9BKV
 
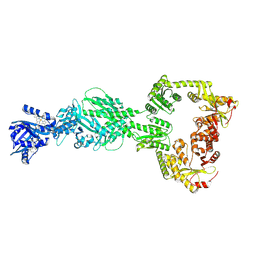 | | DosP R97A bent form | | Descriptor: | Oxygen sensor protein DosP, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kumar, P, Kober, D.L. | | Deposit date: | 2024-04-29 | | Release date: | 2024-11-20 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.43 Å) | | Cite: | Structures of the multi-domain oxygen sensor DosP: remote control of a c-di-GMP phosphodiesterase by a regulatory PAS domain.
Nat Commun, 15, 2024
|
|
9BGV
 
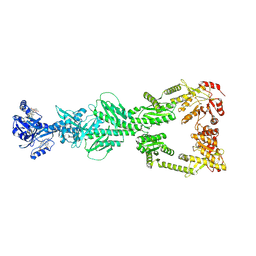 | | DosP R97A straight form | | Descriptor: | Oxygen sensor protein DosP, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Kumar, P, Kober, D.L. | | Deposit date: | 2024-04-19 | | Release date: | 2024-11-20 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.65 Å) | | Cite: | Structures of the multi-domain oxygen sensor DosP: remote control of a c-di-GMP phosphodiesterase by a regulatory PAS domain.
Nat Commun, 15, 2024
|
|
3H9O
 
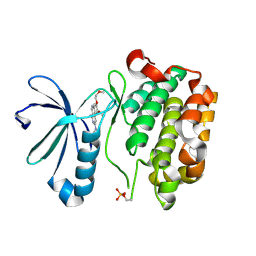 | |
9BAX
 
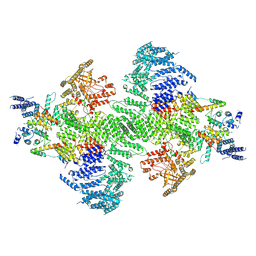 | | PI4KA complex bound to C-terminus of EFR3A | | Descriptor: | Hyccin, Phosphatidylinositol 4-kinase alpha, Protein EFR3 homolog A, ... | | Authors: | Shaw, A.L, Suresh, S, Yip, C.K, Burke, J.E. | | Deposit date: | 2024-04-04 | | Release date: | 2024-11-27 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.65 Å) | | Cite: | Molecular basis for plasma membrane recruitment of PI4KA by EFR3.
Sci Adv, 10, 2024
|
|
8DYF
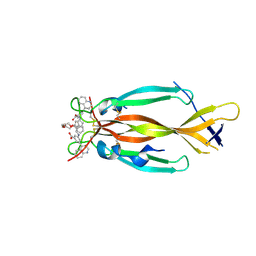 
 | | IL17A homodimer bound to Compound 10 | | Descriptor: | (5M)-3-[({2-[2-(2-{2-[2-({[(5M)-3-carboxy-5-(5,8-dihydroquinolin-4-yl)phenyl]amino}methyl)phenoxy]ethoxy}ethoxy)ethoxy]phenyl}methyl)amino]-5-(quinolin-4-yl)benzoic acid, Interleukin-17A | | Authors: | Argiriadi, M.A, Goedken, E.R. | | Deposit date: | 2022-08-04 | | Release date: | 2022-09-07 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Identification and structure-based drug design of cell-active inhibitors of interleukin 17A at a novel C-terminal site.
Sci Rep, 12, 2022
|
|
8UF3
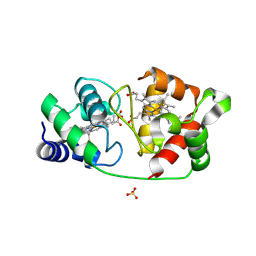 
 | |
6F63
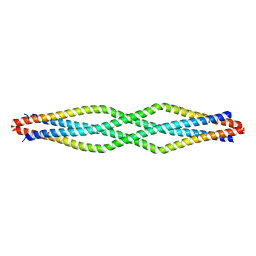 
 | |
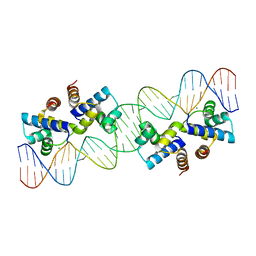
4X4B
 
 | |
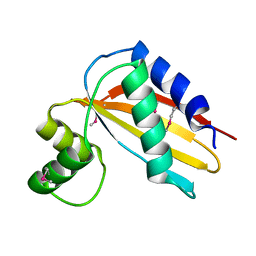
4Q6V
 
 | |
6UKM
 
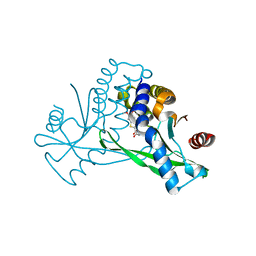 | | STING C-terminal Domain Complexed with Non-cyclic Dinucleotide Compound MSA-2 | | Descriptor: | 4-(5,6-dimethoxy-1-benzothiophen-2-yl)-4-oxobutanoic acid, fusion protein of Ubiquitin-like protein SMT3 and Stimulator of interferon protein c-terminal domain | | Authors: | Lesburg, C.A. | | Deposit date: | 2019-10-05 | | Release date: | 2020-08-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | An orally available non-nucleotide STING agonist with antitumor activity.
Science, 369, 2020
|
|
8S50
 
