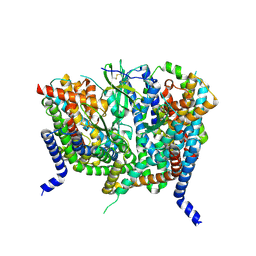 | | Complex of human ASCT2 with Syncytin-1 | | Descriptor: | ALANINE, Neutral amino acid transporter B(0), Syncytin-1 | | Authors: | Khare, S, Reyes, N. | | Deposit date: | 2023-04-23 | | Release date: | 2024-05-01 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (2.62 Å) | | Cite: | Receptor-recognition and antiviral mechanisms of retrovirus-derived human proteins.
Nat.Struct.Mol.Biol., 2024
|
|
8QTO
 
 | | CRYSTAL STRUCTURE OF HOLO-L28H-FNR OF A. FISCHERI | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, FNR type regulator, IRON/SULFUR CLUSTER | | Authors: | Volbeda, A, Fontecilla-Camps, J.C. | | Deposit date: | 2023-10-13 | | Release date: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Probing the Reactivity of [4Fe-4S] Fumarate and Nitrate Reduction (FNR) Regulator with O2 and NO: Increased O2 Resistance and Relative Specificity for NO of the [4Fe-4S] L28H FNR Cluster
Inorganics (Basel), 11, 2023
|
|
8PDJ
 
 | | The phosphatase and C2 domains of SHIP1 with covalent Z56948267 | | Descriptor: | 4-azanyl-3-fluoranyl-benzenethiol, DIMETHYL SULFOXIDE, Phosphatidylinositol 3,4,5-trisphosphate 5-phosphatase 1 | | Authors: | Bradshaw, W.J, Moreira, T, Pascoa, T.C, Bountra, C, Chalk, R, von Delft, F, Brennan, P.E, Gileadi, O. | | Deposit date: | 2023-06-12 | | Release date: | 2023-06-28 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Regulation of inositol 5-phosphatase activity by the C2 domain of SHIP1 and SHIP2.
Structure, 32, 2024
|
|
8PDH
 
 | | The phosphatase and C2 domains of SHIP1 with covalent Z1742148362 | | Descriptor: | (5-phenyl-1,3,4-oxadiazol-2-yl)methanimine, 1,2-ETHANEDIOL, DIMETHYL SULFOXIDE, ... | | Authors: | Bradshaw, W.J, Moreira, T, Pascoa, T.C, Bountra, C, Chalk, R, von Delft, F, Brennan, P.E, Gileadi, O. | | Deposit date: | 2023-06-12 | | Release date: | 2023-06-28 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Regulation of inositol 5-phosphatase activity by the C2 domain of SHIP1 and SHIP2.
Structure, 32, 2024
|
|
8PDG
 
 | | The phosphatase and C2 domains of SHIP1 with covalent Z2738285202 | | Descriptor: | DIMETHYL SULFOXIDE, Phosphatidylinositol 3,4,5-trisphosphate 5-phosphatase 1, ~{N}-(8-chloranylquinolin-2-yl)propanamide | | Authors: | Bradshaw, W.J, Moreira, T, Scacioc, A, Bountra, C, Chalk, R, von Delft, F, Brennan, P.E, Gileadi, O. | | Deposit date: | 2023-06-12 | | Release date: | 2023-06-28 | | Last modified: | 2024-04-17 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Regulation of inositol 5-phosphatase activity by the C2 domain of SHIP1 and SHIP2.
Structure, 32, 2024
|
|
7NTS
 
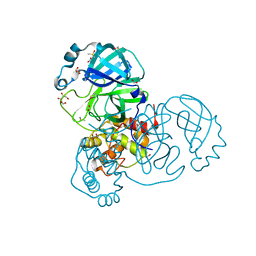 | | Crystal structure of the SARS-CoV-2 Main Protease with oxidized C145 | | Descriptor: | DIMETHYL SULFOXIDE, FORMIC ACID, GLYCEROL, ... | | Authors: | Dupre, E, Villeret, V, Hanoulle, X. | | Deposit date: | 2021-03-10 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.477 Å) | | Cite: | NMR Spectroscopy of the Main Protease of SARS-CoV-2 and Fragment-Based Screening Identify Three Protein Hotspots and an Antiviral Fragment.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
6HQE
 
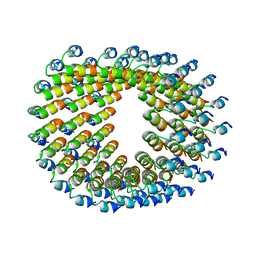 | | Cryo-EM of self-assembly peptide filament LRV_M3delta1 | | Descriptor: | peptide LRV_M3delta1 | | Authors: | Osinski, T, Wang, F, Hughes, S.A, Kreutzberger, M.A.B, Conticello, V.P, Egelman, E.H. | | Deposit date: | 2018-09-24 | | Release date: | 2019-06-26 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Ambidextrous helical nanotubes from self-assembly of designed helical hairpin motifs.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
7OHF
 
