8A09
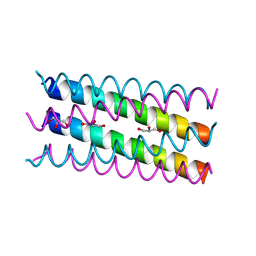 
 | | A hexameric barrel state of a de novo coiled-coil assembly: CC-Type2-(QgLaId)4 | | Descriptor: | A hexameric barrel state of a de novo coiled-coil assembly: CC-Type2-(QgLaId)4, GLYCEROL | | Authors: | Martin, F.J.O, Rhys, G.G, Dawson, W.M, Woolfson, D.N. | | Deposit date: | 2022-05-27 | | Release date: | 2022-06-15 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Differential sensing with arrays of de novo designed peptide assemblies.
Nat Commun, 14, 2023
|
|
8A8X
 
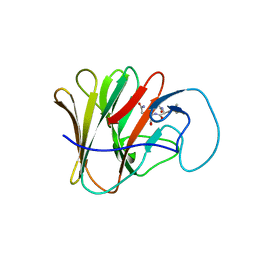 | | TRIM7 PRYSPRY in complex with a MNV1-NS3 peptide HDDFGLQ | | Descriptor: | E3 ubiquitin-protein ligase TRIM7, MNV1-NS3 C term peptide | | Authors: | Luptak, J. | | Deposit date: | 2022-06-24 | | Release date: | 2022-07-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | TRIM7 Restricts Coxsackievirus and Norovirus Infection by Detecting the C-Terminal Glutamine Generated by 3C Protease Processing.
Viruses, 14, 2022
|
|
8A5M
 
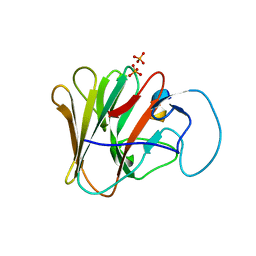 | | TRIM7 PRYSPRY in complex with a MNV1-NS6 peptide LEALEFQ | | Descriptor: | E3 ubiquitin-protein ligase TRIM7, MNV1-NS6 peptide LEALEFQ, PHOSPHATE ION | | Authors: | Luptak, J. | | Deposit date: | 2022-06-15 | | Release date: | 2022-07-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.918 Å) | | Cite: | TRIM7 Restricts Coxsackievirus and Norovirus Infection by Detecting the C-Terminal Glutamine Generated by 3C Protease Processing.
Viruses, 14, 2022
|
|
8A5L
 
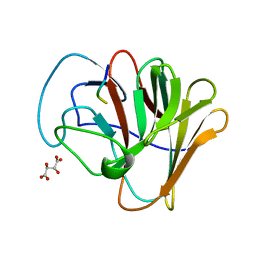 | | TRIM7 PRYSPRY in complex with a 2BC peptide TIEALFQ | | Descriptor: | 2BC peptide TIEALFQ, D(-)-TARTARIC ACID, E3 ubiquitin-protein ligase TRIM7 | | Authors: | Luptak, J. | | Deposit date: | 2022-06-15 | | Release date: | 2022-07-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.622 Å) | | Cite: | TRIM7 Restricts Coxsackievirus and Norovirus Infection by Detecting the C-Terminal Glutamine Generated by 3C Protease Processing.
Viruses, 14, 2022
|
|
7WUE
 
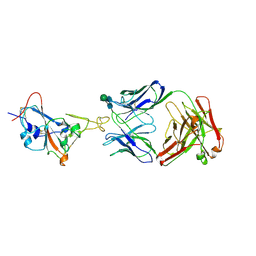 | | Crystal structure of SARS-CoV-2 Receptor Binding Domain in complex with the monoclonal antibody m31A7 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Spike protein S1, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mohapatra, A. | | Deposit date: | 2022-02-08 | | Release date: | 2022-03-16 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Vaccination with SARS-CoV-2 spike protein lacking glycan shields elicits enhanced protective responses in animal models.
Sci Transl Med, 14, 2022
|
|
7WUH
 
