5LTL
 
 | |
3D32
 
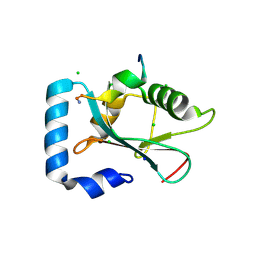 | | Complex of GABA(A) receptor-associated protein (GABARAP) with a synthetic peptide | | Descriptor: | CHLORIDE ION, Gamma-aminobutyric acid receptor-associated protein, K1 peptide, ... | | Authors: | Weiergraeber, O.H, Stangler, T, Willbold, D. | | Deposit date: | 2008-05-09 | | Release date: | 2008-08-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Ligand Binding Mode of GABA(A) Receptor-Associated Protein.
J.Mol.Biol., 381, 2008
|
|
5C6M
 
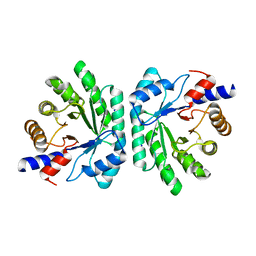 | | Crystal structure of deoxyribose-phosphate aldolase from Shewanella halifaxensis | | Descriptor: | CHLORIDE ION, Deoxyribose-phosphate aldolase, SODIUM ION | | Authors: | Weiergraeber, O.H, Dick, M, Bramski, J, Pietruszka, J. | | Deposit date: | 2015-06-23 | | Release date: | 2016-02-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Trading off stability against activity in extremophilic aldolases.
Sci Rep, 6, 2016
|
|
5C5Y
 
 | |
4CO7
 
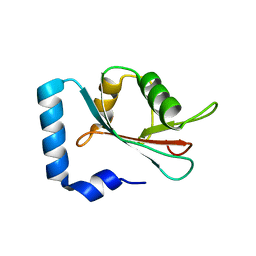 | | Crystal structure of human GATE-16 | | Descriptor: | GAMMA-AMINOBUTYRIC ACID RECEPTOR-ASSOCIATED PROTEIN-LIKE 2 | | Authors: | Weiergraeber, O.H, Ma, P, Willbold, D. | | Deposit date: | 2014-01-27 | | Release date: | 2015-01-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Conformational Polymorphism in Autophagy-Related Protein Gate-16.
Biochemistry, 54, 2015
|
|
5EMU
 
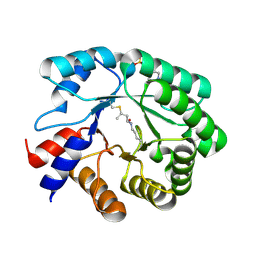 | |
5EL1
 
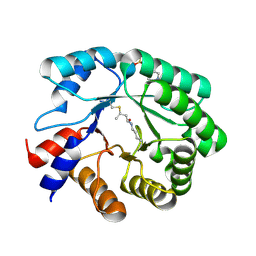 | |
7ZGT
 
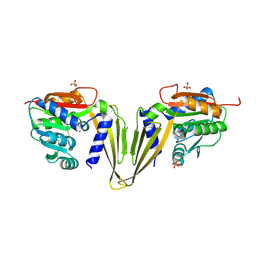 | | C-Methyltransferase PsmD from Streptomyces griseofuscus (apo form) | | Descriptor: | FORMIC ACID, Methyltransferase, PHOSPHATE ION, ... | | Authors: | Weiergraeber, O.H, Amariei, D.A, Pozhydaieva, N, Pietruszka, J. | | Deposit date: | 2022-04-04 | | Release date: | 2022-12-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Enzymatic C3-Methylation of Indoles Using Methyltransferase PsmD-Crystal Structure, Catalytic Mechanism, and Preparative Applications
Acs Catalysis, 2022
|
|
7ZKH
 
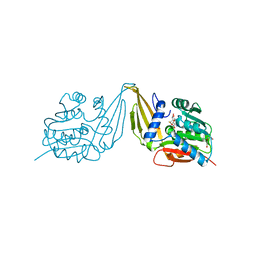 | | C-Methyltransferase PsmD from Streptomyces griseofuscus with bound cofactor (crystal form 1) | | Descriptor: | Methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE, TRIETHYLENE GLYCOL, ... | | Authors: | Weiergraeber, O.H, Amariei, D.A, Pozhydaieva, N, Pietruszka, J. | | Deposit date: | 2022-04-13 | | Release date: | 2022-12-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Enzymatic C3-Methylation of Indoles Using Methyltransferase PsmD-Crystal Structure, Catalytic Mechanism, and Preparative Applications
Acs Catalysis, 2022
|
|
7ZKG
 