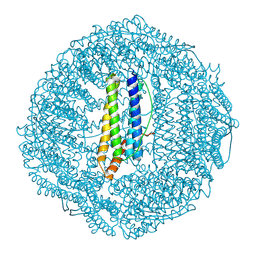 | | Cryo-EM structure of pyrococcus furiosus apoferritin in nanofluidic channels | | Descriptor: | Ferritin | | Authors: | Huber, S.T, Sarajlic, E, Huijink, R, Evers, W.H, Jakobi, A.J. | | Deposit date: | 2021-05-10 | | Release date: | 2021-08-11 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Nanofluidic chips for cryo-EM structure determination from picoliter sample volumes.
Elife, 11, 2022
|
|
6D9S
 
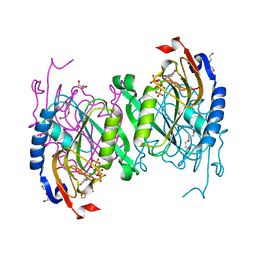 | | The (p)ppGpp-bound crystal structure of HPRT (hypoxanthine phosphoribosyltransferase) | | Descriptor: | DI(HYDROXYETHYL)ETHER, GUANOSINE-5',3'-TETRAPHOSPHATE, Hypoxanthine phosphoribosyltransferase, ... | | Authors: | Satyshur, K.A, Dubiel, K, Anderson, B, Wolak, C, Keck, J.L. | | Deposit date: | 2018-04-30 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.105 Å) | | Cite: | Evolution of (p)ppGpp-HPRT regulation through diversification of an allosteric oligomeric interaction.
Elife, 8, 2019
|
|
8QY4
 
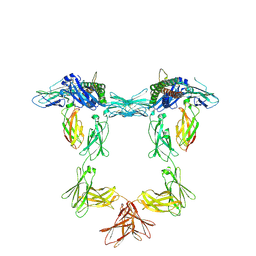 | | Structure of interleukin 11 (gp130 P496L mutant). | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Interleukin-11, ... | | Authors: | Gardner, S, Bubeck, D, Jin, Y. | | Deposit date: | 2023-10-25 | | Release date: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.06 Å) | | Cite: | Structure of interleukin 11 (gp130 P496L mutant).
To Be Published
|
|
8QY5
 
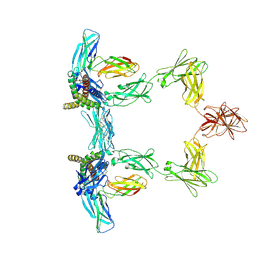 | | Structure of interleukin 6. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Interleukin-6, ... | | Authors: | Gardner, S, Bubeck, D, Jin, Y. | | Deposit date: | 2023-10-25 | | Release date: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structure of interleukin 6.
To Be Published
|
|
6GS2
 
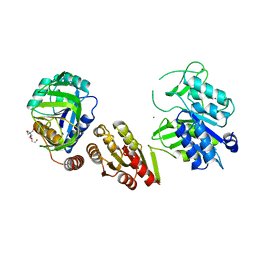 | | Crystal Structure of the GatD/MurT Enzyme Complex from Staphylococcus aureus | | Descriptor: | DI(HYDROXYETHYL)ETHER, MAGNESIUM ION, SA1707 protein, ... | | Authors: | Muckenfuss, L.M, Noeldeke, E.R, Niemann, V, Zocher, G, Stehle, T. | | Deposit date: | 2018-06-13 | | Release date: | 2018-09-05 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Structural basis of cell wall peptidoglycan amidation by the GatD/MurT complex of Staphylococcus aureus.
Sci Rep, 8, 2018
|
|
8QY6
 
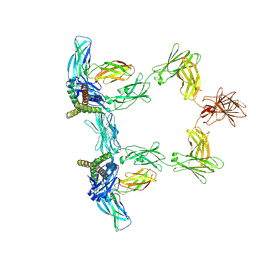 | | Structure of interleukin 6 (gp130 P496L mutant). | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Interleukin-6, ... | | Authors: | Gardner, S, Bubeck, D, Jin, Y. | | Deposit date: | 2023-10-25 | | Release date: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.16 Å) | | Cite: | Structure of interleukin 6 (gp130 P496L mutant).
To Be Published
|
|
6DIY
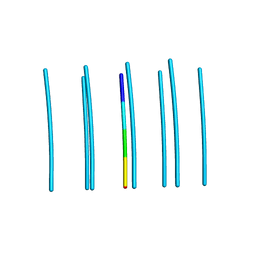 
 | | YTFGQ segment from Human Immunoglobulin Light-Chain Variable Domain, Residues 96-100, assembled as an amyloid fibril | | Descriptor: | YTFGQ segment Light-Chain Variable Domain Kappa AL09 | | Authors: | Brumshtein, B, Esswein, S.R, Sawaya, M.R, Eisenberg, D.S. | | Deposit date: | 2018-05-24 | | Release date: | 2018-10-31 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (0.9 Å) | | Cite: | Identification of two principal amyloid-driving segments in variable domains of Ig light chains in systemic light-chain amyloidosis.
J. Biol. Chem., 293, 2018
|
|
5BQ1
 