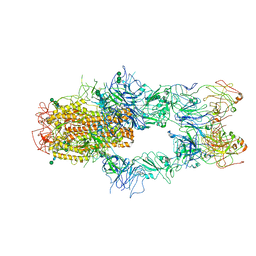 | | SARS-CoV-2 Spike in complex with Fab of m31A7 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Chen, X, Wu, Y.-M. | | Deposit date: | 2022-02-08 | | Release date: | 2022-03-23 | | Last modified: | 2022-07-06 | | Method: | ELECTRON MICROSCOPY (4.7 Å) | | Cite: | Vaccination with SARS-CoV-2 spike protein lacking glycan shields elicits enhanced protective responses in animal models.
Sci Transl Med, 14, 2022
|
|
7ZIR
 
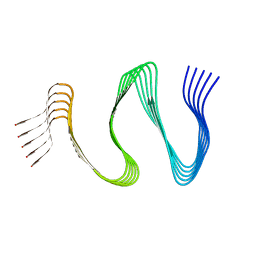 | | Cryo-EM structure of hnRNPDL amyloid fibrils | | Descriptor: | Heterogeneous nuclear ribonucleoprotein D-like | | Authors: | Garcia-Pardo, J, Chaves-Sanjuan, A, Bartolome-Nafria, A, Gil-Garcia, M, Visentin, C, Bolognesi, M, Ricagno, S, Ventura, S. | | Deposit date: | 2022-04-08 | | Release date: | 2022-12-28 | | Last modified: | 2023-12-13 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Cryo-EM structure of hnRNPDL-2 fibrils, a functional amyloid associated with limb-girdle muscular dystrophy D3.
Nat Commun, 14, 2023
|
|
7WKU
 
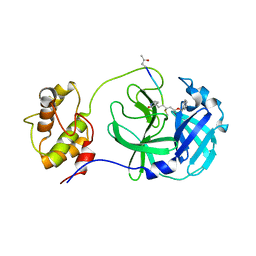 | | Structure of PDCoV Mpro in complex with an inhibitor | | Descriptor: | N-[(5-METHYLISOXAZOL-3-YL)CARBONYL]ALANYL-L-VALYL-N~1~-((1R,2Z)-4-(BENZYLOXY)-4-OXO-1-{[(3R)-2-OXOPYRROLIDIN-3-YL]METHYL}BUT-2-ENYL)-L-LEUCINAMIDE, Peptidase C30 | | Authors: | Wang, F.H, Yang, H.T. | | Deposit date: | 2022-01-11 | | Release date: | 2022-05-25 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Structure of the Porcine Deltacoronavirus Main Protease Reveals a Conserved Target for the Design of Antivirals.
Viruses, 14, 2022
|
|
8A6I
 
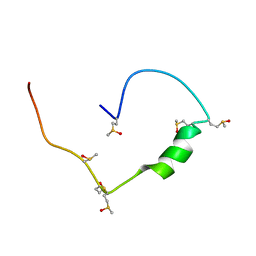 | | Structure of the low complexity domain of TDP-43 (fragment 309-350) with methionine sulfoxide modifications | | Descriptor: | TAR DNA-binding protein 43 | | Authors: | Carrasco, J, Anton, R, Pantoja-Uceda, D, Laurents, D.V, Oroz, J. | | Deposit date: | 2022-06-17 | | Release date: | 2023-02-08 | | Method: | SOLUTION NMR | | Cite: | Metamorphism in TDP-43 prion-like domain determines chaperone recognition.
Nat Commun, 14, 2023
|
|
6JHE
 
