8D6Y
 
 | |
8D6W
 
 | |
8D6V
 
 | |
8D6X
 
 | |
7OJG
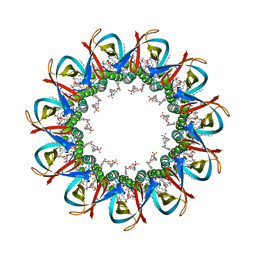 
 | | CRYO-EM STRUCTURE OF UNDECAMERIC SLYB FROM ESCHERICHIA COLI K12 | | Descriptor: | (2~{R},4~{R},5~{R},6~{R})-6-[(1~{R})-1,2-bis(oxidanyl)ethyl]-2-[[(2~{R},3~{S},4~{R},5~{R},6~{R})-5-[[(~{E},3~{R})-3-dodecanoyloxytetradec-5-enoyl]amino]-6-[[(2~{R},3~{S},4~{R},5~{R},6~{R})-3-oxidanyl-5-[[(~{E},3~{R})-3-oxidanyltetradec-11-enoyl]amino]-4-[(~{E},3~{R})-3-oxidanyltetradec-5-enoyl]oxy-6-phosphonooxy-oxan-2-yl]methoxy]-3-phosphonooxy-4-[(~{E},3~{R})-3-tetradecanoyloxytetradec-7-enoyl]oxy-oxan-2-yl]methoxy]-4,5-bis(oxidanyl)oxane-2-carboxylic acid, 2-(HEXADECANOYLOXY)-1-[(PHOSPHONOOXY)METHYL]ETHYL HEXADECANOATE, GLYCEROL, ... | | Authors: | Nguyen, V.S, Remaut, H. | | Deposit date: | 2021-05-14 | | Release date: | 2022-08-03 | | Last modified: | 2023-12-20 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | SlyB encapsulates outer membrane proteins in stress-induced lipid nanodomains
Nature, 2023
|
|
7UA5
 
 | | Structure of dephosphorylated human RyR2 in the closed state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Peptidyl-prolyl cis-trans isomerase FKBP1B, Ryanodine receptor 2, ... | | Authors: | Miotto, M.C, Marks, A.R. | | Deposit date: | 2022-03-11 | | Release date: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (2.83 Å) | | Cite: | Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Sci Adv, 8, 2022
|
|
7U9Z
 
 | |
7U9X
 
 | |
7U9Q
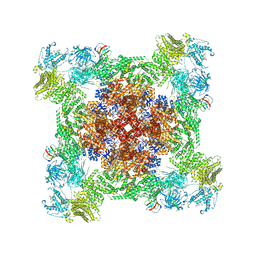 
 | | Structure of PKA phosphorylated human RyR2 in the closed state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Peptidyl-prolyl cis-trans isomerase FKBP1B, Ryanodine receptor 2, ... | | Authors: | Miotto, M.C, Marks, A.R. | | Deposit date: | 2022-03-11 | | Release date: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Sci Adv, 8, 2022
|
|
7U9T
 
 | |
7UA3
 
 | |
7UA9
 
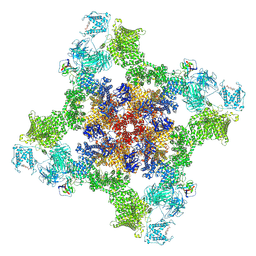 | | Structure of dephosphorylated human RyR2 in the open state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, Peptidyl-prolyl cis-trans isomerase FKBP1B, ... | | Authors: | Miotto, M.C, Marks, A.R. | | Deposit date: | 2022-03-11 | | Release date: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (3.59 Å) | | Cite: | Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Sci Adv, 8, 2022
|
|
7UA1
 
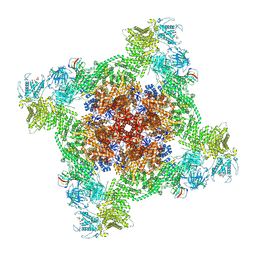 | | Structure of PKA phosphorylated human RyR2-R2474S in the closed state in the presence of ARM210 | | Descriptor: | 4-[(7-methoxy-2,3-dihydro-1,4-benzothiazepin-4(5H)-yl)methyl]benzoic acid, ADENOSINE-5'-TRIPHOSPHATE, Peptidyl-prolyl cis-trans isomerase FKBP1B, ... | | Authors: | Miotto, M.C, Marks, A.R. | | Deposit date: | 2022-03-11 | | Release date: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (2.99 Å) | | Cite: | Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Sci Adv, 8, 2022
|
|
7UA4
 
