1G0Y
 
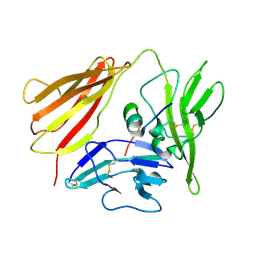 | | IL-1 RECEPTOR TYPE 1 COMPLEXED WITH ANTAGONIST PEPTIDE AF10847 | | Descriptor: | ANTAGONIST PEPTIDE AF10847, INTERLEUKIN-1 RECEPTOR, TYPE I | | Authors: | Vigers, G.P.A, Dripps, D.J, Edwards, C.K, Brandhuber, B.J. | | Deposit date: | 2000-10-09 | | Release date: | 2000-10-25 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | X-ray crystal structure of a small antagonist peptide bound to interleukin-1 receptor type 1.
J.Biol.Chem., 275, 2000
|
|
6N2K
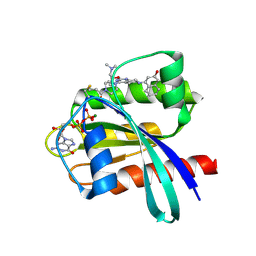 
 | | Tetrahydropyridopyrimidines as Covalent Inhibitors of KRAS-G12C | | Descriptor: | 1-{4-[2-{[(2R)-1-(dimethylamino)propan-2-yl]oxy}-7-(3-hydroxynaphthalen-1-yl)-5,6,7,8-tetrahydropyrido[3,4-d]pyrimidin-4-yl]piperazin-1-yl}propan-1-one, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Vigers, G.P. | | Deposit date: | 2018-11-13 | | Release date: | 2018-12-12 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Discovery of Tetrahydropyridopyrimidines as Irreversible Covalent Inhibitors of KRAS-G12C with In Vivo Activity.
ACS Med Chem Lett, 9, 2018
|
|
1ITB
 
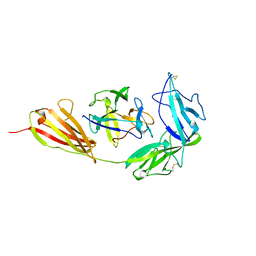 | | TYPE-1 INTERLEUKIN-1 RECEPTOR COMPLEXED WITH INTERLEUKIN-1 BETA | | Descriptor: | INTERLEUKIN-1 BETA, TYPE 1 INTERLEUKIN-1 RECEPTOR | | Authors: | Vigers, G.P.A, Anderson, L.J, Caffes, P, Brandhuber, B.J. | | Deposit date: | 1997-01-15 | | Release date: | 1998-02-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of the type-I interleukin-1 receptor complexed with interleukin-1beta.
Nature, 386, 1997
|
|
6USX
 
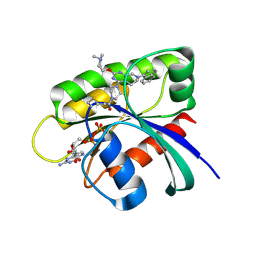 | | Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer | | Descriptor: | 1-{4-[2-{[(2S)-1-methylpyrrolidin-2-yl]methoxy}-7-(naphthalen-1-yl)-5,6,7,8-tetrahydropyrido[3,4-d]pyrimidin-4-yl]piperazin-1-yl}propan-1-one, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Vigers, G.P, Smith, D.J. | | Deposit date: | 2019-10-28 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Identification of the Clinical Development CandidateMRTX849, a Covalent KRASG12CInhibitor for the Treatment of Cancer.
J.Med.Chem., 63, 2020
|
|
6UT0
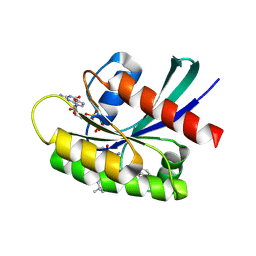 
 | | Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Vigers, G.P, Smith, D.J. | | Deposit date: | 2019-10-29 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Identification of the Clinical Development CandidateMRTX849, a Covalent KRASG12CInhibitor for the Treatment of Cancer.
J.Med.Chem., 63, 2020
|
|
6USZ
 
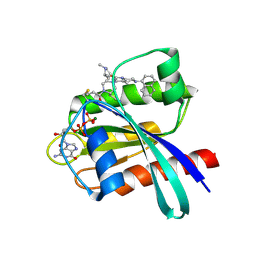 | | Identification of the Clinical Development Candidate MRTX849, a Covalent KRASG12C Inhibitor for the Treatment of Cancer | | Descriptor: | GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, ... | | Authors: | Vigers, G.P, Smith, D.J. | | Deposit date: | 2019-10-28 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Identification of the Clinical Development CandidateMRTX849, a Covalent KRASG12CInhibitor for the Treatment of Cancer.
J.Med.Chem., 63, 2020
|
|
6OPI
 
