1IIQ
 
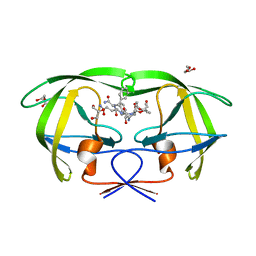 | | CRYSTAL STRUCTURE OF HIV-1 PROTEASE COMPLEXED WITH A HYDROXYETHYLAMINE PEPTIDOMIMETIC INHIBITOR | | Descriptor: | GLYCEROL, N-{(2R,3S)-3-[(tert-butoxycarbonyl)amino]-2-hydroxy-4-phenylbutyl}-L-phenylalanyl-L-glutaminyl-L-phenylalaninamide, PROTEASE RETROPEPSIN | | Authors: | Dohnalek, J, Hasek, J, Duskova, J, Petrokova, H, Hradilek, M, Soucek, M, Konvalinka, J, Brynda, J, Sedlacek, J, Fabry, M. | | Deposit date: | 2001-04-24 | | Release date: | 2002-04-12 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Hydroxyethylamine isostere of an HIV-1 protease inhibitor prefers its amine to the hydroxy group in binding to catalytic aspartates. A synchrotron study of HIV-1 protease in complex with a peptidomimetic inhibitor.
J.Med.Chem., 45, 2002
|
|
1ZTZ
 
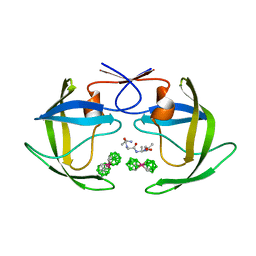 | | Crystal structure of HIV protease- metallacarborane complex | | Descriptor: | COBALT BIS(1,2-DICARBOLLIDE), PROTEASE RETROPEPSIN, autoproteolytic tetrapeptide | | Authors: | Cigler, P, Kozisek, M, Rezacova, P, Brynda, J, Otwinowski, Z, Sedlacek, J, Bodem, J, Kraeusslich, H.-G, Kral, V, Konvalinka, J. | | Deposit date: | 2005-05-28 | | Release date: | 2005-11-01 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | From nonpeptide toward noncarbon protease inhibitors: Metallacarboranes as specific and potent inhibitors of HIV protease
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1FQX
 
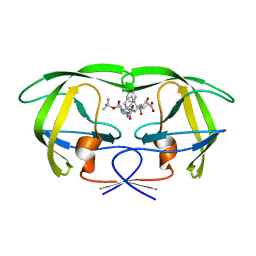 | | CRYSTAL STRUCTURE OF THE COMPLEX OF HIV-1 PROTEASE WITH A PEPTIDOMIMETIC INHIBITOR | | Descriptor: | N-{(2S,3S)-3-[(tert-butoxycarbonyl)amino]-2-hydroxy-4-phenylbutyl}-L-phenylalanyl-L-alpha-glutamyl-L-phenylalaninamide, PROTEASE RETROPEPSIN | | Authors: | Dohnalek, J, Hasek, J, Duskova, J, Petrokova, H, Hradilek, M, Soucek, M, Konvalinka, J, Brynda, J, Sedlacek, J, Fabry, M. | | Deposit date: | 2000-09-07 | | Release date: | 2001-03-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | A distinct binding mode of a hydroxyethylamine isostere inhibitor of HIV-1 protease.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
1JP5
 
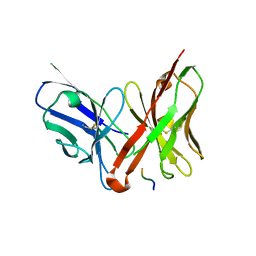 | | Crystal structure of the single-chain Fv fragment 1696 in complex with the epitope peptide corresponding to N-terminus of HIV-1 protease | | Descriptor: | epitope peptide corresponding to N-terminus of HIV-1 protease, single-chain Fv fragment 1696 | | Authors: | Rezacova, P, Lescar, J, Brynda, J, Fabry, M, Horejsi, M, Sedlacek, J, Bentley, G.A. | | Deposit date: | 2001-08-01 | | Release date: | 2001-10-12 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structural basis of HIV-1 and HIV-2 protease inhibition by a monoclonal antibody.
Structure, 9, 2001
|
|
2HKH
 
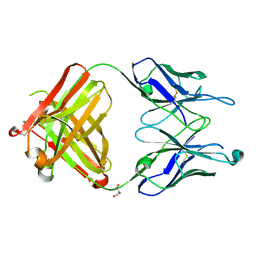 | | Crystal structure of the Fab M75 | | Descriptor: | GLYCEROL, Immunoglobulin Heavy chain Fab fragment, Immunoglobulin Light chain Fab fragment | | Authors: | Kral, V, Mader, P, Stouracova, R, Fabry, M, Sedlacek, J, Brynda, J. | | Deposit date: | 2006-07-04 | | Release date: | 2007-11-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Stabilization of antibody structure upon association to a human carbonic anhydrase IX epitope studied by X-ray crystallography, microcalorimetry, and molecular dynamics simulations.
Proteins, 71, 2008
|
|
1HC0
 
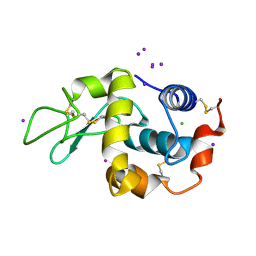 | | structure of lysozyme with periodate | | Descriptor: | CHLORIDE ION, IODIDE ION, LYSOZYME C | | Authors: | Ondracek, J, Jursik, F, Brynda, J, Rezacova, P, Sedlacek, J. | | Deposit date: | 2001-04-25 | | Release date: | 2005-11-14 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Regular Arrangement of Periodates Bound to Lysozyme.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1SP5
 