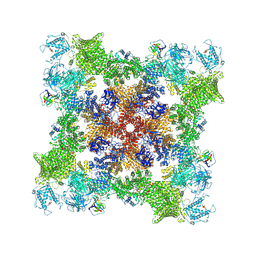 | |
7U9R
 
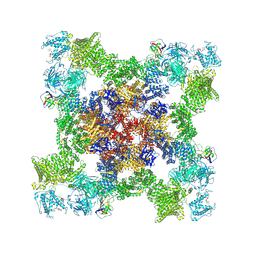 | | Structure of PKA phosphorylated human RyR2 in the open state | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, Peptidyl-prolyl cis-trans isomerase FKBP1B, ... | | Authors: | Miotto, M.C, Marks, A.R. | | Deposit date: | 2022-03-11 | | Release date: | 2022-08-03 | | Method: | ELECTRON MICROSCOPY (3.69 Å) | | Cite: | Structural analyses of human ryanodine receptor type 2 channels reveal the mechanisms for sudden cardiac death and treatment.
Sci Adv, 8, 2022
|
|
7NOZ
 
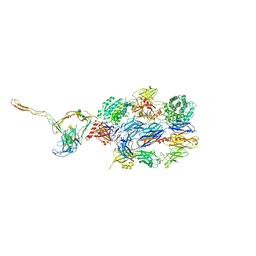 | | Structure of the nanobody stablized properdin bound alternative pathway proconvertase C3b:FB:FP | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Complement C3 alpha chain, Complement C3 beta chain, ... | | Authors: | Lorenzen, J, Pedersen, D.V, Andersen, G.R. | | Deposit date: | 2021-02-26 | | Release date: | 2022-07-27 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Structure determination of an unstable macromolecular complex enabled by nanobody-peptide bridging.
Protein Sci., 31, 2022
|
|
8AB3
 
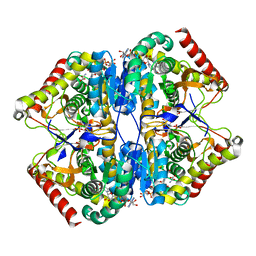 | | Crystal Structure of the Lactate Dehydrogenase of Cyanobacterium Aponinum in complex with oxamate, NADH and FBP. | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, 1,6-di-O-phosphono-beta-D-fructofuranose, L-lactate dehydrogenase, ... | | Authors: | Robin, A.Y, Girard, E, Madern, D. | | Deposit date: | 2022-07-04 | | Release date: | 2022-07-27 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.616 Å) | | Cite: | Deciphering Evolutionary Trajectories of Lactate Dehydrogenases Provides New Insights into Allostery.
Mol.Biol.Evol., 40, 2023
|
|
7P4N
 | |
7TD6
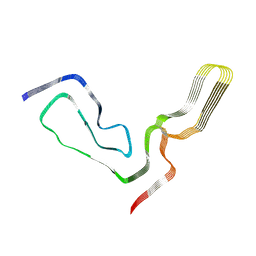 
 | | aRML prion fibril | | Descriptor: | Major prion protein | | Authors: | Hoyt, F, Standke, H.G, Artikis, E, Caughey, B, Kraus, A. | | Deposit date: | 2021-12-30 | | Release date: | 2022-07-20 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Cryo-EM structure of anchorless RML prion reveals variations in shared motifs between distinct strains.
Nat Commun, 13, 2022
|
|
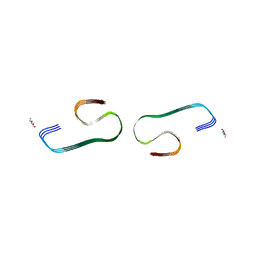
7F29
 
 | |
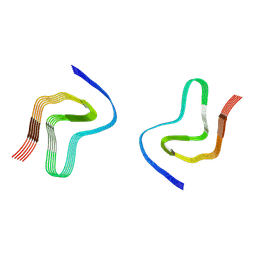
7OZH
 
 | |
7OZG
 
 | |
5SRR
 
 | |
8D5K
 
 | |
8D5E
 
 | | The complex of Gtf2b Peptide TGAASFDEF Presented by H2-Dd | | Descriptor: | Beta-2-microglobulin, GLYCEROL, H-2 class I histocompatibility antigen, ... | | Authors: | Custodio, J.M.F, Baker, B.M. | | Deposit date: | 2022-06-04 | | Release date: | 2022-07-06 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structural and physical features that distinguish tumor-controlling from inactive cancer neoepitopes.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
