3LYS
 
 | | Crystal Structure of the N-terminal domain of the Prophage pi2 protein 01 (integrase) from Lactococcus lactis, Northeast Structural Genomics Consortium Target KR124F | | Descriptor: | Prophage pi2 protein 01, integrase | | Authors: | Forouhar, F, Abashidze, M, Seetharaman, J, Sahdev, S, Xiao, R, Ciccosanti, C, Belote, R.L, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-02-28 | | Release date: | 2010-03-16 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Northeast Structural Genomics Consortium Target KR124F
To be Published
|
|
3LYU
 
 | | Crystal Structure of the C-terminal domain (residues 83-215) of PF1911 hydrogenase from Pyrococcus furiosus, Northeast Structural Genomics Consortium Target PfR246A | | Descriptor: | Putative hydrogenase | | Authors: | Forouhar, F, Abashidze, M, Seetharaman, J, Sahdev, S, Xiao, R, Foote, E.L, Ciccosanti, C, Belote, R.L, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-02-28 | | Release date: | 2010-03-23 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Northeast Structural Genomics Consortium Target PfR246A
To be Published
|
|
3MPR
 
 | | Crystal Structure of endonuclease/exonuclease/phosphatase family protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Consortium Target BtR318A | | Descriptor: | ACETIC ACID, BROMIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Kuzin, A, Su, M, Seetharaman, J, Mao, M, Xiao, R, Ciccosanti, C, Lee, D, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-04-27 | | Release date: | 2010-05-26 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (1.899 Å) | | Cite: | Northeast Structural Genomics Consortium Target BtR318A
To be Published
|
|
3MA5
 
 | | Crystal structure of the tetratricopeptide repeat domain protein Q2S6C5_SALRD from Salinibacter ruber. Northeast Structural Genomics Consortium Target SrR115c. | | Descriptor: | Tetratricopeptide repeat domain protein | | Authors: | Vorobiev, S, Neely, H, Seetharaman, J, Wang, H, Foote, E.L, Ciccosanti, C, Mao, L, Xiao, R, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-03-23 | | Release date: | 2010-04-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the tetratricopeptide repeat domain protein Q2S6C5_SALRD from Salinibacter ruber.
To be Published
|
|
3MJQ
 
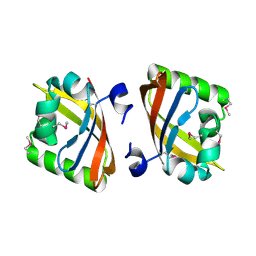 | | Crystal structure of the PAS domain of Q24QT8_DESHY protein from Desulfitobacterium hafniense. Northeast Structural Genomics Consortium Target DhR85c. | | Descriptor: | uncharacterized protein | | Authors: | Vorobiev, S, Neely, H, Seetharaman, J, Wang, D, Ciccosanti, C, Mao, L, Xiao, R, Acton, T.B, Everett, J.K, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-04-13 | | Release date: | 2010-04-28 | | Last modified: | 2021-10-06 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Crystal structure of the PAS domain of Q24QT8_DESHY protein from Desulfitobacterium hafniense.
To be Published
|
|
3MFB
 
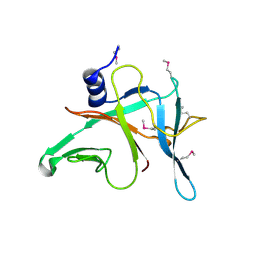 | | Crystal Structure of the S-type Pyocin domain of ECA1669 protein from Erwinia carotovora, Northeast Structural Genomics Consortium Target EwR82C | | Descriptor: | Uncharacterized protein | | Authors: | Forouhar, F, Neely, H, Seetharaman, J, Sahdev, S, Xiao, R, Ciccosanti, C, Lee, D, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-04-01 | | Release date: | 2010-04-14 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Northeast Structural Genomics Consortium Target EwR82C
To be Published
|
|
3MEJ
 