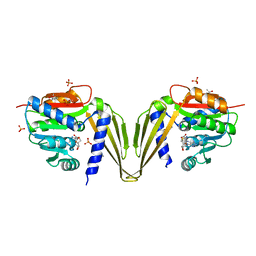 | | C-Methyltransferase PsmD from Streptomyces griseofuscus with bound cofactor (crystal form 2) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Methyltransferase, PHOSPHATE ION, ... | | Authors: | Weiergraeber, O.H, Amariei, D.A, Pozhydaieva, N, Pietruszka, J. | | Deposit date: | 2022-04-13 | | Release date: | 2022-12-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Enzymatic C3-Methylation of Indoles Using Methyltransferase PsmD-Crystal Structure, Catalytic Mechanism, and Preparative Applications
Acs Catalysis, 2022
|
|
8S1M
 
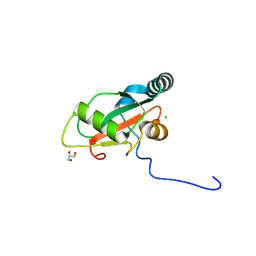 | |
3STO
 
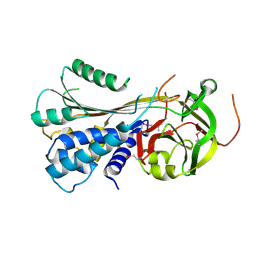 | | Serpin from the trematode Schistosoma Haematobium | | Descriptor: | Serine protease inhibitor | | Authors: | Granzin, J, Weiergraeber, O.H, Lee, X, Blanton, R.E. | | Deposit date: | 2011-07-11 | | Release date: | 2012-05-30 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.41 Å) | | Cite: | Three-dimensional structure of a schistosome serpin revealing an unusual configuration of the helical subdomain.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
7R5Z
 
 | |
4WZG
 
 | | Structure of human ATG101 | | Descriptor: | Autophagy-related protein 101, BETA-MERCAPTOETHANOL | | Authors: | Michel, M, Weiergraeber, O.H. | | Deposit date: | 2014-11-19 | | Release date: | 2015-06-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The mammalian autophagy initiator complex contains 2 HORMA domain proteins.
Autophagy, 11, 2015
|
|
2IF4
 
 | |
7ZL7
 
 | | Human GABARAP in complex with stapled peptide Pen8-ortho | | Descriptor: | CHLORIDE ION, Gamma-aminobutyric acid receptor-associated protein, ORTHO-XYLENE, ... | | Authors: | Ueffing, A, Brown, H, Willbold, D, Kritzer, J.A, Weiergraeber, O.H. | | Deposit date: | 2022-04-14 | | Release date: | 2022-08-31 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure-Based Design of Stapled Peptides That Bind GABARAP and Inhibit Autophagy.
J.Am.Chem.Soc., 144, 2022
|
|
7ZKR
 
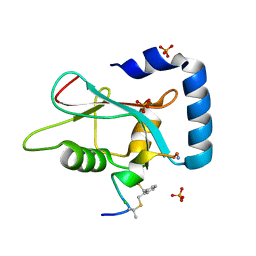 | | Human GABARAP in complex with stapled peptide Pen3-ortho | | Descriptor: | CHLORIDE ION, Gamma-aminobutyric acid receptor-associated protein, ORTHO-XYLENE, ... | | Authors: | Ueffing, A, Brown, H, Willbold, D, Kritzer, J.A, Weiergraeber, O.H. | | Deposit date: | 2022-04-13 | | Release date: | 2022-08-31 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structure-Based Design of Stapled Peptides That Bind GABARAP and Inhibit Autophagy.
J.Am.Chem.Soc., 144, 2022
|
|
2NCN
 