 | | phosphorylated ERK2 with SCH-CPD336 | | Descriptor: | (3R)-N-[3-(2-cyclopropylpyridin-4-yl)-1H-indazol-5-yl]-3-(methoxymethyl)-1-(2-oxo-2-{4-[4-(pyrimidin-2-yl)phenyl]-3,6-dihydropyridin-1(2H)-yl}ethyl)pyrrolidine-3-carboxamide, Mitogen-activated protein kinase 1 | | Authors: | Vigers, G.P, Smith, D. | | Deposit date: | 2019-04-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Activation loop dynamics are controlled by conformation-selective inhibitors of ERK2.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6OPH
 
 | | phosphorylated ERK2 with GDC-0994 | | Descriptor: | 1-[(1~{S})-1-(4-chloranyl-3-fluoranyl-phenyl)-2-oxidanyl-ethyl]-4-[2-[(2-methylpyrazol-3-yl)amino]pyrimidin-4-yl]pyridin-2-one, Mitogen-activated protein kinase 1 | | Authors: | Vigers, G.P, Smith, D. | | Deposit date: | 2019-04-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Activation loop dynamics are controlled by conformation-selective inhibitors of ERK2.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6OPG
 
 | | phosphorylated ERK2 with AMP-PNP | | Descriptor: | MAGNESIUM ION, Mitogen-activated protein kinase 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Vigers, G.P, Smith, D. | | Deposit date: | 2019-04-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Activation loop dynamics are controlled by conformation-selective inhibitors of ERK2.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6OPK
 
 | | Phosphorylated ERK2 with Vertex-11e | | Descriptor: | 4-{2-[(2-chloro-4-fluorophenyl)amino]-5-methylpyrimidin-4-yl}-N-[(1S)-1-(3-chlorophenyl)-2-hydroxyethyl]-1H-pyrrole-2-carboxamide, Mitogen-activated protein kinase 1 | | Authors: | Vigers, G.P, Rudolph, J. | | Deposit date: | 2019-04-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Activation loop dynamics are controlled by conformation-selective inhibitors of ERK2.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
6N2J
 | | Tetrahydropyridopyrimidines as Covalent Inhibitors of KRAS-G12C | | Descriptor: | 1-{4-[7-(naphthalen-1-yl)-5,6,7,8-tetrahydropyrido[3,4-d]pyrimidin-4-yl]piperazin-1-yl}propan-1-one, GTPase KRas, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Vigers, G.P. | | Deposit date: | 2018-11-13 | | Release date: | 2018-12-12 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Discovery of Tetrahydropyridopyrimidines as Irreversible Covalent Inhibitors of KRAS-G12C with In Vivo Activity.
ACS Med Chem Lett, 9, 2018
|
|
4PZX
 
 | | Synthesis, Characterization and PK/PD Studies of a Series of Spirocyclic Pyranochromene BACE1 Inhibitors | | Descriptor: | (4R,4a'R,10a'R)-7'-(5-chloropyridin-3-yl)-3',4',4a',10a'-tetrahydro-1'H-spiro[1,3-oxazole-4,5'-pyrano[3,4-b]chromen]-2-amine, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A. | | Deposit date: | 2014-03-31 | | Release date: | 2014-05-14 | | Last modified: | 2014-10-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Synthesis, characterization, and PK/PD studies of a series of spirocyclic pyranochromene BACE1 inhibitors.
Bioorg.Med.Chem.Lett., 24, 2014
|
|
4PZW
 
 | | Synthesis, Characterization and PK/PD Studies of a Series of Spirocyclic Pyranochromene BACE1 Inhibitors | | Descriptor: | (4R,4a'S,10a'S)-7'-(5-chloropyridin-3-yl)-3',4',4a',10a'-tetrahydro-1'H-spiro[1,3-oxazole-4,5'-pyrano[3,4-b]chromen]-2-amine, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A. | | Deposit date: | 2014-03-31 | | Release date: | 2014-05-14 | | Last modified: | 2014-10-15 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Synthesis, characterization, and PK/PD studies of a series of spirocyclic pyranochromene BACE1 inhibitors.
Bioorg.Med.Chem.Lett., 24, 2014
|
|
4R5N
 
 | | 8-Tetrahydropyran-2-yl chromans: highly selective beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors | | Descriptor: | (4R,4a'S,10a'R)-8'-(2-fluoropyridin-3-yl)-4a'-methyl-3',4',4a',10a'-tetrahydro-2'H-spiro[1,3-oxazole-4,10'-pyrano[3,2-b]chromen]-2-amine, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2014-08-21 | | Release date: | 2014-12-03 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | 8-Tetrahydropyran-2-yl Chromans: Highly Selective Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors.
J.Med.Chem., 57, 2014
|
|
4N00
 