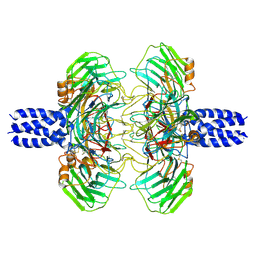 | | Cryo-EM structure of the C terminal region of PTX3 with a section of coiled-coil | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Pentraxin-related protein PTX3 | | Authors: | Snee, M, Shah, A, Lockhart-Cairns, M, Collins, R, Levy, C, Baldock, C, Day, A. | | Deposit date: | 2024-02-22 | | Release date: | 2025-01-22 | | Last modified: | 2025-03-12 | | Method: | ELECTRON MICROSCOPY (3.33 Å) | | Cite: | The structural organisation of pentraxin-3 and its interactions with heavy chains of inter-alpha-inhibitor regulate crosslinking of the hyaluronan matrix.
Matrix Biol., 136, 2025
|
|
260D
 
 | |
7Z21
 
 | | BAF A12T bound to the lamin A/C Ig-fold domain | | Descriptor: | Barrier-to-autointegration factor, N-terminally processed, CHLORIDE ION, ... | | Authors: | Marcelot, A, Legrand, P, Zinn-Justin, S. | | Deposit date: | 2022-02-25 | | Release date: | 2022-08-24 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.629 Å) | | Cite: | The BAF A12T mutation disrupts lamin A/C interaction, impairing robust repair of nuclear envelope ruptures in Nestor-Guillermo progeria syndrome cells.
Nucleic Acids Res., 50, 2022
|
|
6UKU
 
 | | STING C-terminal Domain Complexed with Non-cyclic Dinucleotide Compound 3 | | Descriptor: | 4,4'-[propane-1,3-diylbis(6-methoxy-1-benzothiene-5,2-diyl)]bis(4-oxobutanoic acid), fusion protein of Ubiquitin-like protein SMT3 and Stimulator of interferon protein c-terminal domain | | Authors: | Lesburg, C.A. | | Deposit date: | 2019-10-06 | | Release date: | 2020-08-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | An orally available non-nucleotide STING agonist with antitumor activity.
Science, 369, 2020
|
|
6UL0
 
 | | STING C-terminal Domain Complexed with Non-cyclic Dinucleotide Compound 4 | | Descriptor: | 4-{5-[(1Z)-3-{[2-(3-carboxypropanoyl)-6-methoxy-1-benzothiophen-5-yl]oxy}prop-1-en-1-yl]-6-methoxy-1-benzothiophen-2-yl}-4-oxobutanoic acid, fusion protein of Ubiquitin-like protein SMT3 and Stimulator of interferon protein c-terminal domain | | Authors: | Lesburg, C.A. | | Deposit date: | 2019-10-06 | | Release date: | 2020-08-19 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | An orally available non-nucleotide STING agonist with antitumor activity.
Science, 369, 2020
|
|
4XHV
 
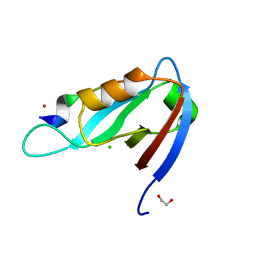 | | Crystal structure of Drosophila Spinophilin-PDZ and a C-terminal peptide of Neurexin | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, LP20995p, ... | | Authors: | Driller, J.H, Muhammad, K.G.H, Reddy, S, Rey, U, Boehme, M.A, Hollmann, C, Ramesh, N, Depner, H, Luetzkendorf, J, Matkovic, T, Bergeron, D, Quentin, C, Schmoranzer, J, Goettfert, F, Holt, M, Wahl, M.C, Hell, S.W, Walter, A, Sigrist, S.J, Loll, B. | | Deposit date: | 2015-01-06 | | Release date: | 2015-07-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.23 Å) | | Cite: | Presynaptic spinophilin tunes neurexin signalling to control active zone architecture and function.
Nat Commun, 6, 2015
|
|
2LM7
 
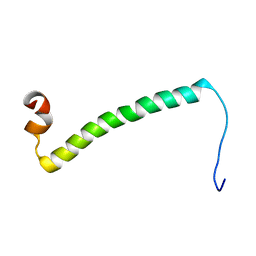 | | NMR structure of the C-terminal domain of VP7 in membrane mimicking micelles | | Descriptor: | Outer capsid glycoprotein VP7 | | Authors: | Elaid, S, Libersou, S, Ouldali, M, Morellet, N, Lepault, J, Bouaziz, S. | | Deposit date: | 2011-11-23 | | Release date: | 2012-10-24 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the C-terminal domain of VP7 in membrane mimicking micelles
To be Published
|
|
7STE
 
