2K9G
 | |
2K9H
 
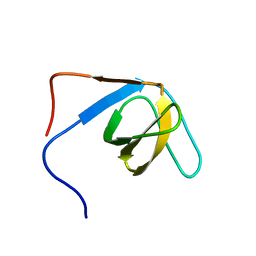
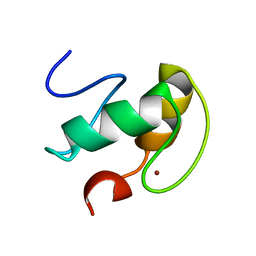 | | The hantavirus glycoprotein G1 tail contains a dual CCHC-type classical zinc fingers | | Descriptor: | Glycoprotein, ZINC ION | | Authors: | Estrada, D.F, Boudreaux, D.M, Zhong, D, St Jeor, S.C, De Guzman, R.N. | | Deposit date: | 2008-10-13 | | Release date: | 2009-01-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The Hantavirus Glycoprotein G1 Tail Contains Dual CCHC-type Classical Zinc Fingers.
J.Biol.Chem., 284, 2009
|
|
2K9I
 | |
2K9J
 
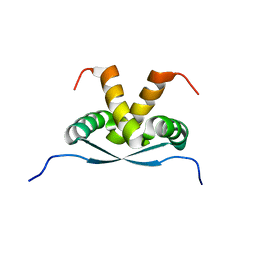
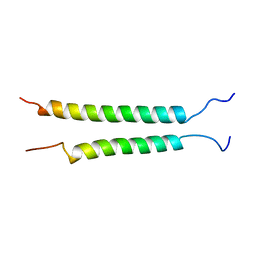 | | Integrin alphaIIb-beta3 transmembrane complex | | Descriptor: | Integrin alpha-IIb light chain, Integrin beta-3 | | Authors: | Lau, T, Kim, C, Ginsberg, M.H, Ulmer, T.S. | | Deposit date: | 2008-10-15 | | Release date: | 2009-03-24 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of the integrin alphaIIbbeta3 transmembrane complex explains integrin transmembrane signalling
Embo J., 28, 2009
|
|
2K9K
 | |
2K9L
 
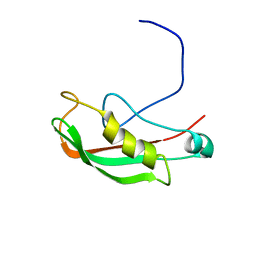
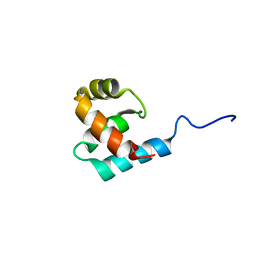 | | Structure of the Core Binding Domain of sigma54 | | Descriptor: | RNA polymerase sigma factor RpoN | | Authors: | Hong, E, Wemmer, D. | | Deposit date: | 2008-10-19 | | Release date: | 2009-05-26 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the RNA polymerase core-binding domain of sigma(54) reveals a likely conformational fracture point
J.Mol.Biol., 390, 2009
|
|
2K9M
 
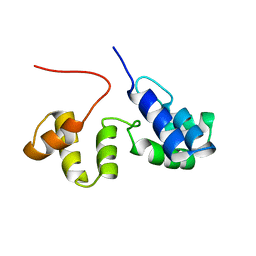 | | Structure of the Core Binding Domain of sigma54 | | Descriptor: | RNA polymerase sigma factor RpoN | | Authors: | Hong, E, Wemmer, D. | | Deposit date: | 2008-10-19 | | Release date: | 2009-05-26 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the RNA polymerase core-binding domain of sigma(54) reveals a likely conformational fracture point
J.Mol.Biol., 390, 2009
|
|
2K9N
 
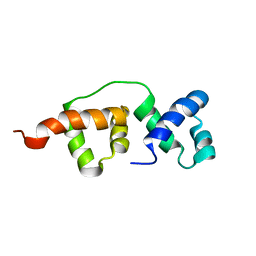 | | Solution NMR structure of the R2R3 DNA binding domain of Myb1 protein from protozoan parasite Trichomonas vaginalis | | Descriptor: | MYB24 | | Authors: | Lou, Y, Wei, S, Rajasekaran, M, Chou, C, Hsu, H, Tai, J, Chen, C. | | Deposit date: | 2008-10-19 | | Release date: | 2009-03-17 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR structural analysis of DNA recognition by a novel Myb1 DNA-binding domain in the protozoan parasite Trichomonas vaginalis.
Nucleic Acids Res., 37, 2009
|
|
2K9O
 