 | | Discovery of 7-THP chromans: BACE1 inhibitors that reduce A-beta in the CNS | | Descriptor: | (4R,4a'S,10a'S)-2-amino-8'-(2-fluoropyridin-3-yl)-1-methyl-3',4',4a',10a'-tetrahydro-1'H-spiro[imidazole-4,10'-pyrano[4,3-b]chromen]-5(1H)-one, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2013-09-30 | | Release date: | 2014-05-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Discovery of 7-tetrahydropyran-2-yl chromans: beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors that reduce amyloid beta-protein (A beta ) in the central nervous system.
J.Med.Chem., 57, 2014
|
|
4JOO
 | | Spirocyclic Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors | | Descriptor: | (4R)-2'-amino-6-bromo-1',2,2-trimethyl-2,3-dihydrospiro[chromene-4,4'-imidazol]-5'(1'H)-one, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2013-03-18 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Spirocyclic beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors: from hit to lowering of cerebrospinal fluid (CSF) amyloid beta in a higher species.
J.Med.Chem., 56, 2013
|
|
4JPE
 | | Spirocyclic Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors | | Descriptor: | (4R)-2-amino-1,3',3'-trimethyl-7'-(pyrimidin-5-yl)-3',4'-dihydro-2'H-spiro[imidazole-4,1'-naphthalen]-5(1H)-one, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2013-03-19 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Spirocyclic beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors: from hit to lowering of cerebrospinal fluid (CSF) amyloid beta in a higher species.
J.Med.Chem., 56, 2013
|
|
4JPC
 
 | | Spirocyclic Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors | | Descriptor: | 3-[(4R)-2'-amino-1',2,2-trimethyl-5'-oxo-1',2,3,5'-tetrahydrospiro[chromene-4,4'-imidazol]-6-yl]benzonitrile, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2013-03-19 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Spirocyclic beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors: from hit to lowering of cerebrospinal fluid (CSF) amyloid beta in a higher species.
J.Med.Chem., 56, 2013
|
|
4JP9
 | | Spirocyclic Beta-Site Amyloid Precursor Protein Cleaving Enzyme 1 (BACE1) Inhibitors | | Descriptor: | (4R)-2'-amino-6-(3-chlorophenyl)-1',2,2-trimethyl-2,3-dihydrospiro[chromene-4,4'-imidazol]-5'(1'H)-one, Beta-secretase 1, NICKEL (II) ION | | Authors: | Vigers, G.P.A, Smith, D. | | Deposit date: | 2013-03-19 | | Release date: | 2013-04-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Spirocyclic beta-site amyloid precursor protein cleaving enzyme 1 (BACE1) inhibitors: from hit to lowering of cerebrospinal fluid (CSF) amyloid beta in a higher species.
J.Med.Chem., 56, 2013
|
|
3D4Q
 
 | | Pyrazole-based inhibitors of B-Raf kinase | | Descriptor: | (1E)-5-(1-piperidin-4-yl-3-pyridin-4-yl-1H-pyrazol-4-yl)-2,3-dihydro-1H-inden-1-one oxime, B-Raf proto-oncogene serine/threonine-protein kinase | | Authors: | Morales, T, Vigers, G.P.A, Brandhuber, B.J. | | Deposit date: | 2008-05-14 | | Release date: | 2008-08-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Potent and selective pyrazole-based inhibitors of B-Raf kinase.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
8U8J
 
 | |
8U8K
 
 | |
4EKL
 
 | | Akt1 with GDC0068 | | Descriptor: | (2S)-2-(4-chlorophenyl)-1-{4-[(5R,7R)-7-hydroxy-5-methyl-6,7-dihydro-5H-cyclopenta[d]pyrimidin-4-yl]piperazin-1-yl}-3-(propan-2-ylamino)propan-1-one, RAC-alpha serine/threonine-protein kinase | | Authors: | Wu, W.-I, Vigers, G.P.A, Morales, T.H, Brandhuber, B.J. | | Deposit date: | 2012-04-09 | | Release date: | 2012-05-23 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | An ATP-Site On-Off Switch That Restricts Phosphatase Accessibility of Akt.
Sci.Signal., 5, 2012
|
|
1ILT
 
 | |
4KTC
 
 | | NS3/NS4A protease with inhibitor | | Descriptor: | (2R,6S,13aR,14aR,16aS)-6-{[(cyclopentyloxy)carbonyl]amino}-14a-[(cyclopropylsulfonyl)carbamoyl]-5,16-dioxooctadecahydrocyclopropa[e]pyrrolo[1,2-a][1,4]diazacyclopentadecin-2-yl 3,4-dihydroisoquinoline-2(1H)-carboxylate, NS4A peptide, Serine protease NS3, ... | | Authors: | Zhang, H, Ballard, J, Vigers, G.P.A, Brandhuber, B.J. | | Deposit date: | 2013-05-20 | | Release date: | 2013-08-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery of Danoprevir (ITMN-191/R7227), a Highly Selective and Potent Inhibitor of Hepatitis C Virus (HCV) NS3/4A Protease.
J.Med.Chem., 57, 2014
|
|