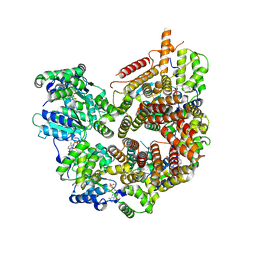 | | Rad24-RFC ADP state | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, Checkpoint protein RAD24, ... | | Authors: | Castaneda, J.C, Schrecker, M, Remus, D, Hite, R.K. | | Deposit date: | 2021-11-12 | | Release date: | 2022-04-06 | | Last modified: | 2024-06-05 | | Method: | ELECTRON MICROSCOPY (2.73 Å) | | Cite: | Mechanisms of loading and release of the 9-1-1 checkpoint clamp.
Nat.Struct.Mol.Biol., 29, 2022
|
|
7CSI
 
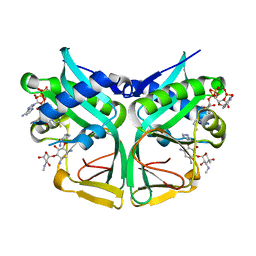 | | Aminoglycoside 2'-N-acetyltransferase from Mycolicibacterium smegmatis-Complex with Coenzyme A and Sisomicin | | Descriptor: | (1S,2S,3R,4S,6R)-4,6-diamino-3-{[(2S,3R)-3-amino-6-(aminomethyl)-3,4-dihydro-2H-pyran-2-yl]oxy}-2-hydroxycyclohexyl 3-deoxy-4-C-methyl-3-(methylamino)-beta-L-arabinopyranoside, Aminoglycoside 2'-N-acetyltransferase, COENZYME A | | Authors: | Jeong, C.S, Hwang, J, Do, H, Lee, J.H. | | Deposit date: | 2020-08-14 | | Release date: | 2021-06-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Structural and biochemical analyses of an aminoglycoside 2'-N-acetyltransferase from Mycolicibacterium smegmatis.
Sci Rep, 10, 2020
|
|
9GLG
 
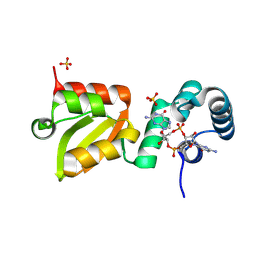 | | X-ray structure of the Thermus thermophilus Q218E mutant of the PilF-GSPIIB domain in the c-di-GMP bound state | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), ACETATE ION, ATP-binding motif-containing protein pilF, ... | | Authors: | Neissner, K, Woehnert, J. | | Deposit date: | 2024-08-27 | | Release date: | 2025-06-25 | | Last modified: | 2025-07-09 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Protonated Glutamate and Aspartate Side Chains Can Recognize Phosphodiester Groups via Strong and Short Hydrogen Bonds in Biomacromolecular Complexes.
Angew.Chem.Int.Ed.Engl., 64, 2025
|
|
6PEX
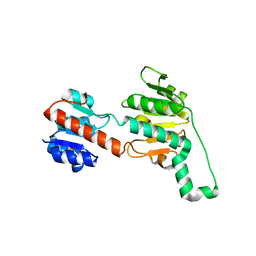 
 | |
6MUD
 
 | |
6S0B
 
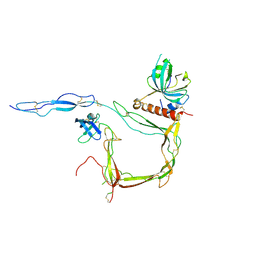 | | Crystal Structure of Properdin in complex with the CTC domain of C3/C3b | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Complement C3, Properdin, ... | | Authors: | van den Bos, R.M, Pearce, N.M, Gros, P. | | Deposit date: | 2019-06-14 | | Release date: | 2019-09-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.312 Å) | | Cite: | Insights Into Enhanced Complement Activation by Structures of Properdin and Its Complex With the C-Terminal Domain of C3b.
Front Immunol, 10, 2019
|
|
6PWK
 
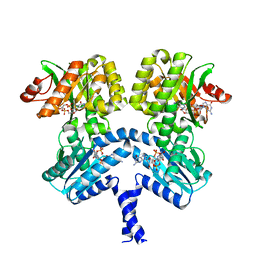 | | Vibrio cholerae LapD S helix-GGDEF-EAL (bound to c-di-GMP) | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), GGDEF and EAL domain-containing protein, MAGNESIUM ION | | Authors: | Giglio, K.M, Cooley, R.B, Sondermann, H. | | Deposit date: | 2019-07-23 | | Release date: | 2019-10-09 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | A Conserved Regulatory Circuit Controls Large Adhesins in Vibrio cholerae.
Mbio, 10, 2019
|
|
8CE8
 
 | | Cytochrome c maturation complex CcmABCDE | | Descriptor: | Cytochrome c biogenesis ATP-binding export protein CcmA, Cytochrome c-type biogenesis protein CcmE, Heme exporter protein B, ... | | Authors: | Ilcu, L, Zhang, L, Einsle, O. | | Deposit date: | 2023-02-01 | | Release date: | 2023-09-06 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (3.81 Å) | | Cite: | Architecture of the Heme-translocating CcmABCD/E complex required for Cytochrome c maturation.
Nat Commun, 14, 2023
|
|