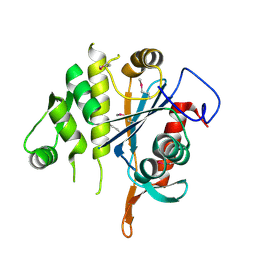 | | Crystal structure of putative transcriptional regulator ywtF from Bacillus subtilis, Northeast Structural Genomics Consortium Target SR736 | | Descriptor: | CHLORIDE ION, transcriptional regulator ywtF | | Authors: | Kuzin, A, Su, M, Seetharaman, J, Mao, M, Xiao, R, Ciccosanti, C, Lee, D, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-03-31 | | Release date: | 2010-04-28 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (2.491 Å) | | Cite: | Northeast Structural Genomics Consortium Target SR736
To be Published
|
|
3MEL
 
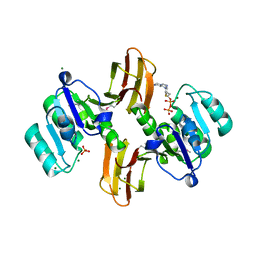 | | Crystal Structure of Thiamin pyrophosphokinase family protein from Enterococcus faecalis, Northeast Structural Genomics Consortium Target EfR150 | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, THIAMINE DIPHOSPHATE, ... | | Authors: | Kuzin, A, Abasidze, M, Seetharaman, J, Mao, M, Xiao, R, Ciccosanti, C, Foote, E.L, Maglaqui, M, Zhao, L, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-03-31 | | Release date: | 2010-05-05 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.788 Å) | | Cite: | Northeast Structural Genomics Consortium Target EfR150
To be Published
|
|
6ZC2
 
 | | Crystal structure of RahU protein in complex with TRIS molecule | | Descriptor: | 1,2-ETHANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, RahU protein | | Authors: | Podobnik, M, Anderluh, G, Lenarcic, T. | | Deposit date: | 2020-06-09 | | Release date: | 2021-04-07 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | Crystal structure of RahU, an aegerolysin protein from the human pathogen Pseudomonas aeruginosa, and its interaction with membrane ceramide phosphorylethanolamine.
Sci Rep, 11, 2021
|
|
6ZC1
 
 | |
2X8B
 
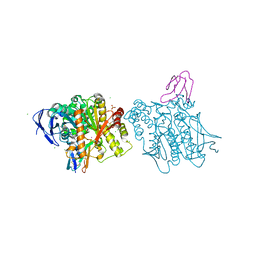 | | Crystal structure of human acetylcholinesterase inhibited by aged tabun and complexed with fasciculin-II | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETHYLCHOLINESTERASE, CHLORIDE ION, ... | | Authors: | Carletti, E, Colletier, J.P, Nachon, F. | | Deposit date: | 2010-03-08 | | Release date: | 2010-04-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural evidence that human acetylcholinesterase inhibited by tabun ages through O-dealkylation.
J. Med. Chem., 53, 2010
|
|
2C1H
 
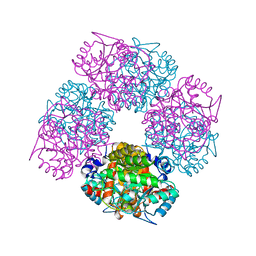 | | The X-ray Structure of Chlorobium vibrioforme 5-Aminolaevulinic Acid Dehydratase Complexed with a Diacid Inhibitor | | Descriptor: | 4,7-DIOXOSEBACIC ACID, DELTA-AMINOLEVULINIC ACID DEHYDRATASE, MAGNESIUM ION | | Authors: | Coates, L, Beaven, G, Erskine, P.T, Beale, S, Wood, S.P, Shoolingin-Jordan, P.M, Cooper, J.B. | | Deposit date: | 2005-09-14 | | Release date: | 2005-12-02 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of Chlorobium Vibrioforme 5-Aminolaevulinic Acid Dehydratase Complexed with a Diacid Inhibitor.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
3CEU
 