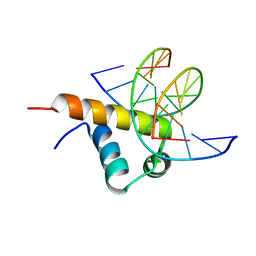 | | Crystal Structure of Bacillus subtilis SigW domain 4 in complexed with -35 element DNA | | Descriptor: | DNA (5'-D(*AP*AP*AP*GP*GP*TP*TP*TP*CP*AP*A)-3'), DNA (5'-D(P*TP*TP*GP*AP*AP*AP*CP*CP*TP*TP*T)-3'), ECF RNA polymerase sigma factor SigW | | Authors: | Kwon, E, Devkota, S.R, Pathak, D, Dahal, P, Kim, D.Y. | | Deposit date: | 2019-02-18 | | Release date: | 2020-01-01 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.101 Å) | | Cite: | Structural analysis of the recognition of the -35 promoter element by SigW from Bacillus subtilis.
Plos One, 14, 2019
|
|
6Q9L
 
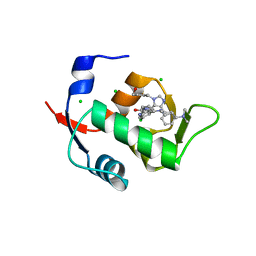 | |
7ZYN
 
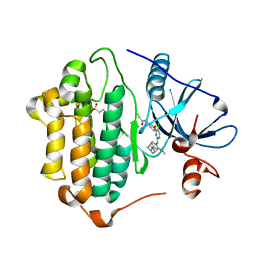 | | Crystal Structure of EGFR-T790M/C797S in Complex with WZ4002 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Epidermal growth factor receptor, N-{3-[(5-chloro-2-{[2-methoxy-4-(4-methylpiperazin-1-yl)phenyl]amino}pyrimidin-4-yl)oxy]phenyl}prop-2-enamide | | Authors: | Niggenaber, J, Kleinboelting, S, Mueller, M.P, Rauh, D. | | Deposit date: | 2022-05-25 | | Release date: | 2023-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Addressing the Osimertinib Resistance Mutation EGFR-L858R/C797S with Reversible Aminopyrimidines.
Acs Med.Chem.Lett., 14, 2023
|
|
7ZYM
 
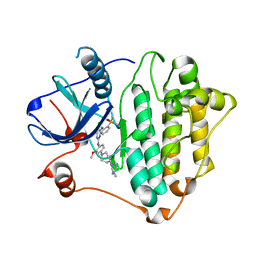 | | Crystal Structure of EGFR-T790M/C797S in Complex with Brigatinib | | Descriptor: | 5-chloro-N~4~-[2-(dimethylphosphoryl)phenyl]-N~2~-{2-methoxy-4-[4-(4-methylpiperazin-1-yl)piperidin-1-yl]phenyl}pyrimidine-2,4-diamine, Epidermal growth factor receptor | | Authors: | Niggenaber, J, Kleinboelting, S, Mueller, M.P, Rauh, D. | | Deposit date: | 2022-05-25 | | Release date: | 2023-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Addressing the Osimertinib Resistance Mutation EGFR-L858R/C797S with Reversible Aminopyrimidines.
Acs Med.Chem.Lett., 14, 2023
|
|
7ZYP
 
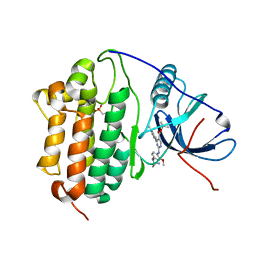 | | Crystal Structure of EGFR-T790M/C797S in Complex with Reversible Aminopyrimidine 9 | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Epidermal growth factor receptor, propan-2-yl 2-[[4-(4-azanylpiperidin-1-yl)-2-methoxy-phenyl]amino]-4-(1-methylindol-3-yl)pyrimidine-5-carboxylate | | Authors: | Niggenaber, J, Kleinboelting, S, Mueller, M.P, Rauh, D. | | Deposit date: | 2022-05-25 | | Release date: | 2023-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Addressing the Osimertinib Resistance Mutation EGFR-L858R/C797S with Reversible Aminopyrimidines.
Acs Med.Chem.Lett., 14, 2023
|
|
7ZYQ
 