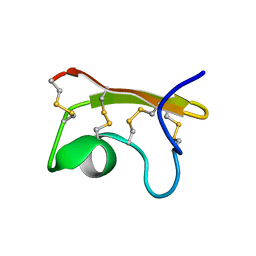 | | Solution structure of Vm24 synthetic scorpion toxin | | Descriptor: | Vm24 SCORPION toxin | | Authors: | del Rio-Portilla, F, Hernandez-Lopez, R, Possani-Postay, L, Gurrola, G. | | Deposit date: | 2008-10-20 | | Release date: | 2009-11-03 | | Last modified: | 2014-05-28 | | Method: | SOLUTION NMR | | Cite: | Structure, function, and chemical synthesis of Vaejovis mexicanus peptide 24: a novel potent blocker of Kv1.3 potassium channels of human T lymphocytes.
Biochemistry, 51, 2012
|
|
2K9P
 
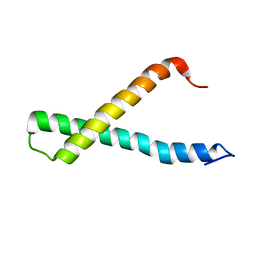 | |
2K9Q
 
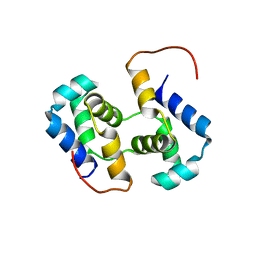 | | Solution NMR structure of HTH_XRE family transcriptional regulator BT_p548217 from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium Target BtR244. | | Descriptor: | uncharacterized protein | | Authors: | Ramelot, T.A, Cort, J.R, Zhao, L, Jiang, M, Foote, E.L, Xiao, R, Nair, R, Baran, M.C, Swapna, G, Acton, T.B, Rost, B, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-10-23 | | Release date: | 2008-11-18 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of HTH_XRE family transcriptional regulator BT_p548217 from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium Target BtR244.
To be Published
|
|
2K9R
 
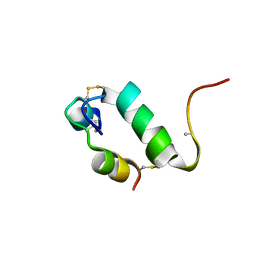 | | Enhancing the activity of insulin by stereospecific unfolding | | Descriptor: | Insulin | | Authors: | Hua, Q.X, Xu, B, Huang, K, Hu, S.Q, Nakarawa, S, Jia, W.H, Philips, N.F.P, Wittaker, L, Wittaker, J, Katsoyannis, P.G, Weiss, M.A. | | Deposit date: | 2008-10-23 | | Release date: | 2008-11-11 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Enhancing the Activity of a Protein by Stereospecific Unfolding: CONFORMATIONAL LIFE CYCLE OF INSULIN AND ITS EVOLUTIONARY ORIGINS.
J.Biol.Chem., 284, 2009
|
|
2K9S
 
 | |
2K9U
 
 | |
2K9X
 
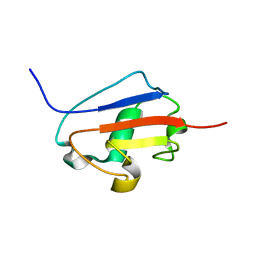 | | Solution structure of Urm1 from Trypanosoma brucei | | Descriptor: | Uncharacterized protein | | Authors: | Zhang, W, Zhang, J, Xu, C, Wang, T, Zhang, X, Tu, X. | | Deposit date: | 2008-10-27 | | Release date: | 2009-03-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Urm1 from Trypanosoma brucei
Proteins, 75, 2009
|
|
2K9Y
 