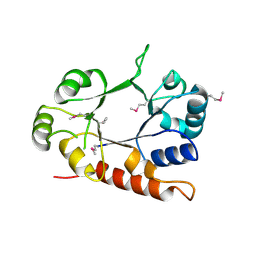 | | Crystal structure of thiamine phosphate pyrophosphorylase (BT_0647) from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR268 | | Descriptor: | Thiamine phosphate pyrophosphorylase | | Authors: | Forouhar, F, Chen, Y, Seetharaman, J, Janjua, H, Mao, L, Xiao, R, Foote, E.L, Maglaqui, M, Ciccosanti, C, Zhao, L, Baran, M.C, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-02-29 | | Release date: | 2008-03-25 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of thiamine phosphate pyrophosphorylase (BT_0647) from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR268.
To be Published
|
|
3CQ9
 
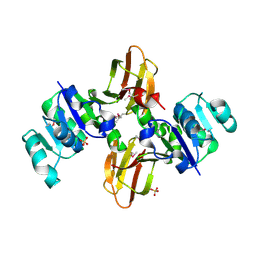 | | Crystal structure of the lp_1622 protein from Lactobacillus plantarum. Northeast Structural Genomics Consortium target LpR114 | | Descriptor: | SULFATE ION, Uncharacterized protein lp_1622 | | Authors: | Vorobiev, S.M, Abashidze, M, Seetharaman, J, Zhao, L, Maglaqui, M, Foote, E.L, Ciccosanti, C, Janjua, H, Xiao, R, Acton, T.B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-04-02 | | Release date: | 2008-04-15 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the lp_1622 protein from Lactobacillus plantarum.
To be Published
|
|
3DC7
 
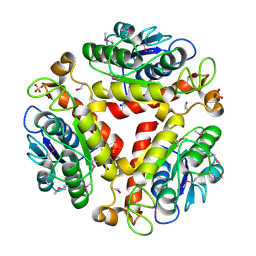 | | Crystal structure of the protein Q88SR8 from Lactobacillus plantarum. Northeast Structural Genomics consortium target LpR109. | | Descriptor: | MAGNESIUM ION, Putative uncharacterized protein lp_3323, SODIUM ION, ... | | Authors: | Kuzin, A.P, Abashidze, M, Vorobiev, S.M, Seetharaman, J, Zhao, L, Mao, L, Ciccosanti, C, Xiao, R, Nair, R, Baran, M.C, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-03 | | Release date: | 2008-08-05 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Crystal structure of the protein Q88SR8 from Lactobacillus plantarum. Northeast Structural Genomics consortium target LpR109. (CASP Target)
To be Published
|
|
3DEV
 
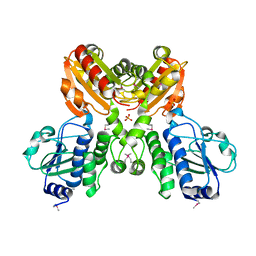 | | Crystal structure of SH1221 protein from Staphylococcus haemolyticus, Northeast Structural Genomics Consortium Target ShR87 | | Descriptor: | MAGNESIUM ION, SH1221, SULFATE ION | | Authors: | Forouhar, F, Neely, H, Seetharaman, J, Mao, L, Xiao, R, Ciccosanti, C, Foote, E.L, Wang, D, Everett, J.K, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-10 | | Release date: | 2008-08-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of SH1221 protein from Staphylococcus haemolyticus, Northeast Structural Genomics Consortium Target ShR87
To be Published
|
|
3DHN
 
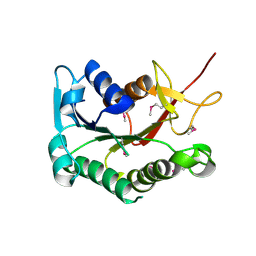 | | Crystal structure of the putative epimerase Q89Z24_BACTN from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR310. | | Descriptor: | NAD-dependent epimerase/dehydratase | | Authors: | Vorobiev, S.M, Su, M, Seetharaman, J, Wang, D, Ciccosanti, C, Foote, L.E, Janjua, H, Xiao, R, Acton, T, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-18 | | Release date: | 2008-08-12 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the putative epimerase Q89Z24_BACTN from Bacteroides thetaiotaomicron.
To be Published
|
|
5OVM
 