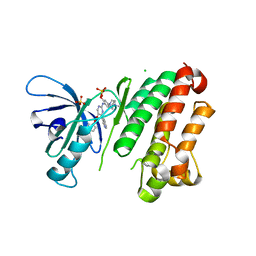 | | Crystal Structure of EGFR-T790M/V948R in Complex with Reversible Aminopyrimidine 13 | | Descriptor: | CHLORIDE ION, Epidermal growth factor receptor, SODIUM ION, ... | | Authors: | Niggenaber, J, Mueller, M.P, Rauh, D. | | Deposit date: | 2022-05-25 | | Release date: | 2023-05-03 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Addressing the Osimertinib Resistance Mutation EGFR-L858R/C797S with Reversible Aminopyrimidines.
Acs Med.Chem.Lett., 14, 2023
|
|
7ZQI
 
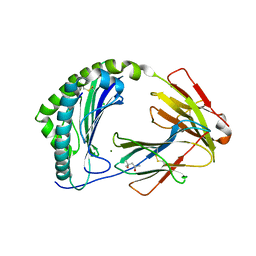 | | MHC class I from a wild bird in complex with a nonameric peptide P2 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Beta-2-microglobulin, CHLORIDE ION, ... | | Authors: | Eltschkner, S, Mellinger, S, Buus, S, Nielsen, M, Paulsson, K.M, Lindkvist-Petersson, K, Westerdahl, H. | | Deposit date: | 2022-04-29 | | Release date: | 2023-05-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The structure of songbird MHC class I reveals antigen binding that is flexible at the N-terminus and static at the C-terminus.
Front Immunol, 14, 2023
|
|
6Q9O
 
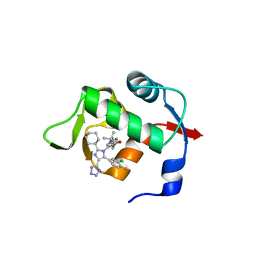 | |
7ZQJ
 
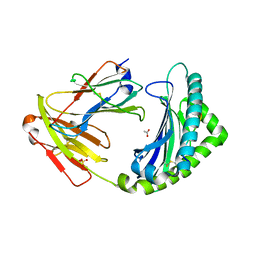 | | MHC class I from a wild bird in complex with a nonameric peptide P3 | | Descriptor: | ACETATE ION, Beta-2-microglobulin, MHC class I antigen, ... | | Authors: | Eltschkner, S, Mellinger, S, Buus, S, Nielsen, M, Paulsson, K.M, Lindkvist-Petersson, K, Westerdahl, H. | | Deposit date: | 2022-04-29 | | Release date: | 2023-05-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The structure of songbird MHC class I reveals antigen binding that is flexible at the N-terminus and static at the C-terminus.
Front Immunol, 14, 2023
|
|
7ZI1
 
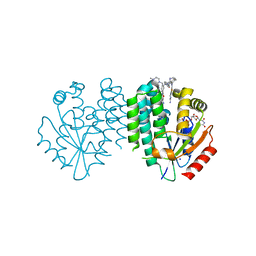 | | Crystal structure of dCK C4S-S74E mutant in complex with UDP and the dCKi1 inhibitor | | Descriptor: | Deoxycytidine kinase, N-[3-[[4-(4-azanylpyrimidin-2-yl)-1,3-thiazol-2-yl]amino]-4-methyl-phenyl]-4-[(4-methylpiperazin-1-yl)methyl]benzamide, SODIUM ION, ... | | Authors: | Saez-Ayala, M, Ben-Yaala, K, Betzi, S, Rebuffet, E, Morelli, X. | | Deposit date: | 2022-04-07 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia.
Nat Commun, 14, 2023
|
|
7ZI3
 
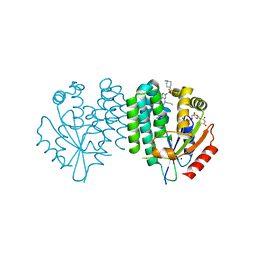 | | Crystal structure of dCK C4S-S74E mutant in complex with UDP and the OR0642 inhibitor | | Descriptor: | 2-[2-[[2-methyl-5-[4-(4-methylpiperazin-1-yl)sulfonyl-2-(trifluoromethyl)phenyl]phenyl]-propyl-amino]-1,3-thiazol-4-yl]pyrimidine-4,6-diamine, Deoxycytidine kinase, SODIUM ION, ... | | Authors: | Ben-Yaala, K, Saez-Ayala, M, Betzi, S, Rebuffet, E, Morelli, X. | | Deposit date: | 2022-04-07 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia.
Nat Commun, 14, 2023
|
|
6Q9W
 