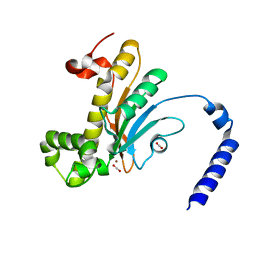 | | Capturing Carbon Dioxide in beta Carbonic Anhydrase | | Descriptor: | CARBON DIOXIDE, Carbonic anhydrase, ZINC ION | | Authors: | Aggarwal, M, Chua, T.K, Pinard, M.A, Szebenyi, D.M, McKenna, R. | | Deposit date: | 2015-05-28 | | Release date: | 2015-10-28 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Carbon Dioxide "Trapped" in a beta-Carbonic Anhydrase.
Biochemistry, 54, 2015
|
|
6FND
 
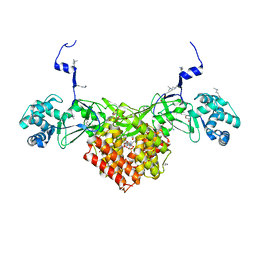 | | Crystal structure of Toxoplasma gondii AKMT | | Descriptor: | 1,2-ETHANEDIOL, Apical complex lysine methyltransferase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Pivovarova, Y, Dong, G. | | Deposit date: | 2018-02-02 | | Release date: | 2018-11-14 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Structure of a Novel Dimeric SET Domain Methyltransferase that Regulates Cell Motility.
J. Mol. Biol., 430, 2018
|
|
5NMN
 
 | |
5NNM
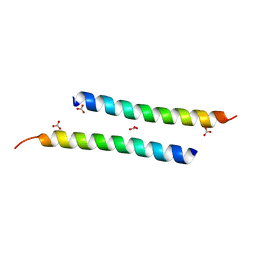 
 | | The crystal structure of dimeric LL-37 | | Descriptor: | CARBONATE ION, Cathelicidin antimicrobial peptide | | Authors: | Zeth, K, Sancho-Vaello, E. | | Deposit date: | 2017-04-10 | | Release date: | 2018-01-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural remodeling and oligomerization of human cathelicidin on membranes suggest fibril-like structures as active species.
Sci Rep, 7, 2017
|
|
5NQ7
 
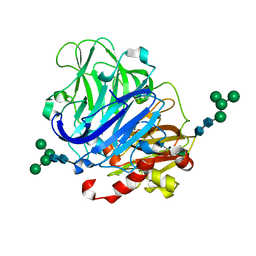 | | Crystal structure of laccases from Pycnoporus sanguineus, izoform I | | Descriptor: | COPPER (II) ION, Laccase, PEROXIDE ION, ... | | Authors: | Orlikowska, M, de J.Rostro-Alanis, M, Bujacz, A, Hernandez-Luna, C, Rubio, R, Parra, R, Bujacz, G. | | Deposit date: | 2017-04-19 | | Release date: | 2017-11-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural studies of two thermostable laccases from the white-rot fungus Pycnoporus sanguineus.
Int. J. Biol. Macromol., 107, 2018
|
|
8R61
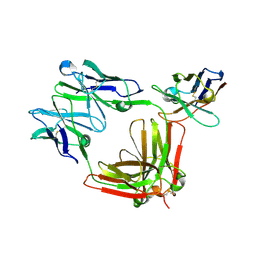 
 | |
6H5E
 
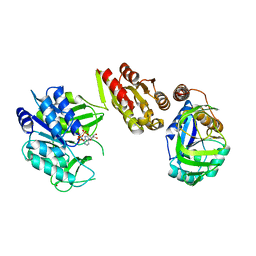 | | Crystal Structure of the GatD/MurT Enzyme Complex from Staphylococcus aureus with bound AMPPNP | | Descriptor: | DI(HYDROXYETHYL)ETHER, DUF1727 domain-containing protein, GLYCEROL, ... | | Authors: | Noeldeke, E.R, Niemann, V, Stoerk, E, Stehle, T. | | Deposit date: | 2018-07-24 | | Release date: | 2018-09-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.139 Å) | | Cite: | Structural basis of cell wall peptidoglycan amidation by the GatD/MurT complex of Staphylococcus aureus.
Sci Rep, 8, 2018
|
|