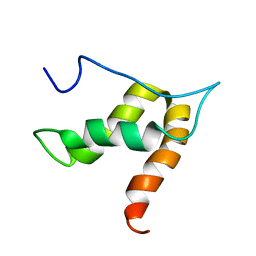 | | Solution structure of lipase binding domain LID1 of foldase from Pseudomonas aeruginosa | | Descriptor: | Lipase chaperone | | Authors: | Viegas, A, Jaeger, K.-E, Etzkorn, M, Gohlke, H, Verma, N, Dollinger, P, Kovacic, F. | | Deposit date: | 2017-08-29 | | Release date: | 2018-12-12 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural and dynamic insights revealing how lipase binding domain MD1 of Pseudomonas aeruginosa foldase affects lipase activation.
Sci Rep, 10, 2020
|
|
6R5M
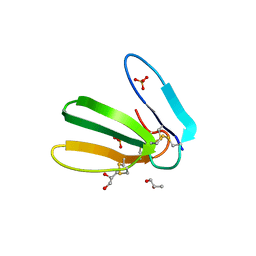 
 | | Crystal structure of toxin MT9 from mamba venom | | Descriptor: | ACETYL GROUP, Dendroaspis polylepis MT9, GLYCEROL, ... | | Authors: | Stura, E.A, Tepshi, L, Ciolek, J, Triquigneaux, M, Zoukimian, C, De Waard, M, Beroud, R, Servent, D, Gilles, N, Legrand, P, Ciccone, L. | | Deposit date: | 2019-03-25 | | Release date: | 2020-02-12 | | Last modified: | 2022-05-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | MT9, a natural peptide from black mamba venom antagonizes the muscarinic type 2 receptor and reverses the M2R-agonist-induced relaxation in rat and human arteries
Biomed Pharmacother, 150, 2022
|
|
4UUQ
 
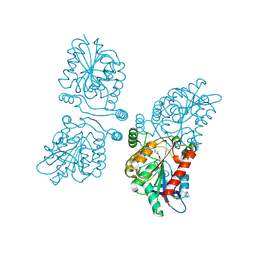 | | Crystal structure of human mono-glyceride lipase in complex with SAR127303 | | Descriptor: | 4-({[(4-chlorophenyl)sulfonyl]amino}methyl)piperidine-1-carboxylic acid, MONOGLYCERIDE LIPASE | | Authors: | Griebel, G, Pichat, P, Beeske, S, Leroy, T, Redon, N, Francon, D, Bert, L, Even, L, Lopez-Grancha, M, Tolstykh, T, Sun, F, Yu, Q, Brittain, S, Arlt, H, He, T, Zhang, B, Wiederschain, D, Bertrand, T, Houtman, J, Rak, A, Vallee, F, Michot, N, Auge, F, Menet, V, Bergis, O.E, George, P, Avenet, P, Mikol, V, Didier, M, Escoubet, J. | | Deposit date: | 2014-07-30 | | Release date: | 2015-01-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Selective Blockade of the Hydrolysis of the Endocannabinoid 2-Arachidonoylglycerol Impairs Learning and Memory Performance While Producing Antinociceptive Activity in Rodents.
Sci.Rep., 5, 2015
|
|
1KI6
 