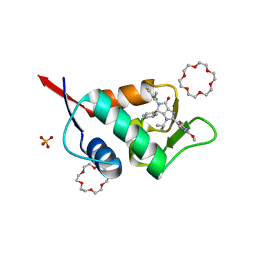 | | X-ray structure of compound 15 bound to HdmX: Structural states of Hdm2 and HdmX: X-ray elucidation of adaptations and binding interactions for different chemical compound classes | | Descriptor: | (4~{S})-4-(4-chlorophenyl)-5-[(1~{S})-1-(3-chlorophenyl)ethyl]-2-(2,4-dimethoxypyrimidin-5-yl)-3-propan-2-yl-4~{H}-pyrrolo[3,4-d]imidazol-6-one, 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, Protein Mdm4, ... | | Authors: | Kallen, J. | | Deposit date: | 2018-12-18 | | Release date: | 2019-05-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural States of Hdm2 and HdmX: X-ray Elucidation of Adaptations and Binding Interactions for Different Chemical Compound Classes.
Chemmedchem, 14, 2019
|
|
7ZI6
 
 | | Crystal structure of dCK C4S-S74E mutant in complex with UDP and the OR0325 inhibitor | | Descriptor: | 4-(4-azanylpyrimidin-2-yl)-N-[2-methyl-5-[4-[2-(4-methylpiperazin-1-yl)ethyl]phenyl]phenyl]-1,3-thiazol-2-amine, Deoxycytidine kinase, SODIUM ION, ... | | Authors: | Saez-Ayala, M, Ben-Yaala, K, Betzi, S, Rebuffet, E, Morelli, X. | | Deposit date: | 2022-04-07 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia.
Nat Commun, 14, 2023
|
|
7ZI9
 
 | | Crystal structure of dCK C4S-S74E mutant in complex with UDP and the OR0624 inhibitor | | Descriptor: | 2-[2-[[2-methyl-5-[4-(4-methylpiperazin-1-yl)sulfonylphenyl]phenyl]-propyl-amino]-1,3-thiazol-4-yl]pyrimidine-4,6-diamine, Deoxycytidine kinase, SODIUM ION, ... | | Authors: | Saez-Ayala, M, Ben-Yaala, K, Betzi, S, Rebuffet, E, Morelli, X. | | Deposit date: | 2022-04-07 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia.
Nat Commun, 14, 2023
|
|
7ZI5
 
 | | Crystal structure of dCK C4S-S74E mutant in complex with UDP and the OR0274 inhibitor | | Descriptor: | 4-(4-azanylpyrimidin-2-yl)-N-[2-methyl-5-[4-[(4-methylpiperazin-1-yl)methyl]phenyl]phenyl]-1,3-thiazol-2-amine, Deoxycytidine kinase, SODIUM ION, ... | | Authors: | Saez-Ayala, M, Ben-Yaala, K, Betzi, S, Rebuffet, E, Morelli, X. | | Deposit date: | 2022-04-07 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | From a drug repositioning to a structure-based drug design approach to tackle acute lymphoblastic leukemia.
Nat Commun, 14, 2023
|
|
7ZPF
 
 | | Three-dimensional structure of AIP56, a short-trip single chain AB toxin from Photobacterium damselae subsp. piscicida. | | Descriptor: | Aip56, GLYCEROL, NICKEL (II) ION, ... | | Authors: | Lisboa, J, Pereira, P.J.B, dos Santos, N.M.S. | | Deposit date: | 2022-04-27 | | Release date: | 2023-05-10 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Unconventional structure and mechanisms for membrane interaction and translocation of the NF-kappa B-targeting toxin AIP56.
Nat Commun, 14, 2023
|
|