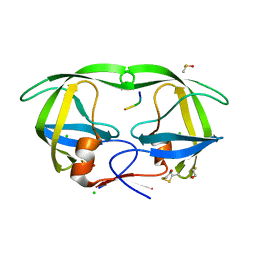 | | Crystal structure of HIV-1 protease complexed with a product of autoproteolysis | | Descriptor: | 5-mer peptide from Protease, BETA-MERCAPTOETHANOL, CHLORIDE ION, ... | | Authors: | Vondrackova, E, Hasek, J, Jaskolski, M, Rezacova, P, Dohnalek, J, Skalova, T, Petrokova, H, Duskova, J, Brynda, J, Sedlacek, J. | | Deposit date: | 2004-03-16 | | Release date: | 2005-07-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Product of enzymatic self-cleavage bound in the active site of HIV protease
To be Published
|
|
1U8G
 
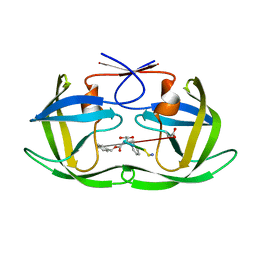 | | Crystal structure of a HIV-1 Protease in complex with peptidomimetic inhibitor KI2-PHE-GLU-GLU-NH2 | | Descriptor: | PROTEASE RETROPEPSIN, peptidomimetic inhibitor KI2-PHE-GLU-GLU-NH2 | | Authors: | Brynda, J, Rezacova, P, Fabry, M, Horejsi, M, Hradilek, M, Soucek, R, Stouracova, R, Konvalinka, J, Sedlacek, J. | | Deposit date: | 2004-08-06 | | Release date: | 2004-11-02 | | Last modified: | 2018-02-14 | | Method: | X-RAY DIFFRACTION (2.201 Å) | | Cite: | Inhibitor binding at the protein interface in crystals of a HIV-1 protease complex.
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1RL8
 
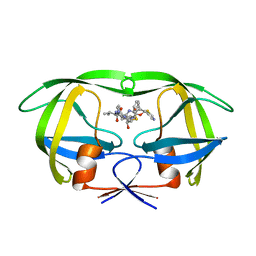 | | Crystal structure of the complex of resistant strain of hiv-1 protease(v82a mutant) with ritonavir | | Descriptor: | RITONAVIR, protease RETROPEPSIN | | Authors: | Rezacova, P, Brynda, J, Sedlacek, J, Konvalinka, J, Fabry, M, Horejsi, M. | | Deposit date: | 2003-11-25 | | Release date: | 2005-04-19 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the complex of resistant strain of hiv-1 protease(v82a mutant) with ritonavir
To be Published, 2005
|
|
1SVZ
 
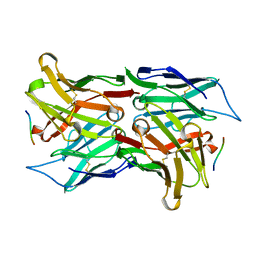 | | Crystal structure of the single-chain Fv fragment 1696 in complex with the epitope peptide corresponding to N-terminus of HIV-2 protease | | Descriptor: | epitope peptide corresponding to N-terminus of HIV-2 protease, single-chain Fv fragment 1696 | | Authors: | Rezacova, P, Brynda, J, Lescar, J, Bentley, G.A, Fabry, M, Horejsi, M, Sedlacek, J. | | Deposit date: | 2004-03-30 | | Release date: | 2005-03-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Crystal structure of a cross-reaction complex between an anti-HIV-1 protease antibody and an HIV-2 protease peptide
J.Struct.Biol., 149, 2005
|
|
1NH0
 
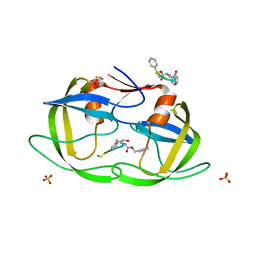 | | 1.03 A structure of HIV-1 protease: inhibitor binding inside and outside the active site | | Descriptor: | BETA-MERCAPTOETHANOL, PROTEASE RETROPEPSIN, SULFATE ION, ... | | Authors: | Brynda, J, Rezacova, P, Fabry, M, Horejsi, M, Hradilek, M, Soucek, M, Konvalinka, J, Sedlacek, J. | | Deposit date: | 2002-12-18 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.03 Å) | | Cite: | A Phenylnorstatine Inhibitor Binding to HIV-1 Protease: Geometry,
Protonation, and Subsite-Pocket Interactions Analyzed at Atomic Resolution
J.Med.Chem., 47, 2004
|
|
1N4X
 
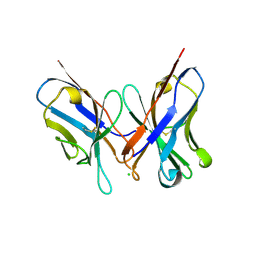 | | Structure of scFv 1696 at acidic pH | | Descriptor: | CHLORIDE ION, immunoglobulin heavy chain variable region, immunoglobulin kappa chain variable region | | Authors: | Lescar, J, Brynda, J, Fabry, M, Horejsi, M, Rezacova, P, Sedlacek, J, Bentley, G.A. | | Deposit date: | 2002-11-02 | | Release date: | 2003-06-10 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure of a single-chain Fv fragment of an antibody that inhibits the HIV-1 and HIV-2 proteases.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