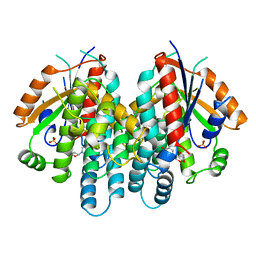 | | CRYSTAL STRUCTURE OF THYMIDINE KINASE FROM HERPES SIMPLEX VIRUS TYPE I COMPLEXED WITH A 5-IODOURACIL ANHYDROHEXITOL NUCLEOSIDE | | Descriptor: | 1',5'-ANHYDRO-2',3'-DIDEOXY-2'-(5-IODOURACIL-1-YL)-D-ABABINO-HEXITOL, SULFATE ION, THYMIDINE KINASE | | Authors: | Champness, J.N, Bennett, M.S, Wien, F, Herdewijn, P, Ostrowski, T, Summers, W.C, Sanderson, M.R. | | Deposit date: | 1998-05-18 | | Release date: | 1998-12-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Exploring the active site of herpes simplex virus type-1 thymidine kinase by X-ray crystallography of complexes with aciclovir and other ligands.
Proteins, 32, 1998
|
|
6RPN
 
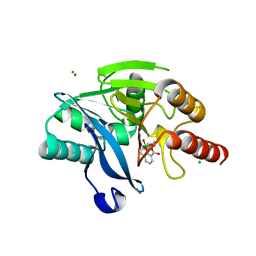 | | Structure of metallo beta lactamase VIM-2 with cyclic boronate APC308. | | Descriptor: | (3~{R})-2,2-bis(oxidanyl)-3-(phenylmethylsulfanyl)-3,4-dihydro-1,2-benzoxaborinin-2-ium-8-carboxylic acid, (3~{S})-2,2-bis(oxidanyl)-3-(phenylmethylsulfanyl)-3,4-dihydro-1,2-benzoxaborinin-2-ium-8-carboxylic acid, Beta-lactamase VIM-2, ... | | Authors: | Parkova, A, Lucic, A, Brem, J, McDonough, M.A, Langley, G.W, Schofield, C.J. | | Deposit date: | 2019-05-14 | | Release date: | 2020-03-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.409 Å) | | Cite: | Broad Spectrum beta-Lactamase Inhibition by a Thioether Substituted Bicyclic Boronate.
Acs Infect Dis., 6, 2020
|
|
6S16
 
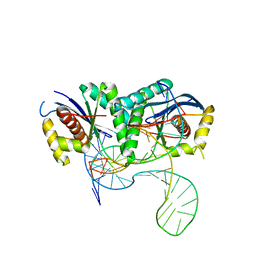 | | T. thermophilus RuvC in complex with Holliday junction substrate | | Descriptor: | CHLORIDE ION, Crossover junction endodeoxyribonuclease RuvC, DNA (33-MER), ... | | Authors: | Gorecka, K.M, Krepl, M, Szlachcic, A, Poznanski, J, Sponer, J, Nowotny, M. | | Deposit date: | 2019-06-18 | | Release date: | 2019-09-25 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (3.409 Å) | | Cite: | RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution.
Nat Commun, 10, 2019
|
|
5OPW
 
 | | Crystal structure of the GroEL mutant A109C | | Descriptor: | 60 kDa chaperonin | | Authors: | Yan, X, Shi, Q, Bracher, A, Milicic, G, Singh, A.K, Hartl, F.U, Hayer-Hartl, M. | | Deposit date: | 2017-08-10 | | Release date: | 2018-01-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | GroEL Ring Separation and Exchange in the Chaperonin Reaction.
Cell, 172, 2018
|
|
5OPX
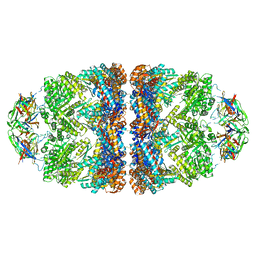 
 | | Crystal structure of the GroEL mutant A109C in complex with GroES and ADP BeF2 | | Descriptor: | 10 kDa chaperonin, 60 kDa chaperonin, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Yan, X, Shi, Q, Bracher, A, Milicic, G, Singh, A.K, Hartl, F.U, Hayer-Hartl, M. | | Deposit date: | 2017-08-10 | | Release date: | 2018-01-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.64 Å) | | Cite: | GroEL Ring Separation and Exchange in the Chaperonin Reaction.
Cell, 172, 2018
|
|