 | |
2K9Z
 
 | | NMR structure of the protein TM1112 | | Descriptor: | uncharacterized protein TM1112 | | Authors: | Mohanty, B, Pedrini, B, Serrano, P, Geralt, M, Horst, R, Herrmann, T, Wilson, I.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-28 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures for the proteins TM1112 and TM1367.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
2KA0
 
 | | NMR structure of the protein TM1367 | | Descriptor: | uncharacterized protein TM1367 | | Authors: | Mohanty, B, Pedrini, B, Serrano, P, Geralt, M, Horst, R, Herrmann, T, Wilson, I.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-27 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures for the proteins TM1112 and TM1367.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
2KA1
 
 | |
2KA2
 | |
2KA3
 
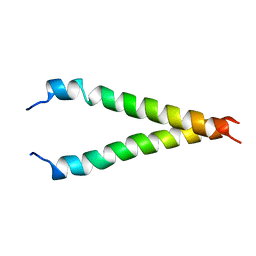
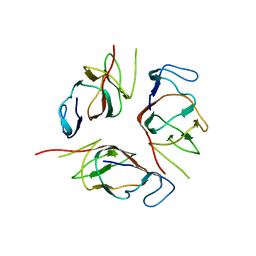 | | Structure of EMILIN-1 C1Q-like domain | | Descriptor: | EMILIN-1 | | Authors: | Verdone, G, Corazza, A, Colebrooke, S.A, Cicero, D.O, Eliseo, T, Boyd, J, Doliana, R, Fogolari, F, Viglino, P, Colombatti, A, Campbell, I.D, Esposito, G. | | Deposit date: | 2008-10-30 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | NMR-based homology model for the solution structure of the C-terminal globular domain of EMILIN1
J.Biomol.Nmr, 43, 2009
|
|
2KA4
 
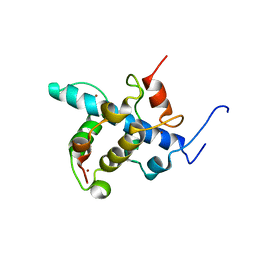 | | NMR structure of the CBP-TAZ1/STAT2-TAD complex | | Descriptor: | Crebbp protein, Signal transducer and activator of transcription 2, ZINC ION | | Authors: | Wojciak, J.M, Martinez-Yamout, M.A, Dyson, H.J, Wright, P.E. | | Deposit date: | 2008-10-30 | | Release date: | 2009-04-21 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structural basis for recruitment of CBP/p300 coactivators by STAT1 and STAT2 transactivation domains
Embo J., 28, 2009
|
|
2KA5
 
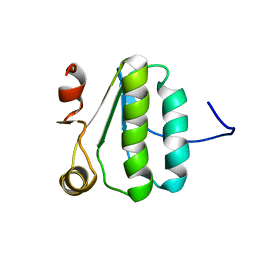 | | NMR Structure of the protein TM1081 | | Descriptor: | Putative anti-sigma factor antagonist TM_1081 | | Authors: | Serrano, P, Geralt, M, Mohanty, B, Pedrini, B, Horst, R, Wuthrich, K, Wilson, I, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-30 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures highlights conformational isomerism in protein active sites.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
2KA6
 
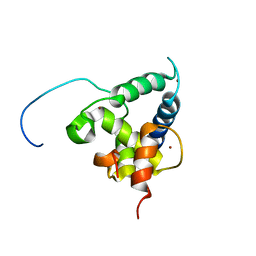 | | NMR structure of the CBP-TAZ2/STAT1-TAD complex | | Descriptor: | CREB-binding protein, Signal transducer and activator of transcription 1-alpha/beta, ZINC ION | | Authors: | Wojciak, J.M, Martinez-Yamout, M.A, Dyson, H.J, Wright, P.E. | | Deposit date: | 2008-10-30 | | Release date: | 2009-04-07 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structural basis for recruitment of CBP/p300 coactivators by STAT1 and STAT2 transactivation domains.
Embo J., 28, 2009
|
|
2KA7
 
 | | NMR solution structure of TM0212 at 40 C | | Descriptor: | Glycine cleavage system H protein | | Authors: | Pedrini, B, Herrmann, T, Mohanty, B, Geralt, M, Wilson, I, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-10-31 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | The J-UNIO protocol for automated protein structure determination by NMR in solution.
J.Biomol.Nmr, 53, 2012
|
|