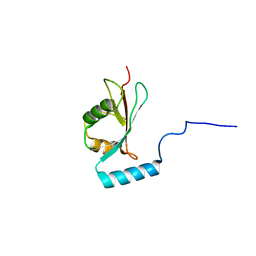 | |
3DOW
 
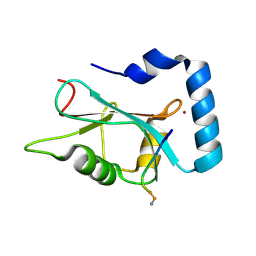 | |
5C2X
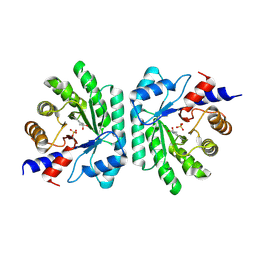 
 | | Crystal structure of deoxyribose-phosphate aldolase from Colwellia psychrerythraea (tetragonal form) | | Descriptor: | CARBONATE ION, Deoxyribose-phosphate aldolase, SULFATE ION, ... | | Authors: | Dick, M, Weiergraeber, O.H, Pietruszka, J. | | Deposit date: | 2015-06-16 | | Release date: | 2016-02-03 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Trading off stability against activity in extremophilic aldolases.
Sci Rep, 6, 2016
|
|
5EKY
 
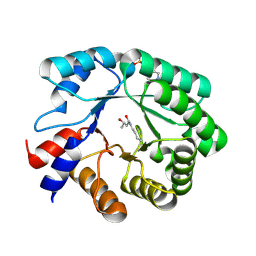 | | Crystal structure of deoxyribose-phosphate aldolase from Escherichia coli (K58E-Y96W mutant) | | Descriptor: | 1,3-BUTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Deoxyribose-phosphate aldolase | | Authors: | Classen, T, Dick, M, Pietruszka, J, Weiergraeber, O.H. | | Deposit date: | 2015-11-04 | | Release date: | 2016-05-04 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Mechanism-based inhibition of an aldolase at high concentrations of its natural substrate acetaldehyde: structural insights and protective strategies.
Chem Sci, 7, 2016
|
|
7YWG
 
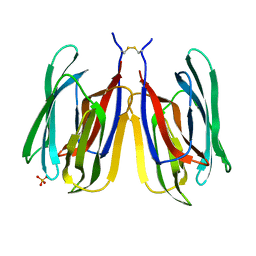 | |
7YWE
 
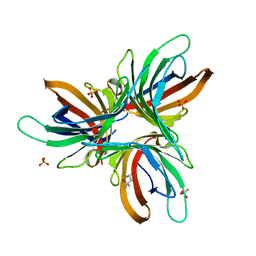 | | Monocot chimeric jacalin JAC1 from Oryza sativa: dirigent domain (crystal form 2) | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Dirigent protein, PHOSPHATE ION | | Authors: | Huwa, N, Classen, T, Weiergraeber, O.H. | | Deposit date: | 2022-02-13 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | The Crystal Structure of the Defense Conferring Rice Protein Os JAC1 Reveals a Carbohydrate Binding Site on the Dirigent-like Domain.
Biomolecules, 12, 2022
|
|
7YWF
 
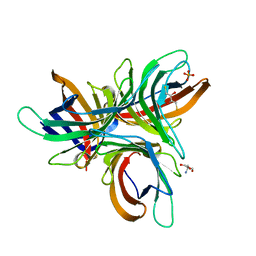 | | Monocot chimeric jacalin JAC1 from Oryza sativa: dirigent domain with bound galactobiose | | Descriptor: | (2R,3S)-1,4-DIMERCAPTOBUTANE-2,3-DIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Dirigent protein, ... | | Authors: | Huwa, N, Classen, T, Weiergraeber, O.H. | | Deposit date: | 2022-02-13 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Crystal Structure of the Defense Conferring Rice Protein Os JAC1 Reveals a Carbohydrate Binding Site on the Dirigent-like Domain.
Biomolecules, 12, 2022
|
|
7YWW
 
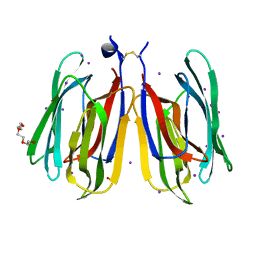 | | Monocot chimeric jacalin JAC1 from Oryza sativa: lectin domain (crystal form 2) | | Descriptor: | Dirigent protein, GLYCEROL, IODIDE ION, ... | | Authors: | Huwa, N, Classen, T, Weiergraeber, O.H. | | Deposit date: | 2022-02-14 | | Release date: | 2022-05-04 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The Crystal Structure of the Defense Conferring Rice Protein Os JAC1 Reveals a Carbohydrate Binding Site on the Dirigent-like Domain.
Biomolecules, 12, 2022
|
|
